Abstract
The sequence of the mouse genome is a key informational tool for understanding the contents of the human genome and a key experimental tool for biomedical research. Here, we report the results of an international collaboration to produce a high-quality draft sequence of the mouse genome. We also present an initial comparative analysis of the mouse and human genomes, describing some of the insights that can be gleaned from the two sequences. We discuss topics including the analysis of the evolutionary forces shaping the size, structure and sequence of the genomes; the conservation of large-scale synteny across most of the genomes; the much lower extent of sequence orthology covering less than half of the genomes; the proportions of the genomes under selection; the number of protein-coding genes; the expansion of gene families related to reproduction and immunity; the evolution of proteins; and the identification of intraspecies polymorphism.
Similar content being viewed by others
Main
With the complete sequence of the human genome nearly in hand1,2, the next challenge is to extract the extraordinary trove of information encoded within its roughly 3 billion nucleotides. This information includes the blueprints for all RNAs and proteins, the regulatory elements that ensure proper expression of all genes, the structural elements that govern chromosome function, and the records of our evolutionary history. Some of these features can be recognized easily in the human sequence, but many are subtle and difficult to discern. One of the most powerful general approaches for unlocking the secrets of the human genome is comparative genomics, and one of the most powerful starting points for comparison is the laboratory mouse, Mus musculus.
Metaphorically, comparative genomics allows one to read evolution's laboratory notebook. In the roughly 75 million years since the divergence of the human and mouse lineages, the process of evolution has altered their genome sequences and caused them to diverge by nearly one substitution for every two nucleotides (see below) as well as by deletion and insertion. The divergence rate is low enough that one can still align orthologous sequences, but high enough so that one can recognize many functionally important elements by their greater degree of conservation. Studies of small genomic regions have demonstrated the power of such cross-species conservation to identify putative genes or regulatory elements3,4,5,6,7,8,9,10,11,12. Genome-wide analysis of sequence conservation holds the prospect of systematically revealing such information for all genes. Genome-wide comparisons among organisms can also highlight key differences in the forces shaping their genomes, including differences in mutational and selective pressures13,14.
Literally, comparative genomics allows one to link laboratory notebooks of clinical and basic researchers. With knowledge of both genomes, biomedical studies of human genes can be complemented by experimental manipulations of corresponding mouse genes to accelerate functional understanding. In this respect, the mouse is unsurpassed as a model system for probing mammalian biology and human disease15,16. Its unique advantages include a century of genetic studies, scores of inbred strains, hundreds of spontaneous mutations, practical techniques for random mutagenesis, and, importantly, directed engineering of the genome through transgenic, knockout and knockin techniques17,18,19,20,21,22.
For these and other reasons, the Human Genome Project (HGP) recognized from its outset that the sequencing of the human genome needed to be followed as rapidly as possible by the sequencing of the mouse genome. In early 2001, the International Human Genome Sequencing Consortium reported a draft sequence covering about 90% of the euchromatic human genome, with about 35% in finished form1. Since then, progress towards a complete human sequence has proceeded swiftly, with approximately 98% of the genome now available in draft form and about 95% in finished form.
Here, we report the results of an international collaboration involving centres in the United States and the United Kingdom to produce a high-quality draft sequence of the mouse genome and a broad scientific network to analyse the data. The draft sequence was generated by assembling about sevenfold sequence coverage from female mice of the C57BL/6J strain (referred to below as B6). The assembly contains about 96% of the sequence of the euchromatic genome (excluding chromosome Y) in sequence contigs linked together into large units, usually larger than 50 megabases (Mb).
With the availability of a draft sequence of the mouse genome, we have undertaken an initial comparative analysis to examine the similarities and differences between the human and mouse genomes. Some of the important points are listed below.
• The mouse genome is about 14% smaller than the human genome (2.5 Gb compared with 2.9 Gb). The difference probably reflects a higher rate of deletion in the mouse lineage.
• Over 90% of the mouse and human genomes can be partitioned into corresponding regions of conserved synteny, reflecting segments in which the gene order in the most recent common ancestor has been conserved in both species.
• At the nucleotide level, approximately 40% of the human genome can be aligned to the mouse genome. These sequences seem to represent most of the orthologous sequences that remain in both lineages from the common ancestor, with the rest likely to have been deleted in one or both genomes.
• The neutral substitution rate has been roughly half a nucleotide substitution per site since the divergence of the species, with about twice as many of these substitutions having occurred in the mouse compared with the human lineage.
• By comparing the extent of genome-wide sequence conservation to the neutral rate, the proportion of small (50–100 bp) segments in the mammalian genome that is under (purifying) selection can be estimated to be about 5%. This proportion is much higher than can be explained by protein-coding sequences alone, implying that the genome contains many additional features (such as untranslated regions, regulatory elements, non-protein-coding genes, and chromosomal structural elements) under selection for biological function.
• The mammalian genome is evolving in a non-uniform manner, with various measures of divergence showing substantial variation across the genome.
• The mouse and human genomes each seem to contain about 30,000 protein-coding genes. These refined estimates have been derived from both new evidence-based analyses that produce larger and more complete sets of gene predictions, and new de novo gene predictions that do not rely on previous evidence of transcription or homology. The proportion of mouse genes with a single identifiable orthologue in the human genome seems to be approximately 80%. The proportion of mouse genes without any homologue currently detectable in the human genome (and vice versa) seems to be less than 1%.
• Dozens of local gene family expansions have occurred in the mouse lineage. Most of these seem to involve genes related to reproduction, immunity and olfaction, suggesting that these physiological systems have been the focus of extensive lineage-specific innovation in rodents.
• Mouse–human sequence comparisons allow an estimate of the rate of protein evolution in mammals. Certain classes of secreted proteins implicated in reproduction, host defence and immune response seem to be under positive selection, which drives rapid evolution.
• Despite marked differences in the activity of transposable elements between mouse and human, similar types of repeat sequences have accumulated in the corresponding genomic regions in both species. The correlation is stronger than can be explained simply by local (G+C) content and points to additional factors influencing how the genome is moulded by transposons.
• By additional sequencing in other mouse strains, we have identified about 80,000 single nucleotide polymorphisms (SNPs). The distribution of SNPs reveals that genetic variation among mouse strains occurs in large blocks, mostly reflecting contributions of the two subspecies Mus musculus domesticus and Mus musculus musculus to current laboratory strains.
The mouse genome sequence is freely available in public databases (GenBank accession number CAAA01000000) and is accessible through various genome browsers (http://www.ensembl.org/Mus_musculus/, http://genome.ucsc.edu/ and http://www.ncbi.nlm.nih.gov/genome/guide/mouse/).
In this paper, we begin with information about the generation, assembly and evaluation of the draft genome sequence, the conservation of synteny between the mouse and human genomes, and the landscape of the mouse genome. We then explore the repeat sequences, genes and proteome of the mouse, emphasizing comparisons with the human. This is followed by evolutionary analysis of selection and mutation in the mouse and human lineages, as well as polymorphism among current mouse strains. A full and detailed description of the methods underlying these studies is provided as Supplementary Information. In many respects, the current paper is a companion to the recent paper on the human genome sequence1. Extensive background information about many of the topics discussed below is provided there.
Background to the mouse genome sequencing project
Origins of the mouse
The precise origin of the mouse and human lineages has been the subject of recent debate. Palaeontological evidence has long indicated a great radiation of placental (eutherian) mammals about 65 million years ago (Myr) that filled the ecological space left by the extinction of the dinosaurs, and that gave rise to most of the eutherian orders23. Molecular phylogenetic analyses indicate earlier divergence times of many of the mammalian clades. Some of these studies have suggested a very early date for the divergence of mouse from other mammals (100–130 Myr23,24,25) but these estimates partially originate from the fast molecular clock in rodents (see below). Recent molecular studies that are less sensitive to the differences in evolutionary rates have suggested that the eutherian mammalian radiation took place throughout the Late Cretaceous period (65–100 Myr), but that rodents and primates actually represent relatively late-branching lineages26,27. In the analyses below, we use a divergence time for the human and mouse lineages of 75 Myr for the purpose of calculating evolutionary rates, although it is possible that the actual time may be as recent as 65 Myr.
Origins of mouse genetics
The origin of the mouse as the leading model system for biomedical research traces back to the start of human civilization, when mice became commensal with human settlements. Humans noticed spontaneously arising coat-colour mutants and recorded their observations for millennia (including ancient Chinese references to dominant-spotting, waltzing, albino and yellow mice). By the 1700s, mouse fanciers in Japan and China had domesticated many varieties as pets, and Europeans subsequently imported favourites and bred them to local mice (thereby creating progenitors of modern laboratory mice as hybrids among M. m. domesticus, M. m. musculus and other subspecies). In Victorian England, ‘fancy’ mice were prized and traded, and a National Mouse Club was founded in 1895 (refs 28, 29).
With the rediscovery of Mendel's laws of inheritance in 1900, pioneers of the new science of genetics (such as Cuenot, Castle and Little) were quick to recognize that the discontinuous variation of fancy mice was analogous to that of Mendel's peas, and they set out to test the new theories of inheritance in mice. Mating programmes were soon established to create inbred strains, resulting in many of the modern, well-known strains (including C57BL/6J)30.
