Abstract
West Nile virus is a mosquito-borne flavivirus closely related to the human epidemic-causing dengue, yellow fever and Japanese encephalitis viruses1. In establishing infection these icosahedral viruses undergo endosomal membrane fusion catalysed by envelope glycoprotein rearrangement of the putative receptor-binding domain III (DIII) and exposure of the hydrophobic fusion loop2,3,4. Humoral immunity has an essential protective function early in the course of West Nile virus infection5,6. Here, we investigate the mechanism of neutralization by the E16 monoclonal antibody that specifically binds DIII. Structurally, the E16 antibody Fab fragment engages 16 residues positioned on four loops of DIII, a consensus neutralizing epitope sequence conserved in West Nile virus and distinct in other flaviviruses. The E16 epitope protrudes from the surface of mature virions in three distinct environments7, and docking studies predict Fab binding will leave five-fold clustered epitopes exposed. We also show that E16 inhibits infection primarily at a step after viral attachment, potentially by blocking envelope glycoprotein conformational changes. Collectively, our results suggest that a vaccine strategy targeting the dominant DIII epitope may elicit safe and effective immune responses against flaviviral diseases.
Similar content being viewed by others
Main
To understand better the mechanism of antibody neutralization of West Nile virus we generated a large panel of envelope glycoprotein-specific monoclonal antibodies8. Domain mapping by yeast surface display revealed that ten out of twelve potently neutralizing monoclonal antibodies selectively bind DIII. We also established that one of these monoclonal antibodies, E16, protects mice from lethal West Nile virus challenge even if administered therapeutically 5 days after infection. Here we have examined the structural basis for E16-mediated neutralization by determining the crystal structure of the Fab fragment in complex with West Nile virus DIII at 2.5 Å resolution (Supplementary Table S1). Our structure reveals that DIII adopts an immunoglobulin-like β-sandwich topology similar to that found in other flavivirus envelope glycoproteins, whereas the E16 Fab adopts a typical quaternary assembly (Fig. 1a). The binding interface has a high degree of shape complementarity (Sc = 0.763) (ref. 9) and occludes 1,550 Å2 of surface area, with VH (variable domain of heavy chain) accounting for 67% of the total antibody-combining site (Fig. 1b). E16 contacts DIII with 18 residues spread along all six of its CDR (complementarity determining region) loops in addition to three VH framework residues (Supplementary Table S2). The interaction between E16 and DIII is dominated by hydrogen bonds, with 16 direct hydrogen bonds and numerous water-mediated networks at the interface of the complex.
a, Ribbon diagram of the complex, with DIII depicted in dark blue, the antibody heavy chain in green and the light chain in cyan. b, Molecular surface representation of the E16–DIII interface highlighting E16 VH (green) and VL (cyan) contact residues (left), as well as DIII contact residues (blue) and the residues defined by yeast surface display (magenta) (right). c, Flow cytometry of yeast cells expressing wild-type or mutant versions of West Nile virus DIII. d, Detailed interactions of DIII residues Ser 306 and Lys 307 with E16, with interfacial waters (red) evident in the composite electron density omit map. e, Interactions of Thr 330 and Thr 332 at the E16 interface.
E16 engages four discontinuous segments of DIII including the amino-terminal region (residues 302–309) and three strand-connecting loops: BC (330–333), DE (365–368) and FG (389–391). E16 contacts a total of 16 DIII residues, which together form a single convex surface patch. Notably, yeast surface display epitope mapping10 of DIII identified four residues at the core of this binding site that are critical for E16 recognition (Fig. 1c). Non-conservative substitution at Ser 306, Lys 307, Thr 330 or Thr 332 disrupts E16 binding but not that of a non-neutralizing DIII-specific monoclonal antibody, E22. These four residues cluster at the centre of the E16–DIII interface (Fig. 1b). The decreased binding associated with mutation of the DIII residues Ser 306, Lys 307 and Thr 332 is most likely attributable to loss of hydrogen-bonding potential with E16, whereas Thr 330 appears to stabilize the DIII N-terminal strand conformation and provides numerous van der Waals contacts with the Fab (Fig. 1d, e). Collectively, our structural studies define the E16 epitope as a large surface patch on DIII created by four distinct secondary structure elements, and yeast mapping highlights the critical contributions of four central residues.
