Abstract
Entamoeba histolytica is an intestinal parasite and the causative agent of amoebiasis, which is a significant source of morbidity and mortality in developing countries1. Here we present the genome of E. histolytica, which reveals a variety of metabolic adaptations shared with two other amitochondrial protist pathogens: Giardia lamblia and Trichomonas vaginalis. These adaptations include reduction or elimination of most mitochondrial metabolic pathways and the use of oxidative stress enzymes generally associated with anaerobic prokaryotes. Phylogenomic analysis identifies evidence for lateral gene transfer of bacterial genes into the E. histolytica genome, the effects of which centre on expanding aspects of E. histolytica's metabolic repertoire. The presence of these genes and the potential for novel metabolic pathways in E. histolytica may allow for the development of new chemotherapeutic agents. The genome encodes a large number of novel receptor kinases and contains expansions of a variety of gene families, including those associated with virulence. Additional genome features include an abundance of tandemly repeated transfer-RNA-containing arrays, which may have a structural function in the genome. Analysis of the genome provides new insights into the workings and genome evolution of a major human pathogen.
Similar content being viewed by others
Main
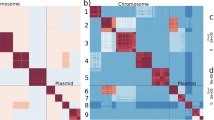
Genome analysis was carried out on a 12.5-fold coverage genome assembly consisting of 23,751,783 base pairs (bp) distributed among 888 scaffolds. The 9,938 predicted genes average 1.17 kilobases (kb) in size and comprise 49% of the genome. One-quarter of E. histolytica genes are predicted to contain introns, with 6% of genes containing multiple introns. No homologues could be identified for a third of predicted proteins (31.8%) from the public databases (see Methods). E. histolytica chromosomes do not condense, and the uncertainty surrounding its ploidy and the extensive length variability observed between homologous chromosomes from different isolates makes the exact chromosome number difficult to determine. The chromosome size variation observed may be due to expansion and contraction of subtelomeric repeats, as in other protists2,3, and it is tempting to speculate that in E. histolytica these regions consist of tRNA-containing arrays. Comprising almost 10% of the sequence reads, 25 types of long tandem array, each containing between one and five tRNA types per repeat unit, could be identified from the genome data. The full complement of tRNAs required for translation has been identified, and all but four of the tRNA genes are encoded exclusively in arrays. These unique tRNA gene arrays are thus predicted to be functional as well as potentially fulfilling a structural role in the genome. No association could be determined between codon usage and the relative copy numbers of their cognate tRNA species.
The metabolism of E. histolytica seems to have been shaped by secondary gene loss and lateral gene transfer (LGT), primarily from bacterial lineages (Fig. 1). E. histolytica is an obligate fermenter, using bacterial-like fermentation enzymes and lacking proteins of the tricarboxylic acid cycle and mitochondrial electron transport chain. An atrophic, mitochondrion-derived organelle has been identified in E. histolytica4, and the genome data support the absence of a mitochondrial genome. Glucose is the main energy source; however, in place of the typical eukaryotic glucose transporters those of E. histolytica are related to the prokaryote glucose/ribose porter family, with the amino- and carboxy-terminal domains switched relative to their prokaryotic counterparts.
Arrows indicate enzyme reactions. Glycolysis and fermentation are the major energy generation pathways. Green arrows represent enzymes encoded by genes that are among the 96 candidates for LGT into the E. histolytica genome. Broken arrows indicate enzymes for which no gene could be identified using searches of the genome data, although the activity is likely to be present. The yellow arrow points to the source of electrons for activation of metronidazole, the major drug for treatment of amoebic liver abscess. DK, pyruvate phosphate dikinase; GlcNAc, N-acetylglucosamine; GPI, glycosylphosphatidylinositol; K, pyruvate kinase; LCFA, long-chain fatty acid; PAPS, phosphoadenosine phosphosulphate; PEP, phosphoenolpyruvate; PP, pyrophosphate; PRPP, phosphoribosyl pyrophosphate; VLCFA, very-long-chain fatty acid.