Genetic mapping in the mouse began with Haldane's report31 in 1915 of linkage between the pink-eye dilution and albino loci on the linkage group that was eventually assigned to mouse chromosome 7, just 2 years after the first report of genetic linkage in Drosophila. The genetic map grew slowly over the next 50 years as new loci and linkage groups were added—chromosome 7 grew to three loci by 1935 and eight by 1954. The accumulation of serological and enzyme polymorphisms from the 1960s to the early 1980s began to fill out the genome, with the map of chromosome 7 harbouring 45 loci by 1982 (refs 29, 31).
The real explosion, however, came with the development of recombinant DNA technology and the advent of DNA-sequence-based polymorphisms. Initially, this involved the detection of restriction-fragment length polymorphisms (RFLPs)32; later, the emphasis shifted to the use of simple sequence length polymorphisms (SSLPs; also called microsatellites), which could be assayed easily by polymerase chain reaction (PCR)33,34,35,36 and readily revealed polymorphisms between inbred laboratory strains.
Origins of mouse genomics
When the Human Genome Project (HGP) was launched in 1990, it included the mouse as one of its five central model organisms, and targeted the creation of genetic, physical and eventually sequence maps of the mouse genome.
By 1996, a dense genetic map with nearly 6,600 highly polymorphic SSLP markers ordered in a common cross had been developed34, providing the standard tool for mouse genetics. Subsequent efforts filled out the map to over 12,000 polymorphic markers, although not all of these loci have been positioned precisely relative to one another. With these and other loci, Haldane's original two-marker linkage group on chromosome 7 had now swelled to about 2,250 loci.
Physical maps of the mouse genome also proceeded apace, using sequence-tagged sites (STS) together with radiation-hybrid panels37,38 and yeast artificial chromosome (YAC) libraries to construct dense landmark maps39. Together, the genetic and physical maps provide thousands of anchor points that can be used to tie clones or DNA sequences to specific locations in the mouse genome.
Other resources included large collections of expressed-sequence tags (EST)40, a growing number of full-length complementary DNAs41,42 and excellent bacterial artificial chromosome (BAC) libraries43. The latter have been used for deriving large sets of BAC-end sequences37 and, as part of this collaboration, to generate a fingerprint-based physical map44. Furthermore, key mouse genome databases were developed at the Jackson (http://www.informatics.jax.org/), Harwell (http://www.har.mrc.ac.uk/) and RIKEN (http://genome.rtc.riken.go.jp/) laboratories to provide the community with access to this information.
With these resources, it became straightforward (but not always easy) to perform positional cloning of classic single-gene mutations for visible, behavioural, immunological and other phenotypes. Many of these mutations provide important models of human disease, sometimes recapitulating human phenotypes with uncanny accuracy. It also became possible for the first time to begin dissecting polygenic traits by genetic mapping of quantitative trait loci (QTL) for such traits.
Continuing advances fuelled a growing desire for a complete sequence of the mouse genome. The development of improved random mutagenesis protocols led to the establishment of large-scale screens to identify interesting new mutants, increasing the need for more rapid positional cloning strategies. QTL mapping experiments succeeded in localizing more than 1,000 loci affecting physiological traits, creating demand for efficient techniques capable of trawling through large genomic regions to find the underlying genes. Furthermore, the ability to perform directed mutagenesis of the mouse germ line through homologous recombination made it possible to manipulate any gene given its DNA sequence, placing an increasing premium on sequence information. In all of these cases, it was clear that genome sequence information could markedly accelerate progress.
Origin of the Mouse Genome Sequencing Consortium
With the sequencing of the human genome well underway by 1999, a concerted effort to sequence the entire mouse genome was organized by a Mouse Genome Sequencing Consortium (MGSC). The MGSC originally consisted of three large sequencing centres—the Whitehead/Massachusetts Institute of Technology (MIT) Center for Genome Research, the Washington University Genome Sequencing Center, and the Wellcome Trust Sanger Institute—together with an international database, Ensembl, a joint project between the European Bioinformatics Institute and the Sanger Institute.
In addition to the genome-wide efforts of the MGSC, other publicly funded groups have been contributing to the sequencing of the mouse genome in specific regions of biological interest. Together, the MGSC and these programmes have so far yielded clone-based draft sequence consisting of 1,859 Mb (74%, although there is redundancy) and finished sequence of 477 Mb (19%) of the mouse genome. Furthermore, Mural and colleagues45 recently reported a draft sequence of mouse chromosome 16 containing 87 Mb (3.5%).
To analyse the data reported here, the MGSC was expanded to include the other publicly funded sequencing groups and a Mouse Genome Analysis Group consisting of scientists from 27 institutions in 6 countries.
Generating the draft genome sequence
Sequencing strategy
Sanger and co-workers developed the strategy of random shotgun sequencing in the early 1980s, and it has remained the mainstay of genome sequencing over the ensuing two decades. The approach involves producing random sequence ‘reads’, generating a preliminary assembly on the basis of sequence overlaps, and then performing directed sequencing to obtain a ‘finished’ sequence with gaps closed and ambiguities resolved46. Ansorge and colleagues47 extended the technique by the use of ‘paired-end sequencing’, in which sequencing is performed from both ends of a cloned insert to obtain linking information, which is then used in sequence assembly. More recently, Myers and co-workers48, and others, have developed efficient algorithms for exploiting such linking information.
A principal issue in the sequencing of large, complex genomes has been whether to perform shotgun sequencing on the entire genome at once (whole-genome shotgun, WGS) or to first break the genome into overlapping large-insert clones and to perform shotgun sequencing on these intermediates (hierarchical shotgun)46. The WGS technique has the advantage of simplicity and rapid early coverage; it readily works for simple genomes with few repeats, but there can be difficulties encountered with genomes that contain highly repetitive sequences (such as the human genome, which has near-perfect repeats spanning hundreds of kilobases). Hierarchical shotgun sequencing overcomes such difficulties by using local assembly, thus decreasing the number of repeat copies in each assembly and allowing comparison of large regions of overlaps between clones. Consequently, efforts to produce finished sequences of complex genomes have relied on either pure hierarchical shotgun sequencing (including those of Caenorhabditis elegans49, Arabidopsis thaliana49 and human1) or a combination of WGS and hierarchical shotgun sequencing (including those of Drosophila melanogaster50, human2 and rice51).
The ultimate aim of the MGSC is to produce a finished, richly annotated sequence of the mouse genome to serve as a permanent reference for mammalian biology. In addition, we wished to produce a draft sequence as rapidly as possible to aid in the interpretation of the human genome sequence and to provide a useful intermediate resource to the research community. Accordingly, we adopted a hybrid strategy for sequencing the mouse genome. The strategy has four components: (1) production of a BAC-based physical map of the mouse genome by fingerprinting and sequencing the ends of clones of a BAC library44; (2) WGS sequencing to approximately sevenfold coverage and assembly to generate an initial draft genome sequence; (3) hierarchical shotgun sequencing of BAC clones covering the mouse genome combined with the WGS data to create a hybrid WGS-BAC assembly; and (4) production of a finished sequence by using the BAC clones as a template for directed finishing. This mixed strategy was designed to exploit the simpler organizational aspects of WGS assemblies in the initial phase, while still culminating in the complete high-quality sequence afforded by clone-based maps.
We chose to sequence DNA from a single mouse strain, rather than from a mixture of strains45, to generate a solid reference foundation, reasoning that polymorphic variation in other strains could be added subsequently (see below). After extensive consultation with the scientific community52, the B6 strain was selected because of its principal role in mouse genetics, including its well-characterized phenotype and role as the background strain on which many important mutations arose. We elected to sequence a female mouse to obtain equal coverage of chromosome X and autosomes. Chromosome Y was thus omitted, but this chromosome is highly repetitive (the human chromosome Y has multiple duplicated regions exceeding 100 kb in size with 99.9% sequence identity53) and seemed an unwise target for the WGS approach. Instead, mouse chromosome Y is being sequenced by a purely clone-based (hierarchical shotgun) approach.
Sequencing and assembly
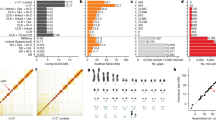
The genome assembly was based on a total of 41.4 million sequence reads derived from both ends of inserts (paired-end reads) of various clone types prepared from B6 female DNA. The inserts ranged in size from 2 to 200 kb (Table 1). The three large MGSC sequencing centres generated 40.4 million reads, and 0.6 million reads were generated at the University of Utah. In addition, we used 0.4 million reads from both ends of BAC inserts reported by The Institute for Genome Research54.
A total of 33.6 million reads passed extensive checks for quality and source, of which 29.7 million were paired; that is, derived from opposite ends of the same clone (Table 1). The assembled reads represent approximately 7.7-fold sequence coverage of the euchromatic mouse genome (6.5-fold coverage in bases with a Phred quality score of >20)55. Together, the clone inserts provide roughly 47-fold physical coverage of the genome.
The sequence reads, together with the pairing information, were used as input for two recently developed sequence-assembly programs, Arachne56,57 and Phusion58. No mapping information and no clone-based sequences were used in the WGS assembly, with the exception of a few reads (<0.1% of the total) derived from a handful of BACs, which were used as internal controls. The assembly programs were tested and compared on intermediate data sets over the course of the project and were thereby refined. The programs produced comparable outputs in the final assembly. The assembly generated by Arachne was chosen as the draft sequence described here because it yielded greater short-range and long-range continuity with comparable accuracy.
