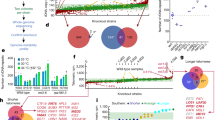
Abstract
The functional importance of the roughly 98% of mammalian genomes not corresponding to protein coding sequences remains largely undetermined1. Here we show that some large-scale deletions of the non-coding DNA referred to as gene deserts2,3,4 can be well tolerated by an organism. We deleted two large non-coding intervals, 1,511 kilobases and 845 kilobases in length, from the mouse genome. Viable mice homozygous for the deletions were generated and were indistinguishable from wild-type littermates with regard to morphology, reproductive fitness, growth, longevity and a variety of parameters assaying general homeostasis. Further detailed analysis of the expression of multiple genes bracketing the deletions revealed only minor expression differences in homozygous deletion and wild-type mice. Together, the two deleted segments harbour 1,243 non-coding sequences conserved between humans and rodents (more than 100 base pairs, 70% identity). Some of the deleted sequences might encode for functions unidentified in our screen; nonetheless, these studies further support the existence of potentially ‘disposable DNA’ in the genomes of mammals.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Bentley, D. R. Genomes for medicine. Nature 429, 440–445 (2004)
Dunham, A. et al. The DNA sequence and analysis of human chromosome 13. Nature 428, 522–528 (2004)
Scherer, S. W. et al. Human chromosome 7: DNA sequence and biology. Science 300, 767–772 (2003)
Venter, J. C. et al. The sequence of the human genome. Science 291, 1304–1351 (2001)
Jasny, B. R. & Kennedy, D. The human genome. Science 291, 1153 (2001)
Gu, Z. et al. Role of duplicate genes in genetic robustness against null mutations. Nature 421, 63–66 (2003)
Winzeler, E. A. et al. Functional characterization of the S. cerevisiae genome by gene deletion and parallel analysis. Science 285, 901–906 (1999)
Nelson, C. E., Hersh, B. M. & Carroll, S. B. The regulatory content of intergenic DNA shapes genome architecture. Genome Biol. 5, R25.1–R25.15 (2004)
Bejerano, G. et al. Ultraconserved elements in the human genome. Science 304, 1321–1325 (2004)
Ramirez-Solis, R., Liu, P. & Bradley, A. Chromosome engineering in mice. Nature 378, 720–724 (1995)
Yu, Y. & Bradley, A. Engineering chromosomal rearrangements in mice. Nature Rev. Genet. 2, 780–790 (2001)
Zhu, Y. et al. Genomic interval engineering of mice identifies a novel modulator of triglyceride production. Proc. Natl Acad. Sci. USA 97, 1137–1142 (2000)
Nobrega, M. A., Ovcharenko, I., Afzal, V. & Rubin, E. M. Scanning human gene deserts for long-range enhancers. Science 302, 413 (2003)
Lettice, L. A. et al. A long-range Shh enhancer regulates expression in the developing limb and fin and is associated with preaxial polydactyly. Hum. Mol. Genet. 12, 1725–1735 (2003)
Kimura-Yoshida, C. et al. Characterization of the pufferfish Otx2 cis-regulators reveals evolutionarily conserved genetic mechanisms for vertebrate head specification. Development 131, 57–71 (2004)
Ovcharenko, I., Loots, G. G. & Stubbs, L. Interpreting mammalian evolution using Fugu genome comparisons. Genomics (in the press)
Kothary, R. et al. Inducible expression of an hsp68–lacZ hybrid gene in transgenic mice. Development 105, 707–714 (1989)
Boffelli, D., Nobrega, M. A. & Rubin, E. M. Comparative genomics at the vertebrate extremes. Nature Rev. Genet. 5, 456–465 (2004)
Zerucha, T. et al. A highly conserved enhancer in the Dlx5/Dlx6 intergenic region is the site of cross-regulatory interactions between Dlx genes in the embryonic forebrain. J. Neurosci. 20, 709–721 (2000)
Frazer, K. A. et al. Noncoding sequences conserved in a limited number of mammals in the SIM2 interval are frequently functional. Genome Res. 14, 367–372 (2004)
Iafrate, A. J. et al. Detection of large-scale variation in the human genome. Nature Genet. 36, 949–951 (2004)
Sebat, J. et al. Large-scale copy number polymorphism in the human genome. Science 305, 525–528 (2004)
Sauer, B. & Henderson, N. Targeted insertion of exogenous DNA into the eukaryotic genome by the Cre recombinase. New Biol. 2, 441–449 (1990)
Hogan, B. C. B. & Lacy, E. Manipulating the Mouse Embryo: A Laboratory Manual (Cold Spring Harbor Laboratory Press, Plainview, New York, 1994)
Rat Genome Sequencing Project Consortium. Genome sequence of the Brown Norway rat yields insights into mammalian evolution. Nature 428, 493–521 (2004)
Mouse Genome Sequencing Consortium. Initial sequencing and comparative analysis of the mouse genome. Nature 420, 520–562 (2002)
Anderson, J. P. et al. HRC is a direct transcriptional target of MEF2 during cardiac, skeletal, and arterial smooth muscle development in vivo. Mol. Cell. Biol. 24, 3757–3768 (2004)
Dodou, E., Xu, S. M. & Black, B. L. mef2c is activated directly by myogenic basic helix–loop–helix proteins during skeletal muscle development in vivo. Mech. Dev. 120, 1021–1032 (2003)
Acknowledgements
We thank I. Ovcharenko, G. Loots and J. Schwartz for help with the identification and annotation of the gene deserts; D. Boffelli, L. Pennacchio, N. Ahituv, J. Bristow and other Rubin laboratory members for suggestions and criticisms on the manuscript; and H. Jacob for providing the clinical chemistry assays. Research was conducted at the E. O. Lawrence Berkeley National Laboratory and at the Joint Genome Institute, with support by grants from the Programs for Genomic Application, the NHLBI and the DOE.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare that they have no competing financial interests.
Supplementary information
Supplementary Methods
Methods used to generate and screen the animals generated in this study, as well as details of the annotation of the genomic intervals targeted for deletion. (DOC 36 kb)
Supplementary Tables
A total of seven tables listing all relevant primer sequences used in the protocols, and results from functional analysis of the ice concerning general fitness, survival and reproduction. (DOC 91 kb)
Supplementary Figure 1
Results from the clinical chemistry assays tested in the various strains of mice. (JPG 53 kb)
Supplementary Figure 2
Results from macroscopic pathology in 12 organs analysed. (JPG 38 kb)
Rights and permissions
About this article
Cite this article
Nóbrega, M., Zhu, Y., Plajzer-Frick, I. et al. Megabase deletions of gene deserts result in viable mice. Nature 431, 988–993 (2004). https://doi.org/10.1038/nature03022
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1038/nature03022
This article is cited by
-
A distant global control region is essential for normal expression of anterior HOXA genes during mouse and human craniofacial development
Nature Communications (2024)
-
Premature ovarian insufficiency is associated with global alterations in the regulatory landscape and gene expression in balanced X-autosome translocations
Epigenetics & Chromatin (2023)
-
The genome loading model for the origin and maintenance of sex in eukaryotes
Biologia Futura (2022)
-
Reverse-genetics studies of lncRNAs—what we have learnt and paths forward
Genome Biology (2020)
-
Trans- and cis-acting effects of Firre on epigenetic features of the inactive X chromosome
Nature Communications (2020)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.