Abstract
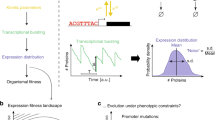
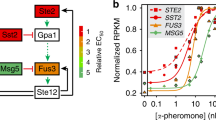
Here we studied the quantitative behaviour and cell-to-cell variability of a prototypical eukaryotic cell-fate decision system, the mating pheromone response pathway in yeast. We dissected and measured sources of variation in system output, analysing thousands of individual, genetically identical cells. Only a small proportion of total cell-to-cell variation is caused by random fluctuations in gene transcription and translation during the response (‘expression noise’). Instead, variation is dominated by differences in the capacity of individual cells to transmit signals through the pathway (‘pathway capacity’) and to express proteins from genes (‘expression capacity’). Cells with high expression capacity express proteins at a higher rate and increase in volume more rapidly. Our results identify two mechanisms that regulate cell-to-cell variation in pathway capacity. First, the MAP kinase Fus3 suppresses variation at high pheromone levels, while the MAP kinase Kss1 enhances variation at low pheromone levels. Second, pathway capacity and expression capacity are negatively correlated, suggesting a compensatory mechanism that allows cells to respond more precisely to pheromone in the presence of a large variation in expression capacity.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
References
Sternberg, P. W. & Horvitz, H. R. Pattern formation during vulval development in C. elegans. Cell 44, 761–772 (1986)
Priess, J. R. & Thomson, J. N. Cellular interactions in early C. elegans embryos. Cell 48, 241–250 (1987)
Kimble, J. & Hirsh, D. The postembryonic cell lineages of the hermaphrodite and male gonads in Caenorhabditis elegans. Dev. Biol. 70, 396–417 (1979)
Kimble, J. Alterations in cell lineage following laser ablation of cells in the somatic gonad of Caenorhabditis elegans. Dev. Biol. 87, 286–300 (1981)
Karp, X. & Greenwald, I. Post-transcriptional regulation of the E/Daughterless ortholog HLH-2, negative feedback, and birth order bias during the AC/VU decision in C. elegans. Genes Dev. 17, 3100–3111 (2003)
Doe, C. Q. & Goodman, C. S. Early events in insect neurogenesis. II. The role of cell interactions and cell lineage in the determination of neuronal precursor cells. Dev. Biol. 111, 206–219 (1985)
Jan, Y. N. & Jan, L. Y. Maggot's hair and bug's eye: role of cell interactions and intrinsic factors in cell fate specification. Neuron 14, 1–5 (1995)
Hoang, T. The origin of hematopoietic cell type diversity. Oncogene 23, 7188–7198 (2004)
Delbrück, M. The burst size distribution in the growth of bacterial viruses (bacteriophages). J. Bacteriol. 50, 131–135 (1945)
Elowitz, M. B. & Leibler, S. A synthetic oscillatory network of transcriptional regulators. Nature 403, 335–338 (2000)
Elowitz, M. B., Levine, A. J., Siggia, E. D. & Swain, P. S. Stochastic gene expression in a single cell. Science 297, 1183–1186 (2002)
Rosenfeld, N., Young, J. W., Alon, U., Swain, P. S. & Elowitz, M. B. Gene regulation at the single-cell level. Science 307, 1962–1965 (2005)
Pedraza, J. M. & van Oudenaarden, A. Noise propagation in gene networks. Science 307, 1965–1969 (2005)
Raser, J. M. & O'Shea, E. K. Control of stochasticity in eukaryotic gene expression. Science 304, 1811–1814 (2004)
Ozbudak, E. M., Thattai, M., Kurtser, I., Grossman, A. D. & van Oudenaarden, A. Regulation of noise in the expression of a single gene. Nature Genet. 31, 69–73 (2002)
Blake, W. J., Kærn, M., Cantor, C. R. & Collins, J. J. Noise in eukaryotic gene expression. Nature 422, 633–637 (2003)
Dohlman, H. G. & Thorner, J. W. Regulation of G protein-initiated signal transduction in yeast: paradigms and principles. Annu. Rev. Biochem. 70, 703–754 (2001)
Roberts, C. J. et al. Signaling and circuitry of multiple MAPK pathways revealed by a matrix of global gene expression profiles. Science 287, 873–880 (2000)
Heiman, M. G. & Walter, P. Prm1p, a pheromone-regulated multispanning membrane protein, facilitates plasma membrane fusion during yeast mating. J. Cell Biol. 151, 719–730 (2000)
Jenness, D. D., Burkholder, A. C. & Hartwell, L. H. Binding of alpha-factor pheromone to Saccharomyces cerevisiae a cells: dissociation constant and number of binding sites. Mol. Cell. Biol. 6, 318–320 (1986)
Bishop, A. C. et al. A chemical switch for inhibitor-sensitive alleles of any protein kinase. Nature 407, 395–401 (2000)
Oehlen, L. J. & Cross, F. R. G1 cyclins CLN1 and CLN2 repress the mating factor response pathway at Start in the yeast cell cycle. Genes Dev. 8, 1058–1070 (1994)
Oehlen, L. J. & Cross, F. R. Potential regulation of Ste20 function by the Cln1-Cdc28 and Cln2-Cdc28 cyclin-dependent protein kinases. J. Biol. Chem. 273, 25089–25097 (1998)
Wassmann, K. & Ammerer, G. Overexpression of the G1-cyclin gene CLN2 represses the mating pathway in Saccharomyces cerevisiae at the level of the MEKK Ste11. J. Biol. Chem. 272, 13180–13188 (1997)
