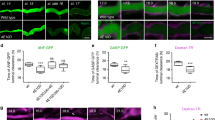
Abstract
Localization of β-actin messenger RNA to sites of active actin polymerization modulates cell migration during embryogenesis, differentiation and possibly carcinogenesis1,2,3,4,5. This localization requires the oncofetal protein ZBP1 (Zipcode binding protein 1), which binds to a conserved 54-nucleotide element in the 3′-untranslated region of the β-actin mRNA known as the ‘zipcode’. ZBP1 promotes translocation of the β-actin transcript to actin-rich protrusions in primary fibroblasts and neurons6,7. It is not known how the ZBP1–RNA complex achieves asymmetric protein sorting by localizing β-actin mRNA. Here we show that chicken ZBP1 modulates the translation of β-actin mRNA. ZBP1 associates with the β-actin transcript in the nucleus and prevents premature translation in the cytoplasm by blocking translation initiation. Translation only occurs when the ZBP1–RNA complex reaches its destination at the periphery of the cell. At the endpoint of mRNA transport, the protein kinase Src promotes translation by phosphorylating a key tyrosine residue in ZBP1 that is required for binding to RNA. These sequential events provide both temporal and spatial control over β-actin mRNA translation, which is important for cell migration and neurite outgrowth.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
References
Lawrence, J. B. & Singer, R. H. Intracellular localization of messenger RNAs for cytoskeletal proteins. Cell 45, 407–415 (1986)
Kislauskis, E. H., Li, Z., Singer, R. H. & Taneja, K. L. Isoform-specific 3′-untranslated sequences sort α-cardiac and β-cytoplasmic actin messenger RNAs to different cytoplasmic compartments. J. Cell Biol. 123, 165–172 (1993)
Zhang, H. L. et al. Neurotrophin-induced transport of a β-actin mRNP complex increases β-actin levels and stimulates growth cone motility. Neuron 31, 261–275 (2001)
Shestakova, E. A., Wyckoff, J., Jones, J., Singer, R. H. & Condeelis, J. Correlation of β-actin messenger RNA localization with metastatic potential in rat adenocarcinoma cell lines. Cancer Res. 59, 1202–1205 (1999)
Shestakova, E. A., Singer, R. H. & Condeelis, J. The physiological significance of β-actin mRNA localization in determining cell polarity and directional motility. Proc. Natl Acad. Sci. USA 98, 7045–7050 (2001)
Ross, A. F., Oleynikov, Y., Kislauskis, E. H., Taneja, K. L. & Singer, R. H. Characterization of a β-actin mRNA zipcode-binding protein. Mol. Cell. Biol. 17, 2158–2165 (1997)
Farina, K. L., Huttelmaier, S., Musunuru, K., Darnell, R. & Singer, R. H. Two ZBP1 KH domains facilitate β-actin mRNA localization, granule formation, and cytoskeletal attachment. J. Cell Biol. 160, 77–87 (2003)
Eom, T., Antar, L. N., Singer, R. H. & Bassell, G. J. Localization of a β-actin messenger ribonucleoprotein complex with zipcode-binding protein modulates the density of dendritic filopodia and filopodial synapses. J. Neurosci. 23, 10433–10444 (2003)
Femino, A. M., Fay, F. S., Fogarty, K. & Singer, R. H. Visualization of single RNA transcripts in situ. Science 280, 585–590 (1998)
Oleynikov, Y. & Singer, R. H. Real-time visualization of ZBP1 association with β-actin mRNA during transcription and localization. Curr. Biol. 13, 199–207 (2003)
Ostareck, D. H. et al. Lipoxygenase mRNA silencing in erythroid differentiation: The 3′UTR regulatory complex controls 60S ribosomal subunit joining. Cell 104, 281–290 (2001)
Schaller, M. D. Paxillin: a focal adhesion-associated adaptor protein. Oncogene 20, 6459–6472 (2001)
Ostareck-Lederer, A. et al. c-Src-mediated phosphorylation of hnRNP K drives translational activation of specifically silenced mRNAs. Mol. Cell. Biol. 22, 4535–4543 (2002)
Lukong, K. E. & Richard, S. Sam68, the KH domain-containing superSTAR. Biochim. Biophys. Acta 1653, 73–86 (2003)
Mayer, B. J. SH3 domains: complexity in moderation. J. Cell Sci. 114, 1253–1263 (2001)