Comparison of available West Nile virus sequences reveals nearly complete conservation of the structurally defined E16 epitope (Fig. 2, and data not shown). Not surprisingly, E16 blocks infection of ten different lineage I and II West Nile virus strains8. Notably, nine other neutralizing monoclonal antibodies from our panel also lose the ability to recognize DIII after mutation of Ser 306, Lys 307, Thr 330 or Thr 332. Importantly, this epitope is also key in the humoral immune response of humans, as E16 Fabs effectively compete with West Nile virus convalescent antibodies for DIII binding8. Sequence analysis of other flaviviruses reveals a high degree of diversity in the four segments of the E16 epitope, with notable variation even between dengue serotypes (Fig. 2). Not surprisingly, E16 does not cross-neutralize dengue, Japanese or St Louis encephalitis viruses. Other groups have also identified flavivirus-specific neutralizing antibodies that localize to an analogous DIII-binding region11,12,13. Thus, the coincident mapping of our and other neutralizing monoclonal antibodies suggests that the structural epitope has a dominant role in flavivirus neutralization.
a, Sequence of the four segments of West Nile virus (WNV) DIII contacted by E16 aligned with the analogous residues of other flaviviruses. The DIII residues contacted by E16 are highlighted in blue and magenta, and deletions are indicated with a hash symbol. DEN, dengue virus; JEV, Japanese encephalitis virus; TBEV, tick-borne encephalitis virus; YFV, yellow fever virus. b, Structure of the West Nile virus dominant neutralizing epitope as defined by the E16–DIII complex.
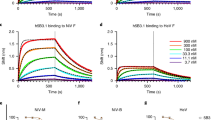
To gain additional insight into the structural basis of E16-mediated neutralization, we docked our Fab–DIII complex onto the structures of the pre-fusion dengue envelope glycoprotein dimer14 and post-fusion trimer3 (Fig. 3a, b). The E16 epitopes are unencumbered in either configuration, although intact E16 antibody is unlikely to bivalently recognize these isolated oligomers due to extensive distal splaying of the Fab tails (Fig. 3a, b). DIII undergoes an ∼70° rotation towards DII in the dimer to trimer transition2,3, and E16 ligation per se could serve to hinder this conformational change. Moreover, E16 binds part of the linker that connects DIII to DI, and the N terminus of our truncated West Nile virus DIII fragment adopts a unique conformation that enables Tyr 302 to make contact with the E16 VH domain (Fig. 3c). In envelope glycoprotein trimers, the flavivirus invariant Tyr 302 interacts with DI in a manner that would be disrupted if re-oriented as observed in the E16–DIII complex. Thus, E16 binding could stabilize the mature state or alternatively restrict transition to the post-fusion conformation.
a, E16 docked onto the DEN-2 envelope glycoprotein dimer (1OAN) through DIII (blue). Super-positioning indicates that binding probably occurs without DI (red) or DII (yellow) contacts. The distances between DIII contact sites (141 Å) and the Fab termini (239 Å) preclude bivalent antibody engagement. b, E16 docked onto the post-fusion DEN-2 envelope glycoprotein trimer (1OK8) indicates accessibility of the binding site. c, DIII from the E16 complex (blue), the pre-fusion DEN-2 (1OKE, light grey) and post-fusion DEN-2 (1OK8, dark grey) reveal the conserved structure of the domains. The interaction of E16 with the flavivirus conserved Tyr 302 in the N-terminal region of West Nile virus DIII is highlighted. d, The E16 structural epitope is mapped in magenta onto the cryo-electron microscopic reconstruction of the West Nile virus virion7 presented as 2.0-Å-radius Cα atoms. e, E16–DIII complexes docked around the icosahedral three-fold axis. f, DIII situated around the outer ring of the five-fold axis is permissive to E16 binding, but the inner ring appears to exclude E16 engagement. g, Saturation binding by E16 is predicted to entail the binding of 120 out of 180 West Nile virus epitopes with exclusion of binding to the inner-five-fold DIIIs (magenta). Panels e and f are magnified views of the boxed regions in g.
