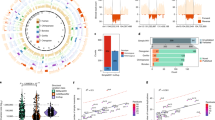
Abstract
We present a global comparison of differences in content of segmental duplication between human and chimpanzee, and determine that 33% of human duplications (> 94% sequence identity) are not duplicated in chimpanzee, including some human disease-causing duplications. Combining experimental and computational approaches, we estimate a genomic duplication rate of 4–5 megabases per million years since divergence. These changes have resulted in gene expression differences between the species. In terms of numbers of base pairs affected, we determine that de novo duplication has contributed most significantly to differences between the species, followed by deletion of ancestral duplications. Post-speciation gene conversion accounts for less than 10% of recent segmental duplication. Chimpanzee-specific hyperexpansion (> 100 copies) of particular segments of DNA have resulted in marked quantitative differences and alterations in the genome landscape between chimpanzee and human. Almost all of the most extreme differences relate to changes in chromosome structure, including the emergence of African great ape subterminal heterochromatin. Nevertheless, base per base, large segmental duplication events have had a greater impact (2.7%) in altering the genomic landscape of these two species than single-base-pair substitution (1.2%).
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Bailey, J. A., Baertsch, R., Kent, W. J., Haussler, D. & Eichler, E. E. Hotspots of mammalian chromosomal evolution. Genome Biol. 5, R23 (2004)
She, X. et al. The structure and evolution of centromeric transition regions within the human genome. Nature 430, 857–864 (2004)
Armengol, L., Pujana, M. A., Cheung, J., Scherer, S. W. & Estivill, X. Enrichment of segmental duplications in regions of breaks of synteny between the human and mouse genomes suggest their involvement in evolutionary rearrangements. Hum. Mol. Genet. 12, 2201–2208 (2003)
Trask, B. et al. Members of the olfactory receptor gene family are contained in large blocks of DNA duplicated polymorphically near the ends of human chromosomes. Hum. Mol. Genet. 7, 13–26 (1998)
Eichler, E. E. et al. Duplication of a gene-rich cluster between 16p11.1 and Xq28: a novel pericentromeric-directed mechanism for paralogous genome evolution. Hum. Mol. Genet. 5, 899–912 (1996)
Ventura, M. et al. Neocentromeres in 15q24–26 map to duplicons which flanked an ancestral centromere in 15q25. Genome Res. 13, 2059–2068 (2003)
Johnson, M. E. et al. Positive selection of a gene family during the emergence of humans and African apes. Nature 413, 514–519 (2001)
Courseaux, A. & Nahon, J. L. Birth of two chimeric genes in the Hominidae lineage. Science 291, 1293–1297 (2001)
Stefansson, H. et al. A common inversion under selection in Europeans. Nature Genet. 37, 129–137 (2005)
Gonzalez, E. et al. The influence of CCL3L1 gene-containing segmental duplications on HIV-1/AIDS susceptibility. Science 307, 1434–1440 (2005)
Bailey, J. A. et al. Recent segmental duplications in the human genome. Science 297, 1003–1007 (2002)
Khaitovich, P. et al. Regional patterns of gene expression in human and chimpanzee brains. Genome Res. 14, 1462–1473 (2004)
Sebat, J. et al. Large-scale copy number polymorphism in the human genome. Science 305, 525–528 (2004)
Iafrate, A. J. et al. Detection of large-scale variation in the human genome. Nature Genet. 36, 949–951 (2004)
Tuzun, E. et al. Fine-scale structural variation of the human genome. Nature Genet. 37, 727–732 (2005)
Sharp, A. J. et al. Segmental duplications and copy number variation in the human genome. Am. J. Hum. Genet. 77, 78–88 (2005)
Fredman, D. et al. Complex SNP-related sequence variation in segmental genome duplications. Nature Genet. 36, 861–866 (2004)
Stankiewicz, P. & Lupski, J. R. Genomic architecture, rearrangements and genomic disorders. Trends Genet. 18, 74–82 (2002)
Zhang, L., Lu, H. H., Chung, W. Y., Yang, J. & Li, W. H. Patterns of segmental duplication in the human genome. Mol. Biol. Evol. 22, 135–141 (2005)
Bailey, J. A., Yavor, A. M., Massa, H. F., Trask, B. J. & Eichler, E. E. Segmental duplications: organization and impact within the current human genome project assembly. Genome Res. 11, 1005–1017 (2001)