Tiruchinapalli, D. M. et al. Activity-dependent trafficking and dynamic localization of zipcode binding protein 1 and β-actin mRNA in dendrites and spines of hippocampal neurons. J. Neurosci. 23, 3251–3261 (2003)
Mingle, L. A. et al. Localization of all seven messenger RNAs for the actin-polymerization nucleator Arp2/3 complex in the protrusions of fibroblasts. J. Cell Sci. 118, 2425–2433 (2005)
Medalia, O. et al. Macromolecular architecture in eukaryotic cells visualized by cryoelectron tomography. Science 298, 1209–1213 (2002)
Huynh, J. R., Munro, T. P., Smith-Litiere, K., Lepesant, J. A. & Johnston, D. S. The Drosophila hnRNPA/B homolog, Hrp48, is specifically required for a distinct step in osk mRNA localization. Dev. Cell 6, 625–635 (2004)
Farina, K. L. & Singer, R. H. The nuclear connection in RNA transport and localization. Trends Cell Biol. 12, 466–472 (2002)
Yano, T., de Quinto, S. L., Matsui, Y., Shevchenko, A. & Ephrussi, A. Hrp48, a Drosophila hnRNPA/B homolog, binds and regulates translation of oskar mRNA. Dev. Cell 6, 637–648 (2004)
Hoek, K. S., Kidd, G. J., Carson, J. H. & Smith, R. hnRNP A2 selectively binds the cytoplasmic transport sequence of myelin basic protein mRNA. Biochemistry 37, 7021–7029 (1998)
Kress, T. L., Yoon, Y. J. & Mowry, K. L. Nuclear RNP complex assembly initiates cytoplasmic RNA localization. J. Cell Biol. 165, 203–211 (2004)
Bromann, P. A., Korkaya, H. & Courtneidge, S. A. The interplay between Src family kinases and receptor tyrosine kinases. Oncogene 23, 7957–7968 (2004)
Playford, M. P. & Schaller, M. D. The interplay between Src and integrins in normal and tumour biology. Oncogene 23, 7928–7946 (2004)
Acknowledgements
We thank A. and D. Ostareck-Lederer for providing Src plasmids and for helpful discussions. We also thank S. Shenoy for help with microscopy and image processing. This research was supported by NIH grants to R.H.S., G.J.B. and J.C., a DOE grant to R.H.S., and by a BMBF (Bundesministerium für Bildung und Forschung) grant to S.H. Author Contributions All of the work in the manuscript was done by S.H., with help from D.Z., X.M. and M. Lederer in the Singer laboratory, and most recently the Hüttelmaier laboratory, except for the work with primary neurons (Fig. 3i–k; J.D. and G.J.B.) and the FRET studies (Fig. 3a–e and Supplementary Fig. S7a–f; M. Lorenz and J.C.).
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Competing interests
Reprints and permissions information is available at npg.nature.com/reprintsandpermissions. The authors declare no competing financial interests.
Supplementary information
Supplementary Notes
This file contains Supplementary Methods, Supplementary Figures 1–9 and accompanying legends. (PDF 1718 kb)
Rights and permissions
About this article
Cite this article
Hüttelmaier, S., Zenklusen, D., Lederer, M. et al. Spatial regulation of β-actin translation by Src-dependent phosphorylation of ZBP1. Nature 438, 512–515 (2005). https://doi.org/10.1038/nature04115
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1038/nature04115
This article is cited by
-
The double homeodomain protein DUX4c is associated with regenerating muscle fibers and RNA-binding proteins
Skeletal Muscle (2023)
-
Oncofetal protein IGF2BPs in human cancer: functions, mechanisms and therapeutic potential
Biomarker Research (2023)
-
A novel hypoxic long noncoding RNA KB-1980E6.3 maintains breast cancer stem cell stemness via interacting with IGF2BP1 to facilitate c-Myc mRNA stability
Oncogene (2021)
-
Intracellular mRNA transport and localized translation
Nature Reviews Molecular Cell Biology (2021)
-
BAD regulates mammary gland morphogenesis by 4E-BP1-mediated control of localized translation in mouse and human models
Nature Communications (2021)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.