Breitkreutz, A., Boucher, L. & Tyers, M. MAPK specificity in the yeast pheromone response independent of transcriptional activation. Curr. Biol. 11, 1266–1271 (2001)
Chou, S., Huang, L. & Liu, H. Fus3-regulated Tec1 degradation through SCFCdc4 determines MAPK signalling specificity during mating in yeast. Cell 119, 981–990 (2004)
Bao, M. Z., Schwartz, M. A., Cantin, G. T., Yates, J. R. & Madhani, H. D. III Pheromone-dependent destruction of the Tec1 transcription factor is required for MAP kinase signalling specificity in yeast. Cell 119, 991–1000 (2004)
Bardwell, L., Cook, J. G., Zhu-Shimoni, J. X., Voora, D. & Thorner, J. Differential regulation of transcription: repression by unactivated mitogen-activated protein kinase Kss1 requires the Dig1 and Dig2 proteins. Proc. Natl Acad. Sci. USA 95, 15400–15405 (1998)
Sabbagh, W. Jr, Flatauer, L. J., Bardwell, A. J. & Bardwell, L. Specificity of MAP kinase signalling in yeast differentiation involves transient versus sustained MAPK activation. Mol. Cell 8, 683–691 (2001)
Paulsson, J. Summing up the noise in gene networks. Nature 427, 415–418 (2004)
Johnston, G. C., Pringle, J. R. & Hartwell, L. H. Coordination of growth with cell division in the yeast Saccharomyces cerevisiae. Exp. Cell Res. 105, 79–98 (1977)
Doi, K. et al. MSG5, a novel protein phosphatase promotes adaptation to pheromone response in S. cerevisiae. EMBO J. 13, 61–70 (1994)
Andersson, J., Simpson, D. M., Qi, M., Wang, Y. & Elion, E. A. Differential input by Ste5 scaffold and Msg5 phosphatase route a MAPK cascade to multiple outcomes. EMBO J. 23, 2564–2576 (2004)
Madhani, H. D. & Fink, G. R. The riddle of MAP kinase signalling specificity. Trends Genet. 14, 151–155 (1998)
Lee, B. N. & Elion, E. A. The MAPKKK Ste11 regulates vegetative growth through a kinase cascade of shared signalling components. Proc. Natl Acad. Sci. USA 96, 12679–12684 (1999)
Ausubel, F. M. et al. (eds) Current Protocols in Molecular Biology (Wiley, New York, 2004)
Guthrie, C. & Fink, G. R. (eds) Methods in Enzymology. Guide to Yeast Genetics and Molecular Biology (Academic, San Diego, 1991)
Colman-Lerner, A., Chin, T. E. & Brent, R. Yeast Cbk1 and Mob2 activate daughter-specific genetic programs to induce asymmetric cell fates. Cell 107, 739–750 (2001)
Acknowledgements
We are grateful to D. Pincus for his help with the MAPK phosphorylation measurements and to L. Lok, K. Benjamin, I. Burbulis and R. Yu for discussions and comments on the manuscript. Work was under the ‘Alpha Project’ at the Center for Genomic Experimentation and Computation, an NIH Center of Excellence in Genomic Science. The Alpha Project is supported by a grant from the National Human Genome Research Institute to R.B. Author Contributions A.C.-L. and A.G. conceived the framework, developed the experimental methods, performed most of the experiments and analysed the results. R.B. provided input regarding problem choice, experimentation and interpretation. A.C.-L., A.G. and R.B. wrote the paper and stand as guarantors of its findings. T.C. made most of the plasmid and yeast strains. E.S. made some plasmids and yeast strains, and some of the measurements in Fig. 2. C.G.P. made the observation that Fus3 regulates pathway variation and collaborated in interpreting its biological implications. O.R. and A.C.-L. made the observation that activated Fus3/Kss1 ratios are α-factor-dependent. D.E. suggested analysis of the stochastic fluctuations in the system and helped with describing the framework.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
Reprints and permissions information is available at npg.nature.com/reprintsandpermissions. The authors declare no competing financial interests.
Supplementary information
Supplementary Notes
This file contains Supplementary Methods, Supplementary Tables and Supplementary Figure Legends. A full list of the contents of this file is included on the first page. (PDF 2152 kb)
Supplementary Figures
This file contains Supplementary Figures S1-S6. (PDF 6254 kb)
Rights and permissions
About this article
Cite this article
Colman-Lerner, A., Gordon, A., Serra, E. et al. Regulated cell-to-cell variation in a cell-fate decision system. Nature 437, 699–706 (2005). https://doi.org/10.1038/nature03998
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature03998
This article is cited by
-
Design principles of improving the dose-response alignment in coupled GTPase switches
npj Systems Biology and Applications (2023)
-
A synthetic circuit for buffering gene dosage variation between individual mammalian cells
Nature Communications (2021)
-
Introns control stochastic allele expression bias
Nature Communications (2021)
-
Cell-to-cell variation in gene expression and the aging process
GeroScience (2021)
-
Single-particle imaging of stress-promoters induction reveals the interplay between MAPK signaling, chromatin and transcription factors
Nature Communications (2020)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.