Abstract
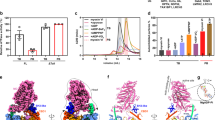
Here we solve a 2.4-Å structure of a truncated version of the reverse-direction myosin motor, myosin VI, that contains the motor domain and binding sites for two calmodulin molecules. The structure reveals only minor differences in the motor domain from that in plus-end directed myosins, with the exception of two unique inserts. The first is near the nucleotide-binding pocket and alters the rates of nucleotide association and dissociation. The second unique insert forms an integral part of the myosin VI converter domain along with a calmodulin bound to a novel target motif within the insert. This serves to redirect the effective ‘lever arm’ of myosin VI, which includes a second calmodulin bound to an ‘IQ motif’, towards the pointed (minus) end of the actin filament. This repositioning largely accounts for the reverse directionality of this class of myosin motors. We propose a model incorporating a kinesin-like uncoupling/docking mechanism to provide a full explanation of the movements of myosin VI.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
References
Berg, J. S., Powell, B. C. & Cheney, R. E. A millennial myosin census. Mol. Biol. Cell 12, 780–794 (2001)
Wells, A. L. et al. Myosin VI is an actin-based motor that moves backwards. Nature 401, 505–508 (1999)
Geisbrecht, E. R. & Montell, D. J. Myosin VI is required for E-cadherin-mediated border cell migration. Nature Cell Biol. 4, 616–620 (2002)
Hasson, T. Myosin-VI: two distinct roles in endocytosis. J. Cell Sci. 116, 3453–3461 (2003)
Buss, F., Spudich, G. & Kendrick-Jones, J. Myosin VI: Cellular functions and motor properties. Annu. Rev. Cell Dev. Biol. 20, 649–676 (2004)
Millo, H., Leaper, K., Lazou, V. & Bownes, M. Myosin VI plays a role in cell-cell adhesion during epithelial morphogenesis. Mech. Dev. 121, 1335–1351 (2004)
Geeves, M. A. & Holmes, K. C. Structural mechanism of muscle contraction. Annu. Rev. Biochem. 68, 687–728 (1999)
Purcell, T. J., Morris, C., Spudich, J. A. & Sweeney, H. L. Role of the lever arm in the processive stepping of myosin V. Proc. Natl Acad. Sci. USA 99, 14159–14164 (2002)
Sakamoto, T. et al. Neck length and processivity of myosin V. J. Biol. Chem. 278, 29201–29207 (2003)
Ruff, C., Furch, M., Brenner, B., Manstein, D. J. & Meyhofer, E. Single-molecule tracking of myosins with genetically engineered amplifier domains. Nature Struct. Biol. 8, 226–229 (2001)
Rock, R. S. et al. Myosin VI is a processive, backwards motor with a large step size. Proc. Natl Acad. Sci. USA 98, 13655–13659 (2001)
Nishikawa, S. et al. Class VI myosin moves processively along actin filaments backward with large steps. Biochem. Biophys. Res. Commun. 290, 311–317 (2002)
Bahloul, A. et al. The unique insert in myosin VI is a structural calcium-calmodulin binding site. Proc. Natl Acad. Sci. USA 101, 4787–4792 (2004)
Rock, R. S. et al. A flexible domain is essential for the large step size and processivity of myosin VI. Mol. Cell 17, 603–609 (2005)
Homma, K., Yoshimura, M., Saito, J., Ikebe, R. & Ikebe, M. The core of the motor domain determines the direction of myosin movement. Nature 412, 831–834 (2001)
Coureux, P.-D. et al. A structural state of the myosin V motor without bound nucleotide. Nature 425, 419–423 (2003)
Reubold, T. F., Eschenburg, S., Becker, A., Kull, F. J. & Manstein, D. J. A structural model for actin-induced nucleotide release in myosin. Nature Struct. Biol. 10, 826–830 (2003)
Coureux, P.-D., Sweeney, H. L. & Houdusse, A. Three myosin V structures delineate essential features of chemo-mechanical transduction. EMBO J. 23, 4527–4537 (2004)
Holmes, K. C., Schröeder, R. R., Sweeney, H. L. & Houdusse, A. The structure of the rigor complex and its implications for the power stroke. Phil. Trans. R. Soc. Lond. B 359, 1819–1828 (2004)
De La Cruz, E. M., Ostap, E. M. & Sweeney, H. L. Kinetic mechanism and regulation of myosin VI. J. Biol. Chem. 276, 32373–32381 (2001)
De La Cruz, E. M., Wells, A. L., Rosenfeld, S. S., Ostap, E. M. & Sweeney, H. L. The kinetic mechanism of myosin V. Proc. Natl Acad. Sci. USA 96, 13726–13731 (1999)