To understand better how E16 recognizes DIII in the context of the mature virus, we docked our structure onto the cryo-electron-microscopy-derived pseudo-atomic model of the intact West Nile virus virion7,16. With three envelope glycoproteins in the asymmetric unit, there are three potential Fab-binding environments (Fig. 3d). Two binding modes are clearly allowed: one that closely circles the three-fold axis and a second disposed symmetrically about the icosahedral dyad that is permuted as an outer five-fold ring (Fig. 3e, f). However, the DIII epitopes are too tightly clustered at the true five-fold axis to permit E16 engagement without steric overlap with adjacent DIII residues. Thus, we postulate that at saturation no more than 120 Fabs can bind the 180 envelope glycoproteins in the mature virion, with exclusion of Fab binding to DIII around the inner five-fold ring (Fig. 3g). Additional exclusions may occur for intact antibody, although we note that E16 Fab alone neutralizes West Nile virus. This incomplete saturation is analogous to that observed in the decoration of papillomavirus by a neutralizing antibody, which blocks infection through a post-attachment mechanism17.
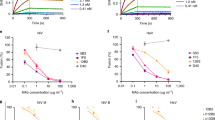
We next tested whether E16 could block cellular attachment of West Nile virus. Binding assays were performed with Vero cells, a cell line permissive for West Nile virus infection. After a 4 °C incubation with West Nile virus in the presence of control (anti-SARS ORF7a), non-neutralizing (E22), or neutralizing monoclonal antibodies that map within (E16 or E24) or outside (E53 or E60) of DIII, cell-associated viral RNA was measured by fluorogenic polymerase chain reaction with reverse transcription (RT–PCR)18. Importantly, the non-binding and non-neutralizing monoclonal antibodies do not inhibit virus binding. In contrast, E53 and E60 block virus attachment by 8–9-fold (P < 0.001) whereas E16 and E24, which recognize the same dominant DIII epitope, only inhibit binding by 3.5-fold (P = 0.003) (Fig. 4a). The observation that E53 and E60 block virus binding more efficiently than E16 was not expected, as E53 and E60 are tenfold less potent in plaque reduction neutralization assays8.
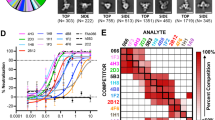
a, Two DI/DII-specific neutralizing monoclonal antibodies (E53 and E60) block cellular attachment significantly more than the DIII-specific neutralizing antibodies (E16 and E24) or controls (no antibody, non-neutralizing monoclonal antibody E22 or anti-SARS ORF7a). Fold-reductions are reported, with standard deviations, as the average of four to seven independent experiments performed in triplicate. b, Dose-dependent blockade of West Nile virus infection by E16 and E53 in pre- and post-adsorption assays. The data are one of three representative experiments performed in duplicate. c, The DIII-specific monoclonal antibodies effectively inhibit West Nile virus infection of macrophages, whereas DI/DII-specific E5 and E60 monoclonal antibodies enhance infection. The data are one of three representative experiments performed in duplicate, with the dotted line representing the limit of sensitivity of the assay. Error bars represent the standard deviation. d, Pre-incubation with unlabelled monoclonal antibodies followed by addition of APC conjugates reveals that both E16 and E60 are self-competitive but not cross-competitive for envelope glycoprotein binding.