The Chimpanzee Sequencing and Analysis Consortium. Initial sequence of the chimpanzee genome and comparison with the human genome. Nature doi:10.1038/nature04072 (this issue)
Tuzun, E., Bailey, J. A. & Eichler, E. E. Recent segmental duplications in the working draft assembly of the brown Norway rat. Genome Res. 14, 493–506 (2004)
Bailey, J. A., Church, D. M., Ventura, M., Rocchi, M. & Eichler, E. E. Analysis of segmental duplications and genome assembly in the mouse. Genome Res. 14, 789–801 (2004)
International Human Genome Sequencing Consortium, Finishing the euchromatic sequence of the human genome. Nature 431, 931–945 (2004)
Rozen, S. et al. Abundant gene conversion between arms of massive palindromes in human and ape Y chromosomes. Nature 423, 873–876 (2003)
Chen, F. C. & Li, W. H. Genomic divergences between humans and other hominoids and the effective population size of the common ancestor of humans and chimpanzees. Am. J. Hum. Genet. 68, 444–456 (2001)
Royle, N. J., Baird, D. M. & Jeffreys, A. J. A subterminal satellite located adjacent to telomeres in chimpanzees is absent from the human genome. Nature Genet. 6, 52–56 (1994)
Yunis, J. J. & Prakash, O. The origin of man: a chromosomal pictorial legacy. Science 215, 1525–1530 (1982)
Fan, Y., Linardopoulou, E., Friedman, C., Williams, E. & Trask, B. J. Genomic structure and evolution of the ancestral chromosome fusion site in 2q13–2q14.1 and paralogous regions on other human chromosomes. Genome Res. 12, 1651–1662 (2002)
Fortna, A. et al. Lineage-specific gene duplication and loss in human and great ape evolution. PLoS Biol. 2, E207 (2004)
Khaitovich, P. et al. Parallel patterns of evolution in the genomes and transcriptomes of humans and chimpanzees. Science (in the press)
Acknowledgements
We thank M. Lachmann, I. Hellman and G. Vessere for technical assistance; the Chimpanzee Sequencing and Analysis Consortium for access to the chimpanzee sequence data before publication; A. Force for discussions; and J. Pecotte, S. Warren and J. Rogers for providing some of the primate material used in this study. This work was supported by grants from the National Human Genome Research Institute, the National Institute of General Medical Sciences, Centro di Eccellenza Geni in campo Biosanitario e Agroalimentare, Ministero Italiano della Università e della Ricerca, the European Commission and the Bundesministerium für Bildung und Forschung.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
Reprints and permissions information is available at npg.nature.com/reprintsandpermissions. The authors declare no competing financial interests.
Supplementary information
Supplementary Methods
Additional description of the methods used in this study. (DOC 60 kb)
Supplementary Figure Legends
Text to accompany the below Supplementary Figures. (DOC 41 kb)
Supplementary Figures
This file contains Supplementary Figures S1-S6, S8. (PPT 7715 kb)
Supplementary Figure S7
Chromosome views of chimpanzee and human segmental duplications, chromosome 1. (See chimpparalogy.gs.washington.edu) (PDF 14982 kb)
Supplementary Figure S7
Chromosome views of chimpanzee and human segmental duplications, chromosome 2. (PDF 15010 kb)
Supplementary Figure S7
Chromosome views of chimpanzee and human segmental duplications, chromosome 3. (PDF 12402 kb)
Supplementary Figure S7
Chromosome views of chimpanzee and human segmental duplications, chromosome 4. (PDF 11967 kb)
Supplementary Figure S7
Chromosome views of chimpanzee and human segmental duplications, chromosome 5. (PDF 11326 kb)
Supplementary Figure S7
Chromosome views of chimpanzee and human segmental duplications, chromosome 6. (PDF 10573 kb)
Supplementary Figure S7
Chromosome views of chimpanzee and human segmental duplications, chromosome 7. (PDF 10055 kb)
Supplementary Figure S7
Chromosome views of chimpanzee and human segmental duplications, chromosome 8. (PDF 9002 kb)
Supplementary Figure S7
Chromosome views of chimpanzee and human segmental duplications, chromosome 9. (PDF 8338 kb)
Supplementary Figure S7
Chromosome views of chimpanzee and human segmental duplications, chromosome 10. (PDF 8433 kb)
Supplementary Figure S7
Chromosome views of chimpanzee and human segmental duplications, chromosome 11. (PDF 8437 kb)
Supplementary Figure S7
Chromosome views of chimpanzee and human segmental duplications, chromosome 12. (PDF 8164 kb)