Sweeney, H. L. & Houdusse, A. The motor mechanism of myosin V: insights for muscle contraction. Phil. Trans. R. Soc. Lond. B 359, 1829–1842 (2004)
Rosenfeld, S. S., Houdusse, A. & Sweeney, H. L. Magnesium regulates ADP dissociation from myosin V. J. Biol. Chem. 280, 6072–6079 (2005)
Rhoads, A. R. & Friedberg, F. Sequence motifs for calmodulin recognition. FASEB J. 11, 331–340 (1997)
Meador, W. E., Means, A. R. & Quiocho, F. A. Target enzyme recognition by calmodulin: 2.4 Å structure of a calmodulin–peptide complex. Science 257, 1251–1255 (1992)
Fallon, J. L. & Quiocho, F. A. A closed compact structure of native Ca2+-calmodulin. Structure 11, 1303–1307 (2003)
Altman, D., Sweeney, H. L. & Spudich, J. A. The mechanism of myosin VI translocation and its load-induced anchoring. Cell 116, 737–749 (2004)
Tsiavaliaris, G., Fujita-Becker, S. & Manstein, D. J. Molecular engineering of a backwards-moving myosin motor. Nature 427, 558–561 (2004)
Rice, S. et al. A structural change in the kinesin motor protein that drives motility. Nature 402, 778–784 (1999)
Rosenfeld, S. S., Fordyce, P. M., Jefferson, G. M., King, P. H. & Block, S. M. Stepping and stretching: how kinesin uses internal strain to walk processively. J. Biol. Chem. 278, 18530–18536 (2003)
Houdusse, A., Kalabokis, V. N., Himmel, D., Szent-Gyorgyi, A. G. & Cohen, C. Atomic structure of scallop myosin subfragment S1 complexed with MgADP: a novel conformation of the myosin head. Cell 97, 459–470 (1999)
Yildiz, A. et al. Myosin VI steps via a hand-over-hand mechanism with its lever arm undergoing fluctuations when attached to actin. J. Biol. Chem. 279, 37223–37226 (2004)
Robblee, J. P., Olivares, A. O. & De la Cruz, E. M. Mechanism of nucleotide binding to actomyosin VI: evidence for allosteric head-head communication. J. Biol. Chem. 279, 38608–38617 (2004)
Sasaki, N., Ohkura, R. & Sutoh, K. Dictyostelium myosin II mutations that uncouple the converter swing and ATP hydrolysis cycle. Biochemistry 42, 90–95 (2003)
Otwinowski, Z. & Minor, W. Processing X-ray diffraction data collected in oscillation mode. Methods Enzymol. 276, 307–326 (1997)
Collaborative Computational Project No. 4., The CCP4 suite: programs for protein crystallography. Acta Crystallogr. D 50, 760–763 (1994)
Navaza, J. AMoRe: an automated package for molecular replacement. Acta Crystallogr. A 50, 157–163 (1994)
Roussel, A. & Cambillaud, C. Silicon Graphics Geometry Partner Directory 77–78 (Silicon Graphics, Mountain View, California, 1989)
Brünger, A. T. et al. Crystallography & NMR system: a new software suite for macromolecular structure determination. Acta Crystallogr. D 54, 905–921 (1998)
Perrakis, A., Morris, R. M. & Lamzin, V. S. Automated protein model building combined with iterative structure refinement. Nature Struct. Biol. 6, 458–463 (1999)
Kraulis, P. J. MOLSCRIPT: a program to produce both detailed and schematic plots of protein structures. J. Appl. Crystallogr. 24, 946–950 (1991)
Houdusse, A., Szent-Györgyi, A. & Cohen, C. Three conformational states of scallop myosin subfragment S1. Proc. Natl Acad. Sci. USA 97, 11238–11243 (2000)
Acknowledgements
We thank the staff of the European Synchrotron Radiation Facility for assistance during data collection, A. Li and D. Garbett for technical assistance in preparing the recombinant proteins, and J. Cicolari for assistance in crystallization experiments. This work was supported by a grant from the National Institutes of Health (NIAMS) to H.L.S and A.H., grants from the CNRS and the ARC to A.H., and predoctoral fellowships from the Quebec government and the FRM to A.B.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
Atomic coordinates and structure factors have been deposited in the Protein Data Bank under the accession numbers 2BKH and r2bkhsf for MDins2 and 2BKI and r2bkisf for long MDins2IQ. Reprints and permissions information is available at npg.nature.com/reprintsandpermissions. The authors declare no competing financial interests.