Because E16 only partially blocks virus binding yet completely neutralizes infection, we tested whether E16 inhibits flavivirus infection by blocking a step after cellular attachment. Using a previously described assay19,20, E16 or E53 was incubated with West Nile virus before, or after, mixing with a monolayer of Vero cells and infection was measured. Pre-binding of West Nile virus with either E16 or E53 significantly protects against infection (Fig. 4b). In contrast, E16 but not E53 significantly inhibits infection when added after virus binding. Because E16-mediated protection is not appreciably affected by the time of addition, we surmise that it acts primarily after West Nile virus cellular attachment.
To define further the mechanism of West Nile virus neutralization, we evaluated whether E16 or other monoclonal antibodies enhance infection in macrophages. Antibody-dependent enhancement of infection occurs when antibody–virus complexes are preferentially internalized through Fcγ receptors on myeloid cells. Although the in vivo consequences remain uncertain, many monoclonal antibodies efficiently enhance flavivirus infection of Fcγ-receptor-bearing cells even when inhibitory in fibroblast neutralization assays21. We tested whether saturating concentrations of non-neutralizing (E5) or neutralizing (E16, E24 or E60) monoclonal antibodies enhance West Nile virus infection in macrophages. We found that whereas E5 and E60 augment infection 270- and 3,000-fold, respectively, E16 potently inhibits macrophage infection at the same concentration. Notably, when E16 is combined with E5 or E60, it completely blocks enhancement as judged by reduction of virus yield (Fig. 4c) or viral RNA (data not shown). E24, which maps to the E16 dominant epitope, also blocks E5- and E60-dependent enhancement. Finally, the blockade of enhancement is not due to epitope competition as E16 and E24 do not cross-compete E5 or E60 for West Nile virus envelope glycoprotein binding (Fig. 4d and data not shown). Collectively, these virological experiments strongly suggest that E16 blocks West Nile virus infection primarily after cellular attachment.
We have shown that E16 binds a dominant neutralizing epitope on West Nile virus envelope glycoprotein defined by four distinct secondary structure elements that create a large surface patch on DIII, a region associated with pH-dependent conformational changes. Modelling studies suggest that E16 is excluded from five-fold clustered DIII epitopes on mature virions, potentially leaving them free to serve in receptor binding. Consistent with this, E16 inhibits West Nile virus infection primarily at a step after virus attachment. Moreover, our data suggest a potential advantage for interfering with post-attachment events, an emerging theme in other viral inhibition studies22,23,24,25. Vaccines that selectively elicit antibodies specific to the critical DIII epitope may neutralize infection regardless of the mode of cellular entry, which may be particularly relevant given the increasing number of putative flavivirus entry receptors26,27,28,29.
Methods
Protein production and crystallization
West Nile virus DIII (residues 296–401) was expressed in bacteria with thrombin-cleavable histidine and BirA tags, and oxidatively re-folded from isolated inclusion bodies. Antibody–antigen complexes were formed by mixing papain-generated E16 Fab with thrombin-cleaved DIII, and purified by gel filtration chromatography. The E16–DIII complex crystallized at 15 mg ml-1 by hanging-drop vapour diffusion at 20 °C using 19% PEG 4000, 100 mM glycine and 100 mM HEPES (pH 8.5). Crystals were cryo-protected with precipitant solution containing 20% ethylene glycol, rapidly cooled in liquid nitrogen, and annealed by two 5-s blockages of the nitrogen stream.