The assembly contains 224,713 sequence contigs, which are connected by at least two read-pair links into supercontigs (or scaffolds). There are a total of 7,418 supercontigs at least 2 kb in length, plus a further 37,125 smaller supercontigs representing <1% of the assembly. The contigs have an N50 length of 24.8 kb, whereas the supercontigs have an N50 length that is approximately 700-fold larger at 16.9 Mb (N50 length is the size x such that 50% of the assembly is in units of length at least x). In fact, most of the genome lies in supercontigs that are extremely large: the 200 largest supercontigs span more than 98% of the assembled sequence, of which 3% is within sequence gaps (Table 2).
Anchoring to chromosomes
We assigned as many supercontigs as possible to chromosomal locations in the proper order and orientation. Supercontigs were localized largely by sequence alignments with the extensively validated mouse genetic map34, with some additional localization provided by the mouse radiation-hybrid map37 and the BAC map44. We found no evidence of incorrect global joins within the supercontigs (that is, multiple markers supporting two discordant locations within the genome), and thus were able to place them directly. Altogether, we placed 377 supercontigs, including all supercontigs >500 kb in length.
Once much of the sequence was anchored, it was possible to exploit additional read-pair and physical mapping information to obtain greater continuity (Table 2). For example, some adjacent supercontigs were connected by BAC-end (or other) links, satisfying appropriate length and orientation constraints, including single links. Furthermore, some adjacent extended supercontigs were connected by means of fingerprint contigs in the BAC-based physical map. These additional links were used to join sequences into ultracontigs. In the end, a total of 88 ultracontigs with an N50 length of 50.6 Mb (exclusive of gaps) contained 95.7% of the assembled sequence (Fig. 1). Continuity near telomeres tends to be lower, and two chromosomes (5 and X) have unusually large numbers of ultracontigs.
The position and extent of the 88 ultracontigs of the MGSCv3 assembly are shown adjacent to ideograms of the mouse chromosomes. All mouse chromosomes are acrocentric, with the centromeric end at the top of each chromosome. The supercontigs of the sequence assembly were anchored to the mouse chromosomes using the MIT genetic map. Neighbouring supercontigs were linked together into ultracontigs using information from single BAC links and the fingerprint and radiation-hybrid maps, resulting in 88 ultracontigs containing 95% of the bases in the euchromatic genome.
Proportion of genome contained in the assembly
This was assessed by comparison with publicly available finished genome sequence and mouse cDNA sequences. Of the 187 Mb of finished mouse sequence, 96% was contained in the anchored assembly. This finished sequence, however, is not a completely random cross-section of the genome (it has been cloned as BACs, finished, and in some cases selected on the basis of its gene content). Of 11,452 cDNA sequences from the curated RefSeq collection, 99.3% of the cDNAs could be aligned to the genome sequence (see Supplementary Information). These alignments contained 96.4% of the cDNA bases. Together, this indicates that the draft genome sequence includes approximately 96% of the euchromatic portion of the mouse genome, with about 95% anchored (Table 1).
Genome size
On the basis of the estimated sizes of the ultracontigs and gaps between them, the total length of the euchromatic mouse genome was estimated to be about 2.5 Gb (see Supplementary Information), or about 14% smaller than that of the euchromatic human genome (about 2.9 Gb) (Table 3). The ultracontigs include spanned gaps, whose lengths are estimated on the basis of paired-end reads and alignment against the human sequence (see below). To test the accuracy of the ultracontig lengths, we compared the actual length of 675 finished mouse BAC sequences (from the B6 strain) with the corresponding estimated length from the draft genome sequence. The ratio of estimated length to actual length had a median value of 0.9994, with 68% of cases falling within 0.99–1.01 and 84% of cases within 0.98–1.02.
Quality assessment at intermediate scale
Although no evidence of large-scale misassembly was found when anchoring the assembly onto the mouse chromosomes, we examined the assembly for smaller errors.
To assess the accuracy at an intermediate scale, we compared the positions of well-studied markers on the mouse genetic map and in the genome assembly (see Supplementary Information). Out of 2,605 genetic markers that were unambiguously mapped to the sequence assembly (BLAST match using 10-100 or better as an E-value to a single location) we found 1.8% in which the chromosomal assignment in the genetic map conflicted with that in the sequence. This is well within the known range of erroneous assignments within the genetic map34. We tested 11 such discrepant markers by re-mapping them in a mouse cross. In ten cases, the data showed that the previous genetic map assignment was erroneous and supported the position in the draft sequence. In one case, the data supported the previous genetic map assignment and contradicted the assembly. By studying the one erroneous case, we recognized that a single 36-kb segment had been erroneously merged into a sequence contig by means of a single overlap of two reads. We screened the entire assembly for similar instances, affecting regions of at least 20 kb. Only 17 additional cases were found, with a median size of the incorrectly merged segment of 34 kb. These are being corrected in the next release of the MGSC sequence. We are continuing to investigate instances involving smaller incorrectly merged segments.
We also found 19 instances (0.7%) of conflicts in local marker order between the genetic map and sequence assembly. A conflict was defined as any instance that would require changing more than a single genotype in the data underlying the genetic map to resolve. We studied ten cases by re-mapping the genetic markers, and eight were found to be due to errors in the genetic map. On the basis of this analysis, we estimate that chromosomal misassignment and local misordering affects <0.3% of the assembled sequence.
Quality assessment at fine scale
We also assessed fine-scale accuracy of the assembly by carefully aligning it to about 10 Mb of finished BAC-derived sequence from the B6 strain. This revealed a total of 39 discrepancies of ≥50 bp in length (median size of 320 bp), reflecting small misassemblies either in the draft sequence or the finished BAC sequences. These discrepancies typically occurred at the ends of contigs in the WGS assembly, indicating that they may represent the incorrect incorporation of a single terminal read.
At the single nucleotide level in the assembly, the observed discrepancy rates varied in a manner consistent with the quality scores assigned to the bases in the WGS assembly (see Supplementary Information). Overall, 96% of nucleotides in the assembly have Arachne quality scores ≥40, corresponding to a predicted error rate of 1 per 10,000 bases. Such bases had an observed discrepancy rate against finished sequence of 0.005%, or 5 errors per 100,000 bases.
Comparison with the draft sequence of chromosome 16
We also compared the sequence reported here to a draft sequence of mouse chromosome 16 recently published by Mural and co-workers45. Because the latter was produced from strain 129 and other mouse strains, it is expected to differ slightly at the nucleotide level but should otherwise show good agreement. The sequences align well at large scales (hundreds of kilobases), although the assembly by Mural and co-workers contains less total sequence (87 compared with 91 Mb) and includes a region of approximately 300 kb that we place on chromosome X. There were differences at intermediate scales, with our draft sequence showing better agreement with finished BAC-derived sequences (approximately fourfold fewer discrepancies of length ≥500 bp; 20 compared with 5 in about 2.8 Mb of finished sequence). These could not be explained by strain differences, as similar results were seen with finished sequence from the B6 and 129 strains.
Collapse of duplicated regions
The human genome contains many large duplicated regions, estimated to comprise roughly 5% of the genome59, with nearly identical sequence. If such regions are also common in the mouse genome, they might collapse into a single copy in the WGS assembly. Such artefactual collapse could be detected as regions with unusually high read coverage, compared with the average depth of 7.4-fold in long assembled contigs. We searched for contigs that were >20 kb in size and contained >10 kb of sequence in which the read coverage was at least twofold higher than the average. Such regions comprised only a tiny fraction (<0.0001) of the total assembly, of which only half had been anchored to a chromosome. None of these windows had coverage exceeding the average by more than threefold. This may indicate that the mouse genome contains fewer large regions of near-exact duplication than the human. Alternatively, regions of near-exact duplication may have been systematically excluded by the WGS assembly programme. This issue is better addressed through hierarchical shotgun than WGS sequencing and will be examined more carefully in the course of producing a finished mouse genome sequence.
Unplaced reads and large tandem repeats
We expected that highly repetitive regions of the genome would not be assembled or would not be anchored on the chromosomes. Indeed, 5.9 million of the 33.6 million passing reads were not part of anchored sequence, with 88% of these not assembled into sequence contigs and 12% assembled into small contigs but not chromosomally localized.
A striking example of unassembled sequence is a large region on mouse chromosome 1 that contains a tandem expansion of sequence containing the Sp100-rs gene fusion. This region is highly variable among mouse species and even laboratory strains, with estimated lengths ranging from 6 to 200 Mb60,61. The bulk of this region was not reliably assembled in the draft genome sequence. The individual sequence reads together were found to contain 493-fold coverage of the Sp100-rs gene, suggesting that there are roughly 60 copies in the B6 genome (corresponding to a region of about 6 Mb). This is consistent with an estimate of 50 copies in B6 obtained by Southern blotting62.
We also examined centromeric sequences, including the euchromatin-proximal major satellite repeat (234 bases) and the telomere-proximal minor repeat (120 bases) found on some chromosomes63,64. (Note that mouse chromosomes are all acrocentric, meaning that the centromere is adjacent to one telomere.) The minor satellite was poorly represented among the sequence reads (present in about 24,000 reads or <0.1% of the total) suggesting that this satellite sequence is difficult to isolate in the cloning systems used. The major satellite was found in about 3.6% of the reads; this is also lower than previous estimates based on density gradient experiments, which found that major satellites comprise about 5.5% of the mouse genome, or approximately 8 Mb per chromosome65.
