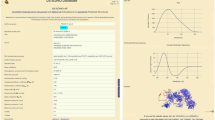
Abstract
The most controversial area in protein folding concerns its earliest stages. Questions such as whether there are genuine folding intermediates, and whether the events at the earliest stages are just rearrangements of the denatured state1 or progress from populated transition states2, remain unresolved. The problem is that there is a lack of experimental high-resolution structural information about early folding intermediates and denatured states under conditions that favour folding because competent states spontaneously fold rapidly. Here we have solved directly the solution structure of a true denatured state by nuclear magnetic resonance under conditions that would normally favour folding, and directly studied its equilibrium and kinetic behaviour. We engineered a mutant of Drosophila melanogaster Engrailed homeodomain that folds and unfolds reversibly just by changing ionic strength. At high ionic strength, the mutant L16A is an ultra-fast folding native protein, just like the wild-type protein; however, at physiological ionic strength it is denatured. The denatured state is a well-ordered folding intermediate, poised to fold by docking helices and breaking some non-native interactions. It unfolds relatively progressively with increasingly denaturing conditions, and so superficially resembles a denatured state with properties that vary with conditions. Such ill-defined unfolding is a common feature of early folding intermediate states and accounts for why there are so many controversies about intermediates versus compact denatured states in protein folding.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Krantz, B. A., Mayne, L., Rumbley, J., Englander, S. W. & Sosnick, T. R. Fast and slow intermediate accumulation and the initial barrier mechanism in protein folding. J. Mol. Biol. 324, 359–371 (2002)
Yang, W. Y. & Gruebele, M. Folding at the speed limit. Nature 423, 193–197 (2003)
Clarke, N. D., Kissinger, C. R., Desjarlais, J., Gilliland, G. L. & Pabo, C. O. Structural studies of the engrailed homeodomain. Protein Sci. 3, 1779–1787 (1994)
Mayor, U. et al. The complete folding pathway of a protein from nanoseconds to microseconds. Nature 421, 863–867 (2003)
Mayor, U., Grossmann, J. G., Foster, N. W., Freund, S. M. & Fersht, A. R. The denatured state of Engrailed Homeodomain under denaturing and native conditions. J. Mol. Biol. 333, 977–991 (2003)
Dill, K. A. & Shortle, D. Denatured states of proteins. Annu. Rev. Biochem. 60, 795–825 (1991)
Oliveberg, M. & Fersht, A. R. Thermodynamics of transient conformations in the folding pathway of barnase: Reorganization of the folding intermediate at low pH. Biochemistry 35, 2738–2749 (1996)
Korzhnev, D. M. et al. Low-populated folding intermediates of Fyn SH3 characterized by relaxation dispersion NMR. Nature 430, 586–590 (2004)
Neri, D., Billeter, M., Wider, G. & Wuthrich, K. NMR determination of residual structure in a urea-denatured protein, the 434-repressor. Science 257, 1559–1563 (1992)
Mok, Y. K., Kay, C. M., Kay, L. E. & Forman-Kay, J. NOE data demonstrating a compact unfolded state for an SH3 domain under non-denaturing conditions. J. Mol. Biol. 289, 619–638 (1999)
Gillespie, J. R. & Shortle, D. Characterization of long-range structure in the denatured state of staphylococcal nuclease. I. Paramagnetic relaxation enhancement by nitroxide spin labels. J. Mol. Biol. 268, 158–159 (1997)
Gillespie, J. R. & Shortle, D. Characterization of long-range structure in the denatured state of staphylococcal nuclease. II. Distance restraints from paramagnetic relaxation and calculation of an ensemble of structures. J. Mol. Biol. 268, 170–184 (1997)
Mohana-Borges, R., Goto, N. K., Kroon, G. J., Dyson, H. J. & Wright, P. E. Structural characterization of unfolded states of apomyoglobin using residual dipolar couplings. J. Mol. Biol. 340, 1131–1142 (2004)