As a phagocytic resident of the human gut, E. histolytica has access to many bacterial and host-derived preformed organic compounds. Most pathways for amino acid biosynthesis have been eliminated, except those for serine and cysteine, which are probably retained for the production of cysteine, the major intracellular thiol. The high levels of cysteine in E. histolytica may compensate for the lack of glutathione and its associated enzymes, a major component of oxidative stress resistance in many organisms5. E. histolytica lacks de novo purine, pyrimidine and thymidylate synthesis and must rely on salvage pathways, similar to G. lamblia and T. vaginalis6. In addition, E. histolytica appears to lack ribonucleotide reductase, a characteristic that it shares with G. lamblia7. E. histolytica is unable to synthesize fatty acids but retains the ability to synthesize a variety of phospholipids. The absence of identifiable pathways for the synthesis of isoprenoids and the sphingolipid head group aminoethylphosphonate suggest the existence of novel pathways. These pathways, once characterized, might represent attractive drug targets. Two unusual enzymes of fatty acid elongation are shared between E. histolytica and G. lamblia, including a predicted acetyl-CoA carboxylase with two carboxyltransferase domains8. We propose that this enzyme removes a carboxyl group from oxaloacetate and transfers it to acetyl-CoA to form malonyl-CoA and pyruvate. E. histolytica also has five members of a fatty acid elongase family, previously identified only in plants, green algae and G. lamblia9,10. Folate is a cofactor essential for thymidylate synthesis and methionine recycling, and genome analysis reveals a complete lack of genes coding for known folate-dependent enzymes and folate transporters. Folate is also required for organelle protein synthesis in mitochondria and chloroplasts, and loss of the mitochondrial genome may have paved the way for the loss of these folate-dependent functions.
LGT is an important force in the evolution of prokaryotes but significantly less is known about its importance in eukaryotic evolution11. We conducted a phylogenetic screen of the Entamoeba genome for cases of relatively recent prokaryote to eukaryote LGT (see Methods), and for 96 genes we believe that this is the simplest explanation for the tree topologies obtained (see Supplementary Information). These genes are embedded among typically eukaryotic genes on E. histolytica scaffolds and do not seem to represent contaminating prokaryotic sequences. Most (58%) of the LGT genes encode a variety of metabolic enzymes, whereas most of the remaining genes (41%) encode proteins of unknown function (Supplementary Fig. 1). The major impact is in the area of carbohydrate and amino acid metabolism, where they have increased the range of substrates available for energy generation including tryptophanase and aspartase, which contribute to the use of amino acids. Several glycosidases and sugar kinases appear to have been acquired through LGT and would probably enable E. histolytica to use sugars other than glucose; for example, fructose and galactose. There is a strong bias in the data for a major donor being in the Cytophaga–Flavobacterium–Bacteroides (CFB) group of the phylum Bacteroidetes; however, this should be interpreted with caution, as current sampling of prokaryotic genomes is still relatively incomplete. It is clear that among the 96 genes, some result in significant enhancements to E. histolytica metabolism, thus contributing to its biology to a greater extent than indicated by the numbers alone.
E. histolytica feed on bacteria in the lumen of the colon and lyse host epithelial cells after invasion of the intestinal wall12. A number of amoebic virulence determinants have been characterized, including a multi-subunit GalGalNAc lectin involved in adhesion to host cells, cysteine proteases that degrade host extracellular matrix, and pore-forming peptides (amoebapores) capable of lysing target cells12. Analysis of the genome reveals redundancy in the genes encoding these virulence factors. Thirty homologues of the intermediate subunit and one homologue of the heavy subunit of the GalGalNAc lectin were identified. Ten new cysteine proteinases with predicted N-terminal transmembrane anchors, which might allow them to be localized on the amoeba cell surface, were identified. In addition to three new amoebapores a homologue of haemolysin III was identified, suggesting that, in addition to amoebapores, haemolysins may have a role in host cell lysis.
Vesicle trafficking has a role in E. histolytica pathogenesis through phagocytosis and the delivery of secreted hydrolytic enzymes and amoebapores to the cell surface13. E. histolytica lacks morphologically identifiable rough endoplasmic reticulum and the Golgi apparatus14 but encodes the basic elements of the vesicle transport machinery common to other eukaryotic cells, with the coat complexes COPI, COPII, clathrin and retromer all being present. Rab and Arf protein family expansions reflect the increased complexity and number of vesicle fusion and recycling steps that have been associated with phagocytosis and pinocytosis in amoebae15. The cytoskeleton has a number of important roles in parasite motility, contact-dependant killing and phagocytosis of host intestinal epithelial cells16. This is reflected in expansions of Rho GTPases and their regulators RhoGAPs and RhoGEFs, which control a number of processes involving the actin cytoskeleton. Five proteins with a unique domain architecture containing both RhoGEF and ArfGAP domains were identified, suggesting a mechanism for direct communication between the regulators of vesicle budding and cytoskeletal rearrangement.