Evaluation of WGS assembly strategy
The WGS assembly described here involved only random reads, without any additional map-based information. By many criteria, the assembly is of very high quality. The N50 supercontig size of 16.9 Mb far exceeds that achieved by any previous WGS assembly, and the agreement with genome-wide maps is excellent. The assembly quality may be due to several factors, including the use of high-quality libraries, the variety of insert lengths in multiple libraries, the improved assembly algorithms, and the inbred nature of the mouse strain (in contrast to the polymorphisms in the human genome sequences). Another contributing factor may be that the mouse differs from the human in having less recent segmental duplication to confound assembly.
Notwithstanding the high quality of the draft genome sequence, we are mindful that it contains many gaps, small misassemblies and nucleotide errors. It is likely that these could not all be resolved by further WGS sequencing, therefore directed sequencing will be needed to produce a finished sequence. The results also suggest that WGS sequencing may suffice for large genomes for which only draft sequence is required, provided that they contain minimal amounts of sequence associated with recent segmental duplications or large, recent interspersed repeat elements.
Adding finished sequence
As a final step, we enhanced the WGS sequence assembly by substituting available finished BAC-derived sequence from the B6 strain. In total, we replaced 3,528 draft sequence contigs with 48.2 Mb of finished sequence from 210 finished BACs available at the time of the assembly. The resulting draft genome sequence, MGSCv3, was submitted to the public databases and is freely available in electronic form through various sources (see below).
The sequence data and assemblies have been freely available throughout the course of the project. The next step of the project, which is already underway, is to convert the draft sequence into a finished sequence. As the MGSC produces additional BAC assemblies and finished sequence, we plan to continue to revise and release enhanced versions of the genome sequence en route to a completely finished sequence66, thereby providing a permanent foundation for biomedical research in the twenty-first century.
Conservation of synteny between mouse and human genomes
With the draft sequence in hand, we began our analysis by investigating the strong conservation of synteny between the mouse and human genomes. Beyond providing insight into evolutionary events that have moulded the chromosomes, this analysis facilitates further comparisons between the genomes.
Starting from a common ancestral genome approximately 75 Myr, the mouse and human genomes have each been shuffled by chromosomal rearrangements. The rate of these changes, however, is low enough that local gene order remains largely intact. It is thus possible to recognize syntenic (literally ‘same thread’) regions in the two species that have descended relatively intact from the common ancestor.
The earliest indication that genes reside in similar relative positions in different mammalian species traces to the observation that the albino and pink-eye dilution mutants are genetically closely linked in both mouse and rat67,68. Significant experimental evidence came from genetic studies of somatic cells69. In 1984, Nadeau and Taylor70 used mouse linkage data and human cytogenetic data to compare the chromosomal locations of orthologous genes. On the basis of a small data set (83 loci), they extrapolated that the mouse and human genomes could be parsed into roughly 180 syntenic regions. During two decades of subsequent work, the density of the synteny map has been increased, but the estimated number of syntenic regions has remained close to the original projection. A recent gene-based synteny map37 used more than 3,600 orthologous loci to define about 200 regions of conserved synteny. However, it is recognized that such maps might still miss regions owing to insufficient marker density.
With a robust draft sequence of the mouse genome and >90% finished sequence of the human genome in hand, it is possible to undertake a more comprehensive analysis of conserved synteny. Rather than simply relying on known human–mouse gene pairs, we identified a much larger set of orthologous landmarks as follows. We performed sequence comparisons of the entire mouse and human genome sequences using the PatternHunter program71 to identify regions having a similarity score exceeding a high threshold (>40, corresponding to a minimum of a 40-base perfect match, with penalties for mismatches and gaps), with the additional property that each sequence is the other's unique match above this threshold. Such regions probably reflect orthologous sequence pairs, derived from the same ancestral sequence.
About 558,000 orthologous landmarks were identified; in the mouse assembly, these sequences have a mean spacing of about 4.4 kb and an N50 length of about 500 bp. The landmarks had a total length of roughly 188 Mb, comprising about 7.5% of the mouse genome. It should be emphasized that the landmarks represent only a small subset of the sequences, consisting of those that can be aligned with the highest similarity between the mouse and human genomes. (Indeed, below we show that about 40% of the human genome can be aligned confidently with the mouse genome.)
The locations of the landmarks in the two genomes were then compared to identify regions of conserved synteny. We define a syntenic segment to be a maximal region in which a series of landmarks occur in the same order on a single chromosome in both species. A syntenic block in turn is one or more syntenic segments that are all adjacent on the same chromosome in human and on the same chromosome in mouse, but which may otherwise be shuffled with respect to order and orientation. To avoid small artefactual syntenic segments owing to imperfections in the two draft genome sequences, we only considered regions above 300 kb and ignored occasional isolated interruptions in conserved order (see Supplementary Information). Thus, some small syntenic segments have probably been omitted—this issue will be addressed best when finished sequences of the two genomes are completed.
Marked conservation of landmark order was found across most of the two genomes (Fig. 2). Each genome could be parsed into a total of 342 conserved syntenic segments. On average, each landmark resides in a segment containing 1,600 other landmarks. The segments vary greatly in length, from 303 kb to 64.9 Mb, with a mean of 6.9 Mb and an N50 length of 16.1 Mb. In total, about 90.2% of the human genome and 93.3% of the mouse genome unambiguously reside within conserved syntenic segments. The segments can be aggregated into a total of 217 conserved syntenic blocks, with an N50 length of 23.2 Mb.
We detected 558,000 highly conserved, reciprocally unique landmarks within the mouse and human genomes, which can be joined into conserved syntenic segments and blocks (defined in text). A typical 510-kb segment of mouse chromosome 12 that shares common ancestry with a 600-kb section of human chromosome 14 is shown. Blue lines connect the reciprocal unique matches in the two genomes. The cyan bars represent sequence coverage in each of the two genomes for the regions. In general, the landmarks in the mouse genome are more closely spaced, reflecting the 14% smaller overall genome size.
The nature and extent of conservation of synteny differs substantially among chromosomes (Fig. 3 and Table 4). In accordance with expectation, the X chromosomes are represented as single, reciprocal syntenic blocks72. Human chromosome 20 corresponds entirely to a portion of mouse chromosome 2, with nearly perfect conservation of order along almost the entire length, disrupted only by a small central segment (Fig. 4a, d). Human chromosome 17 corresponds entirely to a portion of mouse chromosome 11, but extensive rearrangements have divided it into at least 16 segments (Fig. 4b, e). Other chromosomes, however, show evidence of much more extensive interchromosomal rearrangement than these cases (Fig. 4c, f).
For each of three human (a–c) and mouse (d–f) chromosomes, the positions of orthologous landmarks are plotted along the x axis and the corresponding position of the landmark on chromosomes in the other genome is plotted on the y axis. Different chromosomes in the corresponding genome are differentiated with distinct colours. In a remarkable example of conserved synteny, human chromosome 20 (a) consists of just three segments from mouse chromosome 2 (d), with only one small segment altered in order. Human chromosome 17 (b) also shares segments with only one mouse chromosome (11) (e), but the 16 segments are extensively rearranged. However, most of the mouse and human chromosomes consist of multiple segments from multiple chromosomes, as shown for human chromosome 2 (c) and mouse chromosome 12 (f). Circled areas and arrows denote matching segments in mouse and human.
We compared the new sequence-based map of conserved synteny with the most recent previous map based on 3,600 loci30. The new map reveals many more conserved syntenic segments (342 compared with 202) but only slightly more conserved syntenic blocks (217 compared with 170). Most of the conserved syntenic blocks had previously been recognized and are consistent with the new map, but many rearrangements of segments within blocks had been missed (notably on the X chromosome).
The occurrence of many local rearrangements is not surprising. Compared with interchromosomal rearrangements (for example, translocations), paracentric inversions (that is, those within a single chromosome and not including the centromere) carry a lower selective disadvantage in terms of the frequency of aneuploidy among offspring. These are also seen at a higher frequency in genera such as Drosophila, in which extensive cytogenetic comparisons have been carried out73,74.
The block and segment sizes are broadly consistent with the random breakage model of genome evolution75 (Fig. 5). At this gross level, there is no evidence of extensive selection for gene order across the genome. Selection in specific regions, however, is by no means excluded, and indeed seems probable (for example, for the major histocompatibility complex). Moreover, the analysis does not exclude the possibility that chromosomal breaks may tend to occur with higher frequency in some locations.
a, b, The number of segments (a) and blocks (b) with synteny conserved between mouse and human in 5-Mb bins (starting with 0.3–5 Mb) is plotted on a logarithmic scale. The dots indicate the expected values for the exponential curve of random breakage given the number of blocks and segments, respectively.
With a map of conserved syntenic segments between the human and mouse genomes, it is possible to calculate the minimal number of rearrangements needed to ‘transform’ one genome into the other70,76,77. When applied to the 342 syntenic segments above, the most parsimonious path has 295 rearrangements. The analysis suggests that chromosomal breaks may have a tendency to reoccur in certain regions. With only two species, however, it is not yet possible to recover the ancestral chromosomal order or reconstruct the precise pathway of rearrangements. As more mammalian species are sequenced, it should be possible to draw such inferences and study the nature of chromosome rearrangement.
Genome landscape
We next sought to analyse the contents of the mouse genome, both in its own right and in comparison with corresponding regions of the human genome. The poster included with this issue provides a high-level view of the mouse genome, showing such features as genes and gene predictions, repetitive sequence content, (G+C) content, synteny with the human genome, and mouse QTLs.