Crystallographic structure determination
Data were collected at ALS beamline 4.2.2 (Lawrence Berkeley Laboratories) by the oscillation method at a wavelength of 1.55 Å at 100 K with a CCD detector. Data were processed, scaled and merged with d*TREK (http://www.rigakumsc.com/protein/dtrek.html). The crystals belong to space group P212121 with unit cell dimensions of a = 52.4 Å, b = 83.3 Å and c = 110.6 Å, with one E16 Fab–DIII complex per asymmetric unit. Crystallographic phases were obtained by molecular replacement (http://www.ccp4.ac.uk/) using the coordinates of an IgG1 Fab′ (Protein Data Bank (PDB) 2IGF) and the NMR structure of West Nile virus DIII (PDB 1S6N), which provided a correlation coefficient of 0.563 after rigid body refinement. An atomic model was iteratively built in O30 and refined in CNS (http://cns.csb.yale.edu/v1.1/), and contains 532 amino acids (residues 300–400 of DIII, 1–212 of the E16 light chain and 1–228 of the E16 heavy chain) and 256 water molecules. The final 2.5 Å resolution model was refined to an Rcryst = 20.5% and Rfree = 28.1% for all F > 0, with excellent geometry and Ramachandran angles (favoured, additional, generous and disallowed values of 87.5%, 11.9%, 0.4% and 0.2%, respectively).
Epitope mapping by yeast surface display
To define DIII residues critical for E16 recognition we expressed an error-prone PCR-derived library of West Nile virus envelope glycoprotein (residues 296–415) on the surface of yeast as Aga2 fusion proteins8,10. Yeast were screened for selective loss of E16 binding relative to other DIII-specific monoclonal antibodies by multiple rounds of fluorescence-activated cell sorting. Isolated clones were recovered, sequenced and evaluated for binding to E16 and the non-neutralizing, West Nile virus DIII-specific monoclonal antibody E22.
West Nile virus binding to Vero cells
Individual purified monoclonal antibodies (50 µg ml-1 of anti-SARS ORF7a, E16, E22, E24, E53 or E60) or medium alone were incubated with 103 plaque-forming units (p.f.u.) of West Nile virus for 1 h at 4 °C. These virus–antibody mixtures were then added to Vero cells in 12-well plates for 1 h on ice. Unbound virus was removed after six washes with PBS at 4 °C. Cells were lysed with guanidinium isothiocyanate, RNA was purified, and viral RNA was quantified by fluorogenic RT–PCR18.
Pre- and post-adsorption antibody inhibition assay
Increasing concentrations of E16 or E53 were added before or after West Nile virus (102 p.f.u.) binding (1 h on ice) to Vero cells. In the post-adsorption assay, after washing away unbound virus, monoclonal antibody was allowed to bind for an additional hour. All cells were washed and an agarose overlay was added. Three days later, plaques were scored after fixation and staining with crystal violet.
Antibody-dependent enhancement assay
West Nile virus (5 × 102 p.f.u.) was pre-incubated with media, individual monoclonal antibodies (50 µg ml-1 of E5, E16, E24 or E60) or combinations of monoclonal antibodies (E16 plus E5, E60 plus E5, E16 plus E60, E24 plus E5 or E24 plus E60) and then added to a monolayer (105) of J774.2 murine macrophages. After 6 h, cells were washed extensively with PBS to remove unbound virus and monoclonal antibody. After an additional 24 h, supernatants were harvested for a viral plaque assay on Vero cells.
Competitive binding of E16 and E60
Yeast expressing the empty vector pYD1 or the West Nile virus envelope glycoprotein ectodomain (residues 1–415) were incubated with 2.5 µg unlabelled E16 or E60 antibody for 1 h on ice. Unbound antibody was removed after three PBS washes containing 1 mg ml-1 BSA. E16 and E60 were conjugated using an Alexa Flour 647 monoclonal antibody labelling kit (Molecular Probes). Conjugated E16 or E60 (25 µg ml-1) was then added to the cells for 30 min at 4 °C. Yeast were washed three times with PBS, fixed with 1% paraformaldehyde, and analysed using flow cytometry.
