Abstract
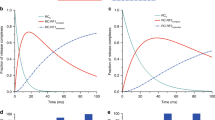
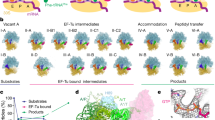
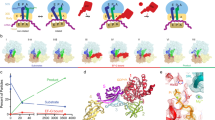
Termination of protein synthesis by the ribosome requires two release factor (RF) classes. The class II RF3 is a GTPase that removes class I RFs (RF1 or RF2) from the ribosome after release of the nascent polypeptide1,2,3. RF3 in the GDP state binds to the ribosomal class I RF complex, followed by an exchange of GDP for GTP and release of the class I RF. As GTP hydrolysis triggers release of RF3 (ref. 4), we trapped RF3 on Escherichia coli ribosomes using a nonhydrolysable GTP analogue. Here we show by cryo-electron microscopy that the complex can adopt two different conformational states. In ‘state 1’, RF3 is pre-bound to the ribosome, whereas in ‘state 2’ RF3 contacts the ribosome GTPase centre. The transfer RNA molecule translocates from the peptidyl site in state 1 to the exit site in state 2. This translocation is associated with a large conformational rearrangement of the ribosome. Because state 1 seems able to accommodate simultaneously both RF3 and RF2, whose position is known from previous studies5,6, we can infer the release mechanism of class I RFs.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Freistroffer, D. V., Pavlov, M. Y., MacDougall, J., Buckingham, R. H. & Ehrenberg, M. Release factor RF3 in E. coli accelerates the dissociation of release factors RF1 and RF2 from the ribosome in a GTP-dependent manner. EMBO J. 16, 4126–4133 (1997)
Grentzmann, G. et al. Release factor RF-3 GTPase activity acts in disassembly of the ribosome termination complex. RNA 4, 973–983 (1998)
Kisselev, L., Ehrenberg, M. & Frolova, L. Termination of translation: interplay of mRNA, rRNAs and release factors? EMBO J. 22, 175–182 (2003)
Zavialov, A. V., Buckingham, R. H. & Ehrenberg, M. A posttermination ribosomal complex is the guanine nucleotide exchange factor for peptide release factor RF3. Cell 107, 115–124 (2001)
Klaholz, B. P. et al. Structure of the Escherichia coli ribosomal termination complex with release factor 2. Nature 421, 90–94 (2003)
Rawat, U. B. et al. A cryo-electron microscopic study of ribosome-bound termination factor RF2. Nature 421, 87–90 (2003)
Yusupov, M. M. et al. Crystal structure of the ribosome at 5.5 Å resolution. Science 292, 883–896 (2001)
Laurberg, M. et al. Structure of a mutant EF-G reveals domain III and possibly the fusidic acid binding site. J. Mol. Biol. 303, 593–603 (2000)
Stark, H. et al. Visualization of elongation factor Tu on the Escherichia coli ribosome. Nature 389, 403–406 (1997)
Valle, M. et al. Cryo-EM reveals an active role for aminoacyl-tRNA in the accommodation process. EMBO J. 21, 3557–3567 (2002)
Stark, H. et al. Ribosome interactions of aminoacyl-tRNA and elongation factor Tu in the codon-recognition complex. Nature Struct. Biol. 9, 849–854 (2002)
Agrawal, R. K., Penczek, P., Grassucci, R. A. & Frank, J. Visualization of elongation factor G on the Escherichia coli 70S ribosome: the mechanism of translocation. Proc. Natl Acad. Sci. USA 95, 6134–6138 (1998)
Stark, H., Rodnina, M. V., Wieden, H. J., van Heel, M. & Wintermeyer, W. Large-scale movement of elongation factor G and extensive conformational change of the ribosome during translocation. Cell 100, 301–309 (2000)
Aevarsson, A. et al. Three-dimensional structure of the ribosomal translocase: elongation factor G from Thermus thermophilus. EMBO J. 13, 3669–3677 (1994)
Czworkowski, J., Wang, J., Steitz, T. A. & Moore, P. B. The crystal structure of elongation factor G complexed with GDP, at 2.7 Å resolution. EMBO J. 13, 3661–3668 (1994)
Wilson, D. N. et al. in The Ribosome. Structure, Function, Antibiotics and Cellular Interactions (ed. Garrett, R. A.) 495–508 (American Society for Microbiology Press, Washington, DC, 2000)
Aevarsson, A. Structure-based sequence alignment of elongation factors Tu and G with related GTPases involved in translation. J. Mol. Evol. 41, 1096–1104 (1995)
Agrawal, R. K., Heagle, A. B., Penczek, P., Grassucci, R. A. & Frank, J. EF-G-dependent GTP hydrolysis induces translocation accompanied by large conformational changes in the 70S ribosome. Nature Struct Biol. 6, 643–647 (1999)