All of the mouse genome information is accessible in electronic form through various browsers: Ensembl (http://www.ensembl.org), the University of California at Santa Cruz (http://genome.ucsc.edu) and the National Center for Biotechnology Information (http://www.ncbi.nlm.nih.gov). These browsers allow users to scroll along the chromosomes and zoom in or out to any scale, as well as to display information at any desired level of detail. The mouse genome information has also been integrated into existing human genome browsers at these same organizations. In this section, we compare general properties of the mouse and human genomes.
Genome expansion and contraction
The projected total length of the euchromatic portion of the mouse genome (2.5 Gb) is about 14% smaller than that of the human genome (2.9 Gb). To investigate the source of this difference, we examined the relative size of intervals between consecutive orthologous landmarks in the human and mouse genomes. The mouse/human ratio has a mean at 0.91 for autosomes, but varies widely, with the mouse interval being larger than the human in 38% of cases (Fig. 6). Chromosome X, by contrast, shows no net relative expansion or contraction, with a mouse/human ratio of 1.03 (Fig. 6 and Table 4). What accounts for the smaller size of the mouse genome? We address this question below in the sections on repeat sequences and on genome evolution.
For each 100-kb region of the mouse genome, the size ratio to the related segment of the human genome was determined. The frequency of the various ratios is plotted on a logarithmic scale for both the autosomes (blue line) and the X chromosome (red line). The ratio for autosomes shows a mean of 0.91 but the ratio varies widely, with the mouse genome larger for 38% of the intervals. The X chromosome by contrast has a mean ratio of just over 1.0. Indeed, chromosome X is slightly smaller in human.
(G+C) content
The overall distribution of local (G+C) content is significantly different between the mouse and human genomes (Fig. 7). Such differences have been noted in biochemical studies78,79,80,81 and in comparative analyses of fourfold degenerate sites in codons of mouse and human genes82,83,84,85, but the availability of nearly complete genome sequences provides the first detailed picture of the phenomenon.
Mouse has a higher mean (G+C) content than human (42% compared with 41%), but human has a larger fraction of windows with either high or low (G+C) content. The distribution was determined using the unmasked genomes in 20-kb non-overlapping windows, with the fraction of windows (y axis) in each percentage bin (x axis) plotted for both human and mouse.
The mouse has a slightly higher overall (G+C) content than the human (42% compared with 41%), but the distribution is tighter. When local (G+C) content is measured in 20-kb windows across the genome, the human genome has about 1.4% of the windows with (G+C) content >56% and 1.3% with (G+C) content <33%. Such extreme deviations are virtually absent in the mouse genome. The contrast is even seen at the level of entire chromosomes. The human has extreme outliers with respect to (G+C) content (the most extreme being chromosome 19), whereas the mouse chromosomes tend to be far more uniform (Fig. 8).
a, The (G+C) content for each of the mouse chromosomes is relatively similar, whereas human chromosomes show more variation; chromosomes 16, 17, 19 and 22 have higher (G+C) content, and chromosome 13 lower (G+C) content. b, Similarly, the density of CpG islands is relatively homogenous for all mouse chromosomes and more variable in human, with the same exceptions. Note that the mouse and human chromosomes are matched by chromosome number, not by regions of conserved synteny.
There is a strong positive correlation in local (G+C) content between orthologous regions in the mouse and human genomes (Fig. 9), but with the mouse regions showing a clear tendency to be less extreme in (G+C) content than the human regions. This tendency is not uniform, with the most extreme differences seen at the tails of the distribution.
a, Scatter plot of mouse (y axis) compared with human (x axis) (G+C) content for all non-overlapping orthologous 100-kb windows. In general, (G+C) content is correlated between the two species, but very few mouse windows have a (G+C) content over 55%, even where the related human window has over 60% (G+C) content. b, Average mouse (G+C) content of 100-kb syntenic windows binned by human (G+C) content (1% intervals). The red line indicates median values with standard deviation and 5% (green) and 95% (blue) confidence intervals. The black line indicates identical (G+C) content in orthologous segments. c–e, Gene content increases with (G+C) content when comparing (G+C) and gene content in 320-kb non-overlapping, unmasked windows for mouse (blue lines) and human (red lines). c, Cumulative proportions of genes (solid lines) and genome (dashed lines) having (G+C) content below a given level. The tighter distribution of (G+C) content in mouse results in the curve for mouse crossing that for human at 45–46% for both genes and total sequence. The tendency for both genomes to be gene-poor at low (G+C) content and gene-rich at high (G+C) content is shown directly in d, which shows the fraction of genes residing within the portion of the genome having (G+C) content below a given level (for example, the half of the genome with the lowest (G+C) content contains 25% of the genes). e, The average number of genes per window is plotted against the (G+C) content of the window for both genomes, showing that the gene density in mouse reaches the same level as in human but at a lower level of (G+C) content.
In mammalian genomes, there is a positive correlation between gene density and (G+C) content81,86,87,88,89. Given the differences in (G+C) content between human and mouse, we compared the distribution of genes—using the sets of orthologous mouse and human genes described below—with respect to (G+C) content for both genomes (Fig. 9). The density of genes differed markedly when expressed in terms of absolute (G+C) content, but was nearly identical when expressed in terms of percentiles of (G+C) content (Fig. 9). For example, both species have 75–80% of genes residing in the (G+C)-richest half of their genome. Mouse and human thus show similar degrees of homogeneity in the distribution of genes, despite the overall differences in (G+C) content. Notably, the mouse shows similar extremes of gene density despite being less extreme in (G+C) content.
What accounts for the differences in (G+C) content between mouse and human? Does it reflect altered selection for (G+C) content90,91, altered mutational or repair processes92,93,94, or possibly both? Data from additional species will probably be needed to address these issues. Any explanation will need to account for various mysterious phenomena. For example, although overall (G+C) content in mouse is slightly higher than in human (42% compared with 41%), the (G+C) content of chromosome X is slightly lower (39.0% compared with 39.4%). The effect is even more pronounced if one excludes lineage-specific repeats (see below), thereby focusing primarily on shared DNA. In that case, mouse autosomes have an overall (G+C) content that is 1.5% higher than human autosomes (41.2% compared with 39.7%) whereas mouse chromosome X has a (G+C) content that is 1% lower than human chromosome X (37.8% compared with 36.8%).
CpG islands
In mammalian genomes, the palindromic dinucleotide CpG is usually methylated on the cytosine residue. Methyl-CpG is mutated by deamination to TpG, leading to approximately fivefold under-representation of CpG across the human1,95 and mouse genomes. In some regions of the genome that have been implicated in gene regulation, CpG dinucleotides are not methylated and thus are not subject to deamination and mutation. Such regions, termed CpG islands, are usually a few hundred nucleotides in length, have high (G+C) content and above average representation of CpG dinucleotides.
We applied a computer program that attempts to recognize CpG islands on the basis of (G+C) and CpG content of arbitrary lengths of sequence96,97 to the non-repetitive portions of human and mouse genome sequences (see Supplementary Information). The mouse genome contains fewer CpG islands than the human genome (about 15,500 compared with 27,000), which is qualitatively consistent with previous reports98. The absolute number of islands identified depends on the precise definition of a CpG island used, but the ratio between the two species remains fairly constant.
The reason for the smaller number of predicted CpG islands in mouse may relate simply to the smaller fraction of the genome with extremely high (G+C) content99 and its effect on the computer algorithm. Approximately 10,000 of the predicted CpG islands in each species show significant sequence conservation with CpG islands in the orthologous intervals in the other species, falling within the orthologous landmarks described above. Perhaps these represent functional CpG islands, a proposition that can now be tested experimentally84.
Repeats
The single most prevalent feature of mammalian genomes is their repetitive sequences, most of which are interspersed repeats representing ‘fossils’ of transposable elements. Transposable elements are a principal force in reshaping the genome, and their fossils thus provide powerful reporters for measuring evolutionary forces acting on the genome. A recent paper on the human genome sequence1 provided extensive background on mammalian transposons, describing their biology and illustrating many applications to evolutionary studies. Here, we will focus primarily on comparisons between the repeat content of the mouse and human genomes.
Mouse has accumulated more new repetitive sequence than human
Approximately 46% of the human genome can be recognized currently as interspersed repeats resulting from insertions of transposable elements that were active in the last 150–200 million years. The total fraction of the human genome derived from transposons may be considerably larger, but it is not possible to recognize fossils older than a certain age because of the high degree of sequence divergence. Because only 37.5% of the mouse genome is recognized as transposon-derived (Table 5), it is tempting to conclude that the smaller size of the mouse genome is due to lower transposon activity since the divergence of the human and mouse lineages. Closer analysis, however, shows that this is not the case. As we discuss below, transposition has been more active in the mouse lineage. The apparent deficit of transposon-derived sequence in the mouse genome is mostly due to a higher nucleotide substitution rate, which makes it difficult to recognize ancient repeat sequences.
Lineage-specific versus ancestral repeats
Interspersed repeats can be divided into lineage-specific repeats (defined as those introduced by transposition after the divergence of mouse and human) and ancestral repeats (defined as those already present in a common ancestor). Such a division highlights the fact that transposable elements have been more active in the mouse lineage than in the human lineage. Approximately 32.4% of the mouse genome (about 818 Mb) but only 24.4% of the human genome (about 695 Mb) consists of lineage-specific repeats (Table 5). Contrary to initial appearances, transposon insertions have added at least 120 Mb more transposon-derived sequence to the mouse genome than to the human genome since their divergence. This observation is consistent with the previous report that the rate of transposition in the human genome has fallen markedly over the past 40 million years1,100.