E. histolytica uses a complex mix of signal transduction systems in order to sense and interact with the different environments it encounters (Fig. 2). Almost 270 putative E. histolytica protein kinases representing members of all seven families of the eukaryotic protein kinase superfamily were identified17. These include tyrosine kinases with SH2 domains, tyrosine kinase-like protein kinases and 90 putative receptor Ser/Thr kinases. These Ser/Thr kinases are uncommon in protists, appear to be absent from Dictyostelium and have previously been described only in plants, animals and Choanoflagellates. The E. histolytica receptor Ser/Thr kinases all contain an N-terminal signal peptide, a predicted extracellular domain and a single transmembrane helix followed by a cytosolic tyrosine kinase-like domain. The receptor kinases fall into three groups on the basis of differences in their predicted extracellular domains. The first group of 50 receptor kinase proteins contains CXXC-rich repeats similar to those found in the intermediate subunit (Igl) of the Gal/GalNAc lectin and G. lamblia variant-specific surface proteins. A second group of 32 proteins encodes cysteine-rich domains containing CXC repeats. The third group of eight receptor kinase-like proteins lacks cysteine-rich extracellular domains. Although no immediate downstream effectors to the amoebic receptor kinases could be identified, E. histolytica contains greater than 100 protein phosphatases, which dephosphorylate proteins. An unusual feature of some of the phosphatases is the presence of varying numbers of leucine-rich repeat (LRR) domains that are involved primarily in protein–protein interactions and have not previously been associated with phosphatases. The E. histolytica genome encodes numerous putative seven-transmembrane receptors and trimeric G proteins, which are probably involved in mediating autocrine stimulation of encystation18. In contrast to autocrine stimulation of Dictyostelium sporulation, which uses secreted cyclic AMP, E. histolytica encystment is self-stimulated by secreted catecholamines18. Finally, E. histolytica has numerous cytosolic proteins involved in signal transduction, including Ras-family proteins, EF-hand calcium-binding proteins, phosphatidylinositol-3-OH kinase and MAP kinases. This represents the most varied set of signal-transduction-related proteins yet described in a single-celled eukaryote.
E. histolytica possesses three types of receptor serine/threonine kinases: one group has CXXC repeats in the extracellular domain; a second has CXC repeats; and a third has non-cysteine rich (NCR) repeats. E. histolytica has cytosolic tyrosine kinases (TyrK), but not receptor tyrosine kinases. Some serine/threonine phosphatases (S/TP) have an attached LRR domain. CaBP, calcium-binding protein; DAG, diacylglycerol; G, G protein; GAP, GTPase-activating protein; GEF, guanine nucleotide exchange factor; IP3, inositol-1,4,5-trisphosphate; PI(3)K, phosphatidylinositol-3-OH kinase; PIP2, phosphatidylinositol-4,5-bisphosphate; PIP3, phosphatidylinositol-3,4,5-trisphosphate; PKC, protein kinase C; PLC, phospholipase C; PTEN, phosphatase and tensin homologue; TyrP, tyrosine phosphatase; 7TM receptors, seven-transmembrane receptors.
In contrast to life in the anoxic colon, E. histolytica encounters a relatively high-oxygen environment during invasive amoebiasis, and coping with this change is therefore an important virulence factor. The importance of this response is underscored by the redundancy of oxygen detoxification mechanisms. E. histolytica has four copies of flavoprotein A, which detoxifies nitric oxide and/or oxygen19 (Fig. 3), and also contains rubrerythrin, which in anaerobic bacteria is protective against intracellular hydrogen peroxide20 (Fig. 3). These oxidative and/or nitrosative stress resistance genes are shared with G. lamblia (with the exception of rubrerythrin) and T. vaginalis, but have generally been associated with anaerobic prokaryotes (Fig. 3).
Enzymes boxed and shaded have previously only been identified in anaerobic prokaryotes and amitochondrial protists. a, Superoxide is detoxified by an iron-containing superoxide dismutase (Fe-SOD). Molecular oxygen is reduced to hydrogen peroxide by the NADPH-flavin oxidoreductase (p34), which also transfers electrons to peroxiredoxin (p29). Rubrerythrin (Rbr) is predicted to convert hydrogen peroxide to water, although the source of electrons for rubrerythrin in E. histolytica is unknown. b, A-type flavoproteins (FprA) detoxify nitric oxide to nitrous oxide. FprA receives electrons from flavoprotein A reductase (Far).