Supplementary Figure S7
Chromosome views of chimpanzee and human segmental duplications, chromosome 13. (PDF 6962 kb)
Supplementary Figure S7
Chromosome views of chimpanzee and human segmental duplications, chromosome 14. (PDF 6348 kb)
Supplementary Figure S7
Chromosome views of chimpanzee and human segmental duplications, chromosome 15. (PDF 5892 kb)
Supplementary Figure S7
Chromosome views of chimpanzee and human segmental duplications, chromosome 16. (PDF 5599 kb)
Supplementary Figure S7
Chromosome views of chimpanzee and human segmental duplications, chromosome 17. (PDF 4884 kb)
Supplementary Figure S7
Chromosome views of chimpanzee and human segmental duplications, chromosome 18. (PDF 4574 kb)
Supplementary Figure S7
Chromosome views of chimpanzee and human segmental duplications, chromosome 19. (PDF 3927 kb)
Supplementary Figure S7
Chromosome views of chimpanzee and human segmental duplications, chromosome 20. (PDF 3946 kb)
Supplementary Figure S7
Chromosome views of chimpanzee and human segmental duplications, chromosome 21. (PDF 2800 kb)
Supplementary Figure S7
Chromosome views of chimpanzee and human segmental duplications, chromosome 22. (PDF 2872 kb)
Supplementary Figure S7
Chromosome views of chimpanzee and human segmental duplications, chromosome X. (PDF 9589 kb)
Supplementary Figure S7
Chromosome views of chimpanzee and human segmental duplications, Chimpanzee Sequence Y. (PDF 1636 kb)
Supplementary Figure S7
Chromosome views of chimpanzee and human segmental duplications, Chimpanzee Sequence, PTR Chr22. (PDF 2732 kb)
Supplementary Figure S7
Chromosome views of chimpanzee and human segmental duplications, Chimpanzee Sequence, PTR Specific Sequence. (PDF 41 kb)
Supplementary Figure S7
Chromosome views of chimpanzee and human segmental duplications, Chimpanzee Sequence, Unaligned PTR Sequence. (PDF 2175 kb)
Supplementary Table S1
Comparison of human and chimpanzee WSSD duplications (XLS 24 kb)
Supplementary Table S2
WSSD duplications and triallelic variants (XLS 18 kb)
Supplementary Table S3
FISH results for shared (CH) duplications (XLS 15 kb)
Supplementary Table S4
Chimpanzee-human array CGH vs. chimpanzee WSSD duplications (XLS 16 kb)
Supplementary Table S5
cDNA/ESTs with copy number verification from Fortna, A. et al. (XLS 19 kb)
Supplementary Table S6
Duplication statistics and duplication shadowing simulation results (XLS 207 kb)
Supplementary Table S7
Human specific gene duplications (XLS 16 kb)
Supplementary Table S8
Chimpanzee specific gene duplications (XLS 52 kb)
Supplementary Table S9
Genes differentially expressed and specifically duplicated in human or chimpanzee (XLS 35 kb)
Supplementary Table S10
Differentially expressed genes between human and chimpanzee vs. duplication regions (XLS 56 kb)
Supplementary Table S11
FISH results with chimpanzee-only duplications. (XLS 44 kb)
Supplementary Table S12
FISH results with chimpanzee-only duplications. (XLS 19 kb)
Supplementary Table S13
Human and chimpanzee single nucleotide variation in unique regions and duplication/unique transition regions. (XLS 15 kb)
Supplementary Table S14
Sequence identity of shared vs. human-only duplications (XLS 16 kb)
Supplementary Table S15
Regions where human copy # exceeds chimp copy# by >5 (XLS 131 kb)
Rights and permissions
About this article
Cite this article
Cheng, Z., Ventura, M., She, X. et al. A genome-wide comparison of recent chimpanzee and human segmental duplications. Nature 437, 88–93 (2005). https://doi.org/10.1038/nature04000
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1038/nature04000
This article is cited by
-
Generation of chimpanzee induced pluripotent stem cell lines for cross-species comparisons
In Vitro Cellular & Developmental Biology - Animal (2024)
-
Variants of a putative baseplate wedge protein extend the host range of Pseudomonas phage K8
Microbiome (2023)
-
Revised time estimation of the ancestral human chromosome 2 fusion
BMC Genomics (2022)
-
Evolutionary history of the human multigene families reveals widespread gene duplications throughout the history of animals
BMC Evolutionary Biology (2019)
-
Genes with human-specific features are primarily involved with brain, immune and metabolic evolution
BMC Bioinformatics (2019)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.