References
Granwehr, B. P. et al. West Nile virus: where are we now? Lancet Infect. Dis. 4, 547–556 (2004)
Bressanelli, S. et al. Structure of a flavivirus envelope glycoprotein in its low-pH-induced membrane fusion conformation. EMBO J. 23, 728–738 (2004)
Modis, Y., Ogata, S., Clements, D. & Harrison, S. C. Structure of the dengue virus envelope protein after membrane fusion. Nature 427, 313–319 (2004)
Mukhopadhyay, S., Kuhn, R. J. & Rossmann, M. G. A structural perspective of the flavivirus life cycle. Nature Rev. Microbiol. 3, 13–22 (2005)
Diamond, M. S., Shrestha, B., Mehlhop, E., Sitati, E. & Engle, M. Innate and adaptive immune responses determine protection against disseminated infection by West Nile encephalitis virus. Viral Immunol. 16, 259–278 (2003)
Wang, T. & Fikrig, E. Immunity to West Nile virus. Curr. Opin. Immunol. 16, 519–523 (2004)
Mukhopadhyay, S., Kim, B. S., Chipman, P. R., Rossmann, M. G. & Kuhn, R. J. Structure of West Nile virus. Science 302, 248 (2003)
Oliphant, T. et al. Development of a humanized monoclonal antibody with therapeutic potential against West Nile virus. Nature Med. 11, 522–530 (2005)
Sundberg, E. J. & Mariuzza, R. A. Molecular recognition in antibody-antigen complexes. Adv. Protein Chem. 61, 119–160 (2002)
Chao, G., Cochran, J. R. & Wittrup, K. D. Fine epitope mapping of anti-epidermal growth factor receptor antibodies through random mutagenesis and yeast surface display. J. Mol. Biol. 342, 539–550 (2004)
Wu, K. P. et al. Structural basis of a flavivirus recognized by its neutralizing antibody: Solution structure of the domain III of the Japanese Encephalitis virus envelope protein. J. Biol. Chem. 278, 46007–46013 (2003)
Volk, D. E. et al. Solution structure and antibody binding studies of the envelope protein domain III from the New York strain of West Nile virus. J. Biol. Chem. 279, 38755–38761 (2004)
Hiramatsu, K., Tadano, M., Men, R. & Lai, C. J. Mutational analysis of a neutralization epitope on the dengue type 2 virus (DEN2) envelope protein: monoclonal antibody resistant DEN2/DEN4 chimeras exhibit reduced mouse neurovirulence. Virology 224, 437–445 (1996)
Modis, Y., Ogata, S., Clements, D. & Harrison, S. C. A ligand-binding pocket in the dengue virus envelope glycoprotein. Proc. Natl Acad. Sci. USA 100, 6986–6991 (2003)
Saphire, E. O. et al. Crystal structure of a neutralizing human IgG against HIV-1: a template for vaccine design. Science 293, 1155–1159 (2001)
Kuhn, R. J. et al. Structure of dengue virus: implications for flavivirus organization, maturation, and fusion. Cell 108, 717–725 (2002)
Booy, F. P., Roden, R. B., Greenstone, H. L., Schiller, J. T. & Trus, B. L. Two antibodies that neutralize papillomavirus by different mechanisms show distinct binding patterns at 13 Å resolution. J. Mol. Biol. 281, 95–106 (1998)
Diamond, M. S., Shrestha, B., Marri, A., Mahan, D. & Engle, M. B cells and antibody play critical roles in the immediate defense of disseminated infection by West Nile encephalitis virus. J. Virol. 77, 2578–2586 (2003)
Crill, W. D. & Roehrig, J. T. Monoclonal antibodies that bind to domain III of dengue virus E glycoprotein are the most efficient blockers of virus adsorption to Vero cells. J. Virol. 75, 7769–7773 (2001)
Hung, S. L. et al. Analysis of the steps involved in Dengue virus entry into host cells. Virology 257, 156–167 (1999)
Halstead, S. B. & O'Rourke, E. J. Antibody-enhanced dengue virus infection in primate leukocytes. Nature 265, 739–741 (1977)
Gollins, S. W. & Porterfield, J. S. A new mechanism for the neutralization of enveloped viruses by antiviral antibody. Nature 321, 244–246 (1986)
Zolla-Pazner, S. Identifying epitopes of HIV-1 that induce protective antibodies. Nature Rev. Immunol. 4, 199–210 (2004)