E. histolytica is the first amoeba genome to be fully sampled, and comparisons with other genomes will assist in resolving fundamental issues relating to eukaryote and amoeba phylogeny, as well as how LGT affects eukaryotes. Despite a lack of representative genome sampling from amitochondrial protist lineages it is already clear that these unrelated anaerobic eukaryotes seem to use convergent metabolic strategies imposed by their environments. As a first insight into an amitochondrial protist genome, analysis of these data and particularly the bacterial-like proteins contained therein should illuminate future efforts aimed at the development of diagnostics and therapeutics of these luminal parasites.
Methods
Genome sequencing and assembly
The E. histolytica genome sequence was generated by the whole-genome shotgun method. As the chromosomes of E. histolytica could not be resolved by pulsed field gel electrophoresis (PFGE) and the A + T content precluded making large or medium insert libraries in bacterial artificial chromosomes (BACs), we were required to use the whole-genome shotgun approach to sequence the genome. Genomic DNA was prepared from E. histolytica strain HM-1:IMSS (ATCC number 30459) grown axenically in TYI-S-33 medium20. At TIGR 390,000 reads were produced from a small (1.5–2.0 kb) and a medium insert library (8–10 kb) generated in the pHOS2 vector. At the Sanger Institute, 200,000 reads were generated from a pUC18 library with average insert size of 2.5 kb plus 6,500 reads from a BAC library with an average insert size of 10 kb (the high A + T content of the genomic DNA prevented cloning of larger fragments). To avoid assembly problems, reads containing episomal-derived rDNA or tRNA-containing sequences (170,000 reads (29%)) were excluded from the whole-genome assembly process. The average edited read length was 645 bp, giving an approximate 12.5-fold genome coverage. Genome assembly was carried out at the Sanger Centre using the program phusion21. All scaffolds smaller than 2 kb (327) were subsequently removed, leaving 1,425 scaffolds with a combined size of 25,393,225 bp. The remaining scaffolds were analysed to remove redundancy that may have resulted as a consequence of allelic differences or aneuploidy. We removed all scaffolds smaller than 5 kb that shared 98% or more nucleotide sequence identity over greater than 95% of their lengths. Removal of these scaffolds left 888 scaffolds remaining, with a total length of 23,751,783 bp. All scaffolds removed during the clean-up process as well as any singleton reads, although not used in the annotation process, were used in determining the presence or absence of genes in the E. histolytica genome. Unfortunately, there is no map to order the scaffolds generated by the assembly; however, the sequence generated by this project should assist in making maps for this genome in the future, and although the large-scale structure of the genome has been lost, the vast majority of the genes that have been predicted are full length with intact 3′ and 5′ untranslated regions.
Annotation
The Combiner algorithm was used for gene structure identification22 using two genefinder programs, phat23 and GlimmerHMM24, trained using a set of published E. histolytica gene sequences, alignments of protein homologues to the genomic sequence and alignment of a set of E. histolytica complementary DNA sequences (provided by N. Guillén) to the genomic sequence. The Combiner gene predictions were then manually curated. Functional annotations for the predicted proteins were automatically generated using a combination of numerous sources of evidence including searches against a non-redundant protein database and identification of functional domains by searches against the Pfam database25. tRNAs were detected using the tRNAscan-SE26 program with default parameters.
Identification of sequence homologues in other species
Sequence homologues from other species were identified by searching the predicted proteins from the E. histolytica genome against the publicly available nr database of GenBank using BlastP (http://www.ncbi.nlm.nih.gov/BLAST/) and filtering search results with an e-value of 10-5 or less, which was chosen because of the relatively large divergence between E. histolytica and other organisms for which the genomes have been sequenced and for which protein data are available.