The overall lower interspersed repeat density in mouse is the result of an apparent lack of ancestral repeats: they comprise only 5% of the mouse genome compared with 22% of the human genome. The ancestral repeats recognizable in mouse tend to be those of more recent origin, that is, those that originated closest to the mouse–human divergence. This difference may be due partly to a higher deletion rate of non-functional DNA in the mouse lineage, so that more of the older interspersed repeats have been lost. However, the deficit largely reflects a much higher neutral substitution rate in the mouse lineage than in the human lineage, rendering many older ancestral repeats undetectable with available computer programs.
Higher substitution rate in mouse lineage
The hypothesis that the neutral substitution rate is higher in mouse than in human was suggested as early as 1969 (refs 101–103). The idea has continued to be challenged on the basis that the apparent differences may be due to inaccuracies in mammalian phylogenies104,105. The explanation, however, remains unclear, with some attributing it to generation time101,106 and others pointing to a closer correlation with body size107,108.
Ancestral repeats provide a powerful measure of neutral substitution rates, on the basis of comparing thousands of current copies to the inferred consensus sequence of the ancestral element. The large copy number and ubiquitous distribution of ancestral repeats overcome issues of local variation in substitution rates (see below). Most notably, differences in divergence levels are not affected by phylogenetic assumptions, as the time spent by an ancestral repeat family in either lineage is necessarily identical.
The median divergence levels of 18 subfamilies of interspersed repeats that were active shortly before the human–rodent speciation (Table 6) indicates an approximately twofold higher average substitution rate in the mouse lineage than in the human lineage, corresponding closely to an early estimate by Wu and Li109. In human, the least-diverged ancestral repeats have about 16% mismatch to their consensus sequences, which corresponds to approximately 0.17 substitutions per site. In contrast, mouse repeats have diverged by at least 26–27% or about 0.34 substitutions per site, which is about twofold higher than in the human lineage. The total number of substitutions in the two lineages can be estimated at 0.51. Below, we obtain an estimate of a combined rate of 0.46–0.47 substitutions per site, on the basis of an analysis that counts only substitutions since the divergence of the species (see Supplementary Information concerning the methods used).
Assuming a speciation time of 75 Myr, the average substitution rates would have been 2.2 × 10-9 and 4.5 × 10-9 in the human and mouse lineages, respectively. This is in accord with previous estimates of neutral substitution rates in these organisms. (Reports of highly similar substitution rates in human and mouse lineages relied on a much earlier divergence time of rodents from other mammals104.)
Comparison of ancestral repeats to their consensus sequence also allows an estimate of the rate of occurrence of small (<50 bp) insertions and deletions (indels). Both species show a net loss of nucleotides (with deleted bases outnumbering inserted bases by at least 2–3-fold), but the overall loss owing to small indels in ancestral repeats is at least twofold higher in mouse than in human. This may contribute a small amount (1–2%) to the difference in genome size noted above.
It should be noted that the roughly twofold higher substitution rate in mouse represents an average rate since the time of divergence, including an initial period when the two lineages had comparable rates. Comparison with more recent relatives (mouse–rat and human–gibbon, each about 20–25 Myr) indicate that the current substitution rate per year in mouse is probably much higher, perhaps about fivefold higher (see Supplementary Information). Also, note that these estimates refer to substitution rate per year, rather than per generation. Because the human generation time is much longer than that of the mouse (by at least 20-fold), the substitution rate is greater in human than mouse when measured per generation.
Higher substitution rate obscures old repeats
We measured the impact of the higher substitution rate in mouse on the ability to detect ancestral repeats in the mouse genome. By computer simulation, the ability of the RepeatMasker100 program to detect repeats was found to fall off rapidly for divergence levels above about 37%. If we simulate the events in the mouse lineage by adjusting the ancestral repeats in the human genome for the higher substitution levels that would have occurred in the mouse genome, the proportion of the genome that would still be recognizable as ancestral repeats falls to only 6%. This is in close agreement with the proportion actually observed for the mouse. Thus, the current analysis of repeated sequences allows us to see further back into human history (roughly 150–200 Myr) than into mouse history (roughly 100–120 Myr).
A higher rate of interspersed repeat insertion does not explain the larger size of the human genome. Below, we suggest that the explanation lies in a higher rate of large deletions in the mouse lineage.
Comparison of mouse and human repeats
All mammals have essentially the same four classes of transposable elements: (1) the autonomous long interspersed nucleotide element (LINE)-like elements; (2) the LINE-dependent, short RNA-derived short interspersed nucleotide elements (SINEs); (3) retrovirus-like elements with long terminal repeats (LTRs); and (4) DNA transposons. The first three classes procreate by reverse transcription of an RNA intermediate (retroposition), whereas DNA transposons move by a cut-and-paste mechanism of DNA sequence (see refs 1, 100 for further information about these classes).
A comparison of these repeat classes in the mouse and human genomes can be enlightening. On the one hand, differences between the two species reveal the dynamic nature of transposable elements; on the other hand, similarities in the location of lineage-specific elements point to common biological factors that govern insertion and retention of interspersed repeats.
Differences between mouse and human
The most notable difference is in the changing rate of transposition over time: the rate has remained fairly constant in mouse, but markedly increased to a peak at about 40 Myr in human, and then plummeted. This phenomenon was noted in our initial analysis of the human genome; the availability of the mouse genome sequence now confirms and sharpens the observation (Fig. 10). Beyond this overall tendency, there are specific differences in each of the four repeat classes.
This is an update of Fig. 18 in the IHGC human genome paper1. a, b, Distribution for mouse and human of copies of each repeat class in bins corresponding to 1% increments in substitution level calculated using Jukes–Cantor formula (K = -3/4 ln(1 - Drest*4/3)) (see Supplementary Information for definition). The first bin for mouse is artificially low because the WGS assembly used for mouse excludes a larger percentage of very recent repeats. c, d, Interspersed repeats grouped into bins of approximately equal time periods after adjusting for the different rates of substitution in the two genomes. On average, the substitution level has been twofold higher in the mouse than in the human lineage (Table 6), but the difference was initially less and has increased over time. The present rates may differ over fourfold. The activity of transposable elements in the mouse lineage has been quite uniform compared with the human lineage, where an overall decline was interrupted temporarily by a burst of Alu activity. The apparent absence of <2% diverged interspersed repeats in mouse is primarily due to the shotgun sequencing strategy; long, closely similar interspersed repeats very often were not assembled. This is supported by an up to tenfold higher concentration of young L1 and ERV elements at the edges of gaps. The gradually decreasing density of repeats beyond a 30% substitution level reflects in part the limits of the detection method.
The first class that we discuss is LINEs. Copies of LINE1 (L1) form the single largest fraction of interspersed repeat sequence in both human and mouse. No other LINE seems to have been active in either lineage. The extant L1 elements in both species derive from a common ancestor (L1MA6 in Table 6) by means of a series of subfamilies defined primarily by the rapidly evolving 3′ non-coding sequences110. The L1 5′-untranslated regions (UTRs) in both lineages have been even more variable, occasionally through acquisition of entirely new sequences111. Indeed, the three active subfamilies in mouse, which are otherwise >97% identical, have unrelated or highly diverged 5′ ends112,113,114. L1 seems to have remained highly active in mouse, whereas it has declined in the human lineage. Goodier and co-workers113 estimated that the mouse genome contains at least 3,000 potentially active elements (full-length with two intact open reading frames (ORFs)). The current draft sequence of the mouse genome contains only 400 young, full-length elements; of these only 12 have two intact ORFs. This is probably a reflection of the WGS shotgun approach used to assemble the genome. Indeed, most of the young elements in the draft genome sequence are incomplete owing to internal sequence gaps, reflecting the difficulty that WGS assembly has with highly similar repeat sequences. This is a notable limitation of the draft sequence.
The second repeat class is SINEs. Whereas only a single SINE (Alu) was active in the human lineage, the mouse lineage has been exposed to four distinct SINEs (B1, B2, ID, B4). Each is thought to rely on L1 for retroposition, although none share sequence similarity, as is the rule for other LINE–SINE pairs115,116. The mouse B1 and human Alu SINEs are unique among known SINEs in being derived from 7SL RNA; they probably have a common origin117. The mouse B2 is typical among SINEs in having a transfer RNA-derived promoter region. Recent ID elements seem to be derived from a neuronally expressed RNA gene called BC1, which may itself have been recruited from an earlier SINE. This subfamily is minor in mouse, with 2–4,000 copies, but has expanded rapidly in rat where it has produced more than 130,000 copies since the mouse–rat speciation118. Both B2 and ID closely resemble Ala-tRNA, but seem to have independent origins. The B4 family resembles a fusion between B1 and ID119,120. We found that 25% of the 75,000 identified ID elements were located within 50 bp of a B1 element of similar orientation, suggesting that perhaps most older ID elements are mislabelled or truncated B4 SINEs.
More rodent-specific SINEs are present in the mouse genome than Alu SINEs in human (1.4 and 1.1 million, respectively), but they occupy a smaller portion of the genome (7.6% and 10.7%, respectively) because of their smaller sizes. The existence of four families in mouse provides independent opportunities to investigate the properties of SINEs (see below).