Supplementary information
Supplementary Methods
Additional information on the methods used in this study. (DOC 78 kb)
Supplementary Data
Analysis of the role of insert1 in myosin VI kinetics. (DOC 41 kb)
Supplementary Table S1
Steady-state and transient kinetic parameters of myosin VI short MDins2IQ with and without insert-1. (DOC 33 kb)
Supplementary Table S2
Data collection and refinement statistics (DOC 53 kb)
Supplementary Figure S1
Lever arm of myosin VI MDins2IQ. Shown is a ribbon representation of the lever arm of the myosin VI MDins2IQ structure. The electron density map clearly shows that the long heavy chain helix found for the insert2 sequence (dark purple) remains straight and extends to form the first part of the IQ motif (cyan), which binds the C-terminal semi-open lobe of an apo-calmodulin (yellow). Variability is however observed in the orientation of the second part of the IQ motif and in most of the helices of the N-terminal lobe of its associated CaM. These regions of higher disorder were not included in the final coordinates and are shown in white in this figure. They are modelled using the coordinates of a Ca2+-free CaM bound to the first IQ motif from mouse myosin V (A.H. unpublished data). (PDF 1360 kb)
Supplementary Movie S1
The 50kDa cleft is not totally closed in this structure of the nucleotide-free Myosin VI motor. Cleft closure occur via interactions between the U50kDa (blue) and L50kDa (white) residues on either side of the cleft. These are only found in the rigor-like myosin V structure. The cleft is totally open in the myosin V post-rigor state. In the myosin VI structure, the cleft is slightly more closed than that of the Dictyostelium discoideum myosin II structure in the absence of nucleotide, but these two structures differ from the myosin V structure and do not correspond to the rigor-like state. (GIF 762 kb)
Supplementary Movie S2
The distortion of the central β-sheet that controls rearrangements in the myosin motor domain differs between Myosin V and VI. (GIF 342 kb)
Supplementary Movie S3
The SH1 helix / N-terminal subdomain interface differs between Myosin V and VI. (GIF 501 kb)
Supplementary Movie S4
Directionality and power-stroke in myosin motors. Animation related to Figure 5. (GIF 1547 kb)
Rights and permissions
About this article
Cite this article
Ménétrey, J., Bahloul, A., Wells, A. et al. The structure of the myosin VI motor reveals the mechanism of directionality reversal. Nature 435, 779–785 (2005). https://doi.org/10.1038/nature03592
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1038/nature03592
This article is cited by
-
How myosin VI traps its off-state, is activated and dimerizes
Nature Communications (2023)
-
Diverse functions of myosin VI in spermiogenesis
Histochemistry and Cell Biology (2021)
-
Synthetic biology approaches to dissecting linear motor protein function: towards the design and synthesis of artificial autonomous protein walkers
Biophysical Reviews (2020)
-
Structure of Myosin VI/Tom1 complex reveals a cargo recognition mode of Myosin VI for tethering
Nature Communications (2019)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.