Phylogenetic analysis
We modified a published suite of scripts and modules called PyPhy27 to make an automated genome-wide primary screen for LGT. PyPhy was used to make bootstrap (100 replicates) consensus p-distance trees from edited alignments of 5,740 E. histolytica proteins; that is, those for which there were sufficient homologues (> 4) in SwissProt and TrEMBL to make trees. The trees were analysed to identify cases where the nearest neighbour to the E. histolytica protein was a prokaryotic sequence. As an additional screen for LGT we identified all proteins for which a prokaryote was the top Blast hit. After manual inspection of the alignments, Blast outputs, tree support values and sequence identities, 279 cases of potential LGT were retained for more detailed phylogenetic analyses. Each candidate LGT was analysed by MrBayes28 using the WAG matrix, a gamma correction for site rate variation and a proportion (pinvar) of invariant sites. The analyses were run for 600,000 generations and sampled every 100 generations, with the first 2,000 samples discarded as burn-in. A consensus tree was made from the remaining samples. Because posterior probabilities—the support values used by bayesian analysis to indicate confidence in groups—have been criticized29, we also used bootstrapping to provide an additional indication of support for relationships. Each data set was bootstrapped (100 replicates) and used to make distance matrices under the same evolutionary model as in the bayesian analysis, using custom (P4) software (available on request). Trees were made from the distance matrices using FastME30 and a bootstrap consensus tree made using P4. On the basis of these analyses we identified 96 genes in which the tree topology is consistent with prokaryote to eukaryote LGT. Blast summary statistics, trees and support values for these 96 candidate LGT are provided as Supplementary Information.
References
Stanley, S. L. Jr Amoebiasis. Lancet 361, 1025–1034 (2003)
Patarapotikul, J. & Langsley, G. Chromosome size polymorphism in Plasmodium falciparum can involve deletions of the subtelomeric pPFrep20 sequence. Nucleic Acids Res. 16, 4331–4340 (1988)
Melville, S. E., Gerrard, C. S. & Blackwell, J. M. Multiple causes of size variation in the diploid megabase chromosomes of African trypanosomes. Chromosome Res. 7, 191–203 (1999)
Leon-Avila, G. & Tovar, J. Mitosomes of Entamoeba histolytica are abundant mitochondrion-related remnant organelles that lack a detectable organellar genome. Microbiology 150, 1245–1250 (2004)
Fahey, R. C., Newton, G. L., Arrick, B., Overdank-Bogart, T. & Aley, S. B. Entamoeba histolytica: a eukaryote without glutathione metabolism. Science 224, 70–72 (1984)
Abrahamsen, M. S. et al. Complete genome sequence of the apicomplexan, Cryptosporidium parvum . Science 304, 441–445 (2004)
Baum, K. F., Berens, R. L., Marr, J. J., Harrington, J. A. & Spector, T. Purine deoxynucleoside salvage in Giardia lamblia . J. Biol. Chem. 264, 21087–21090 (1989)
Jordan, I. K., Henze, K., Fedorova, N. D., Koonin, E. V. & Galperin, M. Y. Phylogenomic analysis of the Giardia intestinalis transcarboxylase reveals multiple instances of domain fusion and fission in the evolution of biotin-dependent enzymes. J. Mol. Microbiol. Biotechnol. 5, 172–189 (2003)
James, D. W. Jr et al. Directed tagging of the Arabidopsis fatty acid elongation1 (FAE1) gene with the maize transposon activator. Plant Cell 7, 309–319 (1995)
Azachi, M. et al. Salt induction of fatty acid elongase and membrane lipid modifications in the extreme halotolerant alga Dunaliella salina . Plant Physiol. 129, 1320–1329 (2002)
Lawrence, J. G. & Hendrickson, H. Lateral gene transfer: when will adolescence end? Mol. Microbiol. 50, 739–749 (2003)
Huston, C. D. Parasite and host contributions to the pathogenesis of amebic colitis. Trends Parasitol. 20, 23–26 (2004)
Welter, B. H. & Temesvari, L. A. A unique Rab GTPase, EhRabA, of Entamoeba histolytica, localizes to the leading edge of motile cells. Mol. Biochem. Parasitol. 135, 185–195 (2004)
Mazzuco, A., Benchimol, M. & De Souza, W. Endoplasmic reticulum and Golgi-like elements in Entamoeba . Micron 28, 241–247 (1997)
Duhon, D. & Cardelli, J. The regulation of phagosome maturation in Dictyostelium . J. Muscle Res. Cell Motil. 23, 803–808 (2002)