The third repeat class is LTR elements. All interspersed LTR-containing elements in mammals are derivatives of the vertebrate-specific retrovirus clade of retrotransposons. The earliest infectious retroviruses probably originated from endogenous retroviral-like (ERV) elements that acquired mechanisms for horizontal transmission121, whereas many current endogenous retroviral elements have probably arisen from infection by retroviruses.
Endogenous retroviruses fall into three classes (I–III), which show a markedly dissimilar evolutionary history in human and mouse (see Fig. 10). Notably, ERVs are nearly extinct in human whereas all three classes have active members in mouse.
Class III accounts for 80% of recognized LTR element copies predating the human–mouse speciation. This class includes the non-autonomous MaLRs: with 388,000 recognizable copies in mouse, it is the single most successful LTR element. It is still active in mouse (represented by MERVL and the MT and ORR1 MaLRs), but died out some 50 Myr in human122.
Copies of class II elements are tenfold denser in mouse than in human. Among the active class II elements in mouse are two abundant and active groups, the intracisternal-A particles (IAP) and the early-transposons (ETn). About 15% of all spontaneous mouse mutants have an allele associated with IAP or ETn insertion, demonstrating the functional consequences of class I element activity in mice. A third active class, the mouse mammary tumour virus, is present in only a few copies123 (see Supplementary Information). In human, there is evidence for at most a few active elements (HERVK10 and HERBK113 (ref. 124)). No class II ERVs are known to predate the human–mouse speciation.
In contrast, class I element copies are fourfold more common in the human than the mouse genome (although it is possible that some have not yet been recognized in mouse). In mouse, this class includes active ERVs, such as the murine leukaemia virus, MuRRS, MuRVY and VL30 (several of which have caused insertional mutations in mouse)—no similar activity is known to exist in human. It is unclear why the class I ERVs have been more successful in the human lineage whereas the class II ERVs have flourished in the mouse lineage.
The fourth repeat class is the DNA transposons. Although most transposable elements have been more active in mouse than human, DNA transposons show the reverse pattern. Only four lineage-specific DNA transposon families could be identified in mouse (the mariner element MMAR1, and the hAT elements URR1, RMER30 and RChar1), compared with 14 in the primate lineage.
For evolutionary survival, DNA transposons are thought to depend on frequent horizontal transfer to new host genomes by means of vectors such as viruses and other intracellular parasites116,125. The mammalian immune system probably forms a large obstacle to the successful invasion of DNA transposons. Perhaps the rodent germ line has been harder to infiltrate by horizontal transfer than the primate genome. Alternatively, it is possible that highly diverged families active in early rodent evolution have not been detected yet. Notably, most copies in the human genome were deposited early in primate evolution.
An interesting case is the mariner element, which seems to have infiltrated independently both the rodent and human genomes. The mariner element is represented by elements (MMAR1 in mouse and HSMAR1 in human) that are 97% identical. The average substitution level outside CpG sites of HSMAR1 is 8% and of MMAR1 is 22%, both well below the divergence of elements predating the human–mouse speciation (Table 6).
Some of the above differences in the nature of interspersed repeats in human and mouse could reflect systematic factors in mouse and human biology, whereas others may represent random fluctuations. Deeper understanding of the biology of transposable elements and detailed knowledge of interspersed repeat populations in other mammals should clarify these issues.
Similar repeats accumulate in orthologous locations
One of the most notable features about repeat elements is the contrast in the genomic distribution of LINEs and SINEs. Whereas LINEs are strongly biased towards (A+T)-rich regions, SINEs are strongly biased towards (G+C)-rich regions. The contrast is all the more notable because both elements are inserted into the genome through the action of the same endonuclease126,127.
Such preferences were studied in detail in the initial analysis of the human genome1, and essentially equivalent preferences are seen in the mouse genome (Fig. 11). With the availability of two mammalian genomes, however, it is possible to extend this analysis to explore whether (A+T) and (G+C) content are truly causative factors or merely reflections of an underlying biological process.
In both species, there is a strong increase in SINE density and a decrease in L1 density with increasing (G+C) content, with the latter particularly marked in the mouse. Another notable contrast is that in mouse, overall interspersed repeat density gradually decreases 2.5-fold with increasing (G+C) content, whereas in human the overall repeat density remains quite uniform. This reflects both the abundance of L1 elements in the mouse (G+C)-poor regions and the unusually high density of Alu in human (G+C)-rich regions.
Towards that end, we studied the insertion of lineage-specific repeat elements in orthologous segments in the human and mouse genomes (Fig. 12). Each insertion represents a new, independent event occurring in one lineage, and thus any correlation between the two species reflects underlying proclivity to insert or retain repeats in particular regions. Visual inspection reveals a strong correlation in the sites of lineage-specific repeats of the various classes (Fig. 12). Lineage-specific repeats also correlate with other genomic features, as discussed in the section on genome evolution.
For each mouse chromosome, its (G+C) content is depicted as a greyscale (centre, right), with darker shades indicating (G+C)-richer regions. Rodent-specific repeats are shown as cumulative histograms (far right), with red, green and blue indicating SINEs, LINEs and other repeats, respectively. The (G+C) content of the orthologous human sequence is similarly shown (centre, left) as well as the primate-specific repeats (far left). Gaps in the human sequence appear opposite those regions of the mouse genome lacking assigned conserved syntenic segments. Note the correlation in (G+C) and repeat content between orthologous regions of the two genomes. Many abrupt shifts in (G+C) content and repeat density are clearly associated with syntenic breaks, which are therefore more likely to be breaks associated with the rodent lineage45.
The correlation of local lineage-specific SINE density is extremely strong (Fig. 13a). Moreover, local SINE density in one species is better predicted by SINE density in the other species than it is by local (G+C) content (Table 7). The local density of each distinct rodent-specific type of SINE is a strong predictor of Alu density at the orthologous locus in human, although the Alu equivalent B1 SINEs show the strongest correlation (r2 = 0.784) (Table 7).
SINE and LINE densities were calculated for 4,126 orthologous pairs with a constant size of 500 kb in mouse. a, b, Strong linear correlation of Alu density in human, and both the Alu-like B1 SINEs (a) and the unrelated B2 SINEs (b) densities in mouse. These correlations are stronger than the correlation of SINE density with (G+C) level (c). d, The relationship of LINE1 density in human and mouse orthologous regions is not linear, reflecting the more extreme bias of LINE1 for (A+T)-rich DNA in mouse.
We interpret these results to mean that SINE density is influenced by genomic features that are correlated with (G+C) content but that are distinct from (G+C) content per se. The fact that (G+C) content alone does not determine SINE density is consistent with the observation that some (G+C)-rich regions of the human genome are not Alu rich128,129.
Lineage-specific LINE density is also clearly correlated between mouse and human (Fig. 13b), although the relationship does not seem to be linear and it is not as strong (Spearman rank analysis, r2 = 0.45). (G+C) content seems to contribute as an independent variable (increasing r2 to 0.52), suggesting that (G+C) content itself directly affects LINE integration.
Genomic outliers
In addition to examining the general correlation in repeat density between mouse and human, we also considered some of the extreme examples. In the human genome, the four homeobox clusters (HOXA, HOXB, HOXC and HOXD) are by far the most repeat-poor regions of the human genome, with repeat content in the range of 1%. These same four regions are exceptions in the mouse genome as well. The strong selective constraints against insertion in these regions probably reflect dense, long-range regulatory information across this developmentally important gene cluster. Other repeat-poor loci in the human genome1 (about 100-kb regions on human chromosomes 1p36, 8q21 and 18q22) have independently remained repeat-poor in mouse (3.6, 6.5 and 7%, respectively) over roughly 75 million years of evolution; we speculate that this similarly reflects dense regulatory information in the region.
Conversely, we searched the mouse genome for repeat-poor regions of at least 100 kb. Again, the outliers show a clear tendency to be repeat-poor in human (see Supplementary Information). A notable feature is that in half of the selected loci the repeat-poor region is confined almost exactly to the extent of a single gene. Figure 14 shows this for the Zfhx1b locus, and also shows coincidence of exclusion of interspersed repeats with high conservation between human and mouse.
The repeat content for mouse (blue) and human (red) in 50-kb windows is shown for a 1-Mb region surrounding the Zfhx1b gene (green). Dotted lines indicate genome average for repeat content in mouse (blue) and human (red). The repeat-poor regions (<10% repeat content in mouse and human) coincide with the location of the 150-kb-long gene and regions of high conservation between human and mouse.
LINE elements prefer sex chromosomes
A conspicuous feature of the repeat distribution is that LINE elements in both human and mouse show a preference for accumulating on sex chromosomes (Figs 12 and 15). Mouse chromosome X contains almost twice the density of lineage-specific L1 copies as the mouse autosomes (28.5% compared with 14.6%). Human sex chromosomes show an even stronger bias (17.5% on X and 18.0% on Y compared with 7.5% for the autosomes). The enrichment is still highly significant even after accounting for the generally higher (A+T) content of the sex chromosomes (Fig. 15).