Lancaster, L., Kiel, M. C., Kaji, A. & Noller, H. F. Orientation of ribosome recycling factor in the ribosome from directed hydroxyl radical probing. Cell 111, 129–140 (2002)
Mora, L., Zavialov, A., Ehrenberg, M. & Buckingham, R. H. Stop codon recognition and interactions with peptide release factor RF3 of truncated and chimeric RF1 and RF2 from Escherichia coli. Mol. Microbiol. 50, 1467–1476 (2003)
Zavialov, A. V., Mora, L., Buckingham, R. H. & Ehrenberg, M. Release of peptide promoted by the GGQ motif of class 1 release factors regulates the GTPase activity of RF3. Mol. Cell 10, 789–798 (2002)
van Heel, M. et al. Single-particle cryo electron microscopy: towards atomic resolution. Quart. Rev. Biophys. 33, 307–369 (2000)
van Heel, M., Harauz, G., Orlova, E. V., Schmidt, R. & Schatz, M. A new generation of the IMAGIC image processing system. J. Struct. Biol. 116, 17–24 (1996)
Rosenthal, P. B. & Henderson, R. Optimal determination of particle orientation, absolute hand, and contrast loss in single-particle electron cryomicroscopy. J. Mol. Biol. 31, 721–745 (2003)
Jones, T. A., Zou, J. Y., Cowan, S. W. & Kjeldgaard, M. Improved methods for building protein models in electron density maps and the location of errors in these models. Acta Crystallogr. A 47, 110–119 (1991)
Brünger, A. T. et al. Crystallography & NMR system: A new software suite for macromolecular structure determination. Acta Crystallogr. D 54, 905–921 (1998)
Plewniak, F. et al. PipeAlign: a new toolkit for protein family analysis. Nucleic Acids Res. 31, 3829–3832 (2003)
Matadeen, R. et al. The Escherichia coli large ribosomal subunit at 7.5 Å resolution. Structure Fold Des. 15, 1575–1583 (1999)
Acknowledgements
We thank M. Ehrenberg and A. Zavialov for purified RF3 and 70S ribosome RCs; T. Pape for assistance in the early phase of this work; S. Chen, M. Schatz, R. Schmidt, and G. Willoughby for support; and R. Buckingham for sharing data before publication. B.P.K. thanks D. Moras, P. Schultz, M. Yusupov, G. Yusupova and B. Rees for support, discussions and comments on the manuscript; O. Poch for discussions on multiple sequence alignments; E. Orlova for initial image processing suggestions; and H. Saibil for the loan of a Tecnai 12 cryo-holder. The project was financed in part by grants from the Biotechnology and Biological Sciences Research Council, the European Union, and the Centre National de la Recherche Scientifique.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
M.v.H. is a shareholder of Image Science Software GmbH, distributors of the IMAGIC-5 software.
Supplementary information
Supplementary Figure 1
Principle of local multivariate statistical analysis exemplified on a typical view (side-view of the 70S ribosome); this approach was applied to all class averages (and to the particles included in these), and allowed particle separation enabling the analysis of two specific intermediate states. (JPG 39 kb)
Supplementary Figure 2
Some representative class averages (a and c) and their corresponding re-projections from the final 3D reconstructions (b and d) of states 1 and 2, respectively. (JPG 70 kb)
Rights and permissions
About this article
Cite this article
Klaholz, B., Myasnikov, A. & van Heel, M. Visualization of release factor 3 on the ribosome during termination of protein synthesis. Nature 427, 862–865 (2004). https://doi.org/10.1038/nature02332
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1038/nature02332
This article is cited by
-
Quality control of protein synthesis in the early elongation stage
Nature Communications (2023)
-
Visualization of translation termination intermediates trapped by the Apidaecin 137 peptide during RF3-mediated recycling of RF1
Nature Communications (2018)
-
Mechanism of eIF6 release from the nascent 60S ribosomal subunit
Nature Structural & Molecular Biology (2015)
-
The palindromic DNA-bound USP/EcR nuclear receptor adopts an asymmetric organization with allosteric domain positioning
Nature Communications (2014)
-
Structure of the full human RXR/VDR nuclear receptor heterodimer complex with its DR3 target DNA
The EMBO Journal (2012)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.