Voigt, H. & Guillen, N. New insights into the role of the cytoskeleton in phagocytosis of Entamoeba histolytica . Cell. Microbiol. 1, 195–203 (1999)
Hunter, T. Protein kinase classification. Methods Enzymol. 200, 3–37 (1991)
Coppi, A., Merali, S. & Eichinger, D. The enteric parasite Entamoeba uses an autocrine catecholamine system during differentiation into the infectious cyst stage. J. Biol. Chem. 277, 8083–8090 (2002)
Gomes, C. M. et al. A novel type of nitric-oxide reductase. Escherichia coli flavorubredoxin. J. Biol. Chem. 277, 25273–25276 (2002)
Sztukowska, M., Bugno, M., Potempa, J., Travis, J. & Kurtz, D. M. Jr Role of rubrerythrin in the oxidative stress response of Porphyromonas gingivalis . Mol. Microbiol. 44, 479–488 (2002)
Mullikin, J. C. & Ning, Z. The phusion assembler. Genome Res. 13, 81–90 (2003)
Allen, J. E., Pertea, M. & Salzberg, S. L. Computational gene prediction using multiple sources of evidence. Genome Res. 14, 142–148 (2004)
Cawley, S. E., Wirth, A. I. & Speed, T. P. Phat–a gene finding program for Plasmodium falciparum . Mol. Biochem. Parasitol. 118, 167–174 (2001)
Majoros, W. H., Pertea, M. & Salzberg, S. L. TigrScan and GlimmerHMM: two open-source ab initio eukaryotic gene-finders. Bioinformatics 20, 2878–2879 (2004)
Bateman, A. et al. The Pfam protein families database. Nucleic Acids Res. 32, D138–D141 (2004)
Lowe, T. M. & Eddy, S. R. tRNAscan-SE: a program for improved detection of transfer RNA genes in genomic sequence. Nucleic Acids Res. 25, 955–964 (1997)
Sicheritz-Ponten, T. & Andersson, S. G. A phylogenomic approach to microbial evolution. Nucleic Acids Res. 29, 545–552 (2001)
Huelsenbeck, J. P. & Ronquist, F. MRBAYES: Bayesian inference of phylogenetic trees. Bioinformatics 17, 754–755 (2001)
Cummings, M. P. et al. Comparing bootstrap and posterior probability values in the four-taxon case. Syst. Biol. 52, 477–487 (2003)
Desper, R. & Gascuel, O. Theoretical foundation of the balanced minimum evolution method of phylogenetic inference and its relationship to weighted least-squares tree fitting. Mol. Biol. Evol. 21, 587–598 (2004)
Acknowledgements
This work was supported by grants from the National Institute of Allergy and Infectious Disease and the Wellcome Trust.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare that they have no competing financial interests.
Supplementary information
Supplementary Data
This file contains a table with 96 candidate LGT genes, a pie chart with functional categorization of LGT genes and a list of .pdf files containing phylogenetic trees for each candidate. (DOC 217 kb)
Supplementary Notes
This file contains the GenBank accessions. (XLS 44 kb)
Supplementary Figures
This zipped file contains pdf files of phylogenetic trees for 96 E. histolytica genes. (ZIP 1439 kb)
Rights and permissions
This article is distributed under the terms of the Creative Commons Attribution-Non-Commercial-Share Alike licence (http://creativecommons.org/licenses/by-nc-sa/3.0/), which permits distribution, and reproduction in any medium, provided the original author and source are credited. This licence does not permit commercial exploitation, and derivative works must be licensed under the same or similar licence.
About this article
Cite this article
Loftus, B., Anderson, I., Davies, R. et al. The genome of the protist parasite Entamoeba histolytica. Nature 433, 865–868 (2005). https://doi.org/10.1038/nature03291
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1038/nature03291
This article is cited by
-
Glycerol biosynthetic pathway plays an essential role in proliferation and antioxidative defense in the human enteric protozoan parasite Entamoeba histolytica
Scientific Reports (2023)
-
The draft genome of Cochliopodium minus reveals a complete meiosis toolkit and provides insight into the evolution of sexual mechanisms in Amoebozoa
Scientific Reports (2022)
-
The non-LTR retrotransposons of Entamoeba histolytica: genomic organization and biology
Molecular Genetics and Genomics (2022)
-
Detection of horizontal gene transfer in the genome of the choanoflagellate Salpingoeca rosetta
Scientific Reports (2021)
-
Possible role played by the SINE2 element in gene regulation, as demonstrated by differential processing and polyadenylation in avirulent strains of E. histolytica
Antonie van Leeuwenhoek (2021)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.