Dyson, H. J. & Wright, P. E. Nuclear magnetic resonance methods for elucidation of structure and dynamics in disordered states. Methods Enzymol. 339, 258–270 (2001)
Wuthrich, K. NMR of Proteins and Nucleic Acids (John Wiley & Sons, New York, 1986)
Farrow, N. A., Zhang, O., Szabo, A., Torchia, D. A. & Kay, L. E. Spectral density function mapping using 15N relaxation data exclusively. J. Biomol. NMR 6, 153–162 (1995)
Farrow, N. A., Zhang, O., Forman-Kay, J. D. & Kay, L. E. Comparison of the backbone dynamics of a folded and an unfolded SH3 domain existing in equilibrium in aqueous buffer. Biochemistry 34, 868–878 (1995)
Ishima, R. & Nagayama, K. Protein backbone dynamics revealed by quasi spectral density function analysis of amide N-15 nuclei. Biochemistry 34, 3162–3171 (1995)
Koharudin, L. M. I., Bonvin, A., Kaptein, R. & Boelens, R. Use of very long-distance NOEs in a fully deuterated protein: an approach for rapid protein fold determination. J. Mag. Res. 163, 228–235 (2003)
Brunger, A. T. et al. Crystallography & NMR system: A new software suite for molecular structure determination. Acta Crystallogr. D 54, 905–921 (1998)
Bonvin, A. M. J. J. & Brunger, A. T. Conformational variability of solution nucelar magnetic resonance structures. J. Mol. Biol. 250, 80–93 (1995)
Bonvin, A. M. J. J. & Brunger, A. T. Do NOE distances contain enough information to assess the relative populations of multi-conformer structures? J. Biomol. NMR 7, 72–76 (1996)
Fushman, D., Tjandra, N. & Cowburn, D. Direct measurement of N-15 chemical shift anisotropy in solution. J. Am. Chem. Soc. 120, 10947–10952 (1998)
Rios, M. A. D. & Plaxco, K. W. Apparent Debye-Huckel electrostatic effects in the folding of a simple, single domain protein. Biochemistry 44, 1243–1250 (2005)
Acknowledgements
T.L.R. was supported by a Trinity College Internal Graduate Studentship, and J.S.M. by a Gates Cambridge Scholarship. We would like to thank T. J. Rutherford and L. Kowalik for help with NMR, and J. M. Perez Canadillas for help with protein expression.Author Contributions T.L.R. was responsible for the major experimental work and planning of this study, with further contributions from J.S.M., U.M. and S.M.V.F. A.R.F. wrote the paper with T.L.R. and performed data analysis.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
Reprints and permissions information is available at npg.nature.com/reprintsandpermissions. The authors declare no competing financial interests.
Supplementary information
Supplementary Figure S1
HN-N RDCs for L16A at 298 K (calculated as D – J coupling). A: 9% acrylamide. B: 12.5% Pf1 phage at pH 7.0. (JPG 21 kb)
Supplementary Notes
This file contains Supplementary Methods and Supplementary Tables S1 and S2. (DOC 45 kb)
Rights and permissions
About this article
Cite this article
Religa, T., Markson, J., Mayor, U. et al. Solution structure of a protein denatured state and folding intermediate. Nature 437, 1053–1056 (2005). https://doi.org/10.1038/nature04054
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1038/nature04054
This article is cited by
-
Modeling protein folding in vivo
Biology Direct (2018)
-
Ion–ion interactions in the denatured state contribute to the stabilization of CutA1 proteins
Scientific Reports (2018)
-
Antimicrobial peptide capsids of de novo design
Nature Communications (2017)
-
The energy cost of polypeptide knot formation and its folding consequences
Nature Communications (2017)
-
Resonance assignment of disordered protein with repetitive and overlapping sequence using combinatorial approach reveals initial structural propensities and local restrictions in the denatured state
Journal of Biomolecular NMR (2016)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.