Skehel, J. J. & Wiley, D. C. Receptor binding and membrane fusion in virus entry: the influenza hemagglutinin. Annu. Rev. Biochem. 69, 531–569 (2000)
Earp, L. J., Delos, S. E., Park, H. E. & White, J. M. The many mechanisms of viral membrane fusion proteins. Curr. Top. Microbiol. Immunol. 285, 25–66 (2005)
Chen, Y. et al. Dengue virus infectivity depends on envelope protein binding to target cell heparan sulfate. Nature Med. 3, 866–871 (1997)
Tassaneetrithep, B. et al. DC-SIGN (CD209) mediates dengue virus infection of human dendritic cells. J. Exp. Med. 197, 823–829 (2003)
Halstead, S. B. Neutralization and antibody-dependent enhancement of dengue viruses. Adv. Virus Res. 60, 421–467 (2003)
Tio, P. H., Jong, W. W. & Cardosa, M. J. Two dimensional VOPBA reveals laminin receptor (LAMR1) interaction with dengue virus serotypes 1, 2 and 3. Virol. J. 2, 25 (2005)
Jones, T. A., Zou, J. Y., Cowan, S. W. & Kjeldgaard, M. Improved methods for binding protein models in electron density maps and the location of errors in these models. Acta Crystallogr. A 47, 110–119 (1991)
Acknowledgements
We thank J. Alexander for yeast surface display expertise, O. Naidenko for expression plasmids, R. Kinney for the West Nile virus infectious clone, and R. Kuhn and M. Rossmann for coordinates of the West Nile virus mature virion. The authors also thank H. Virgin, T. Hansen and E. Unanue for critical comments on the manuscript. The work was supported by grants from the NIH and the Pediatric Dengue Vaccine Initiative (M.S.D. and D.H.F.). An NIH Medical Scientist Training Program Grant supported G.E.N. Author Contributions G.E.N. carried out the protein biochemistry, crystallography and data analysis under the supervision of D.H.F. Epitope mapping and virology assays were performed by T.O. and M.S.D. E16 was sequenced by S.J. and Fabs were generated by S.B. The manuscript was written by G.E.N., M.S.D. and D.H.F.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare competing financial interests: details accompany the paper on www.nature.com/nature.G.E.N., T.O., M.S.D. and D.H.F. have filed provisional patents based on this and other studies. S.J. and S.B. are employed by and M.S.D. consults for MacroGenics.
Supplementary information
Supplementary Table S1
This table describes the crystallographic data collection and refinement statistics for the E16 Fab/WNV DIII complex. (RTF 24 kb)
Supplementary Table S2
This table describes van der Waals contacts and putative direct and water mediated hydrogen bonds at the E16 Fab/WNV DIII interface. (RTF 44 kb)
Rights and permissions
About this article
Cite this article
Nybakken, G., Oliphant, T., Johnson, S. et al. Structural basis of West Nile virus neutralization by a therapeutic antibody. Nature 437, 764–769 (2005). https://doi.org/10.1038/nature03956
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1038/nature03956
This article is cited by
-
Neutralizing antibodies targeting a novel epitope on envelope protein exhibited broad protection against flavivirus without risk of disease enhancement
Journal of Biomedical Science (2023)
-
Potential neutralizing antibodies discovered for novel corona virus using machine learning
Scientific Reports (2021)
-
The distribution of important sero-complexes of flaviviruses in Malaysia
Tropical Animal Health and Production (2019)
-
Characterization of a potent and highly unusual minimally enhancing antibody directed against dengue virus
Nature Immunology (2018)
-
A protective human monoclonal antibody targeting the West Nile virus E protein preferentially recognizes mature virions
Nature Microbiology (2018)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.