Error bars depict standard deviation over all autosomes (circles). Diamonds, X chromosomes; squares, human Y chromosome. The mouse Y chromosome is not represented in the whole-genome assembly, and too little clone-based information is available to be included. a, The number of lineage-specific L1 copies per megabase declines 13- to 20-fold from lowest to highest (G+C) content. This relationship is stronger in mouse and on the sex chromosomes. Note that, for the same (G+C) content, L1 density is 1.5- to twofold higher on the sex chromosomes. b, The average length of lineage-specific L1 copies peaks at around the 39% (G+C) level, where it is three- (human) to fourfold (mouse) higher than in the (G+C)-richest regions. The average length in mouse is underestimated owing to the bias against full-length young elements in the shotgun assembly. On average, L1 copies are longer on human Y than on either X chromosome or the autosomes.
The higher density of L1 on sex chromosomes had been noted in early hybridization experiments130,131 and has led to the suggestion that L1 copies may help facilitate X inactivation132,133. For chromosome Y, the accumulation probably reflects a greater tolerance for insertion (owing to the paucity of genes) and the inability to purge deleterious mutations by recombination. Consistent with the latter explanation, chromosome Y also shows a threefold higher density of full-length L1 copies (which are rapidly eliminated elsewhere in the genome134) and an overall excess of LTR element insertions. Chromosome X shows an excess of L1 copies, but not a marked excess of either full-length L1 or LTR copies. The explanation for this preferential accumulation of L1 elements on chromosome X in both the mouse and human lineages remains unclear.
Simple sequence repeats
Mammalian genomes are scattered with simple sequence repeats (SSRs), consisting of short perfect or near-perfect tandem repeats that presumably arise through slippage during DNA replication. SSRs have had a particularly important role as genetic markers in linkage studies in both mouse and human, because their lengths tend to be polymorphic in populations and can be readily assayed by PCR. It is possible that such SSRs, arising as they do through replication errors, would be largely equivalent between mouse and human; however, there are impressive differences between the two species135.
Overall, mouse has 2.25–3.25-fold more short SSRs (1–5 bp unit) than human (Table 8); the precise ratio depends on the percentage identity required in defining a tandem repeat. The mouse seems to represent an exception among mammals on the basis of comparison with the small amount of genomic sequence available from dog (4 Mb) and pig (5 Mb), both of which show proportions closer to human136 (E. Green, unpublished data; Table 8).
The analysis can be refined, however, by excluding transposable elements that contain SSRs at their 3′ ends. For example, 90% of A-rich SSRs in human are provided by or spawned from poly(A) tails of Alu and L1 elements, and 15% of (CA)n-like SSRs in mouse are contained in B2 element tails. When these sources are eliminated, the contrast between mouse and human grows to roughly fourfold.
The reason for the greater density of SSRs in mouse is unknown. Table 9 shows that SSRs of >20 bp are not only more frequent, but are generally also longer in the mouse than in the human genome, suggesting that this difference is due to extension rather than to initiation. The equilibrium distribution of SSR length has been proposed137 to be determined by slippage between exact copies of the repeat during meiotic recombination138. The shorter lengths of SSRs in human may result from the higher rate of point substitutions per generation (see above), which disrupts the exactness of the repeats.
Apart from the absolute number of SSRs, there are also some marked differences in the frequency of certain SSR classes (Table 9)136. The most extreme is the tetramer (ACAG)n, which is 20-fold more common in mouse than human (even after eliminating copies associated with B2 and B4 SINEs); the sequence does not occur in large clusters, but rather is distributed throughout the genome. In general, SSRs in which one strand is a polypurine tract and the other a polypyrimidine tract are much more common and extended in mouse than human. For the six such di-, tri- and tetramer SSRs (AG, AAG, AGG, AAAG, AAGG, AGGG), copies with at least 20 bp and 95% identity are 1.6-fold longer and tenfold more common in mouse than human.
Analysis of the distribution of SSRs across chromosomes also reveals an interesting feature common to both organisms (see Supplementary Information). In both human and mouse, there is a nearly twofold increase in density of SSRs near the distal ends of chromosome arms. Because mouse chromosomes are acrocentric, they show the effect only at one end. The increased density of SSRs in telomeric regions may reflect the tendency towards higher recombination rates in subtelomeric regions1.
Mouse genes
Genes comprise only a small portion of the mammalian genome, but they are understandably the focus of greatest interest. One of the most notable findings of the initial sequencing and analysis of the human genome1 was that the number of protein-coding genes was only in the range of 30,000–40,000, far less than the widely cited textbook figure of 100,000, but in accord with more recent, rigorous estimates55,139,140,141. The lower gene count was based on the observed and predicted gene counts, statistically adjusted for systematic under- and overcounting.
Our goal here is to produce an improved catalogue of mammalian protein-coding genes and to revisit the gene count. Genome analysis has been enhanced by a number of recent developments. These include burgeoning mammalian EST and cDNA collections, knowledge of the genomes and proteomes of a growing number of organisms, increasingly complete coverage of the mouse and human genomes in high-quality sequence assemblies, and the ability to use de novo gene prediction methodologies that exploit information from two mammalian genomes to avoid potential biases inherent in using known transcripts or homology to known genes.
We focus here on protein-coding genes, because the ability to recognize new RNA genes remains rudimentary. As used below, the terms ‘gene catalogue’ and ‘gene count’ refer to protein-coding genes only. We briefly discuss RNA genes at the end of the section.
Evidence-based gene prediction
We constructed catalogues of human and mouse gene predictions on the basis of available experimental evidence. The main computational tool was the Ensembl gene prediction pipeline142 augmented with the Genie gene prediction pipeline143. Briefly, the Ensembl system uses three tiers of input. First, known protein-coding cDNAs are mapped onto the genome. Second, additional protein-coding genes are predicted on the basis of similarity to proteins in any organism using the GeneWise program144. Third, de novo gene predictions from the GENSCAN program145 that are supported by experimental evidence (such as ESTs) are considered. These three strands of evidence are reconciled into a single gene catalogue by using heuristics to merge overlapping predictions, detect pseudogenes and discard misassemblies. These results are then augmented by using conservative predictions from the Genie system, which predicts gene structures in the genomic regions delimited by paired 5′ and 3′ ESTs on the basis of cDNA and EST information from the region.
We also examined predictions from a variety of other computational systems (see Supplementary Information). These methods tended to have significant overlap with the above-generated gene catalogues, but each tended to introduce significant numbers of predictions that were unsupported by other methods and that appeared to be false positives. Accordingly, we did not add these predictions to our gene catalogues; however, we did use them to fill in missing exons in existing predictions (see Supplementary Information).
The computational pipeline produces predicted transcripts, which may represent fragmentary products or alternative products of a gene. They may also represent pseudogenes, which can be difficult in some cases to distinguish from real genes. The predicted transcripts are then aggregated into predicted genes on the basis of sequence overlaps (see Supplementary Information). The computational pipeline remains imperfect and the predictions are tentative.
Initial and current human gene catalogue
The initial human gene catalogue1 contained about 45,000 predicted transcripts, which were aggregated into about 32,000 predicted genes containing a total of approximately 170,000 distinct exons (Table 10). Many of the predicted transcripts clearly represented only gene fragments, because the overall set contained considerably fewer exons per gene (mean 4.3, median 3) than known full-length human genes (mean 10.2, median 8).
This initial gene catalogue was used to estimate the number of human protein-coding genes, on the basis of estimates of the fragmentation rate, false positive rate and false negative rate for true human genes. Such corrections were particularly important, because a typical human gene was represented in the predictions by about half of its coding sequence or was significantly fragmented. The analysis suggested that the roughly 32,000 predicted genes represented about 24,500 actual human genes (on the basis of fragmentation and false positive rates) out of the best-estimate total of approximately 31,000 human protein-coding genes on the basis of estimated false negatives1. We suggested a range of 30,000–40,000 to allow for additional genes.
Several papers have re-analysed the initial gene catalogue and argued for a substantially larger human gene count146,147. Most of these analyses, however, did not account for the incomplete nature of the catalogoue148, the complexities arising from alternative splicing, and the difficulty of interpreting evidence from fragmentary messenger RNAs (such as ESTs and serial analysis of gene expression (SAGE) tags) that may not represent protein-coding genes149.
Since the initial paper1, the human gene catalogue has been refined as sequence becomes more complete and methods are revised. The current catalogue (Ensembl build 29) contains 27,049 predicted transcripts aggregated into 22,808 predicted genes containing about 199,000 distinct exons (Table 10). The predicted transcripts are larger, with the mean number of exons roughly doubling (to 8.7), and the catalogue has increased in completeness, with the total number of exons increasing by nearly 20%. We return below to the issue of estimating the mammalian gene count.
Mouse gene catalogue
We sought to create a mouse gene catalogue using the same methodology as that used for the human gene catalogue (Table 10). An initial catalogue was created by using the same evidence set as for the human analysis, including cDNAs and proteins from various organisms. This set included a previously published collection of mouse cDNAs produced at the RIKEN Genome Center41.
We also created an extended mouse gene catalogue by including a much larger set of about 32,000 mouse cDNAs with significant ORFs (see Supplementary Information) that were sequenced by RIKEN (see ref. 150). These additional mouse cDNAs improved the catalogue by increasing the average transcript length through the addition of exons (raising the total from about 191,000 to about 213,000, including many from untranslated regions) and by joining fragmented transcripts. The set contributed roughly 1,200 new predicted genes. The total number of predicted genes did not change significantly, however, because the increase was offset by a decrease due to mergers of predicted genes. These mouse cDNAs have not yet been used to extend the human gene catalogue. Accordingly, comparisons of the mouse and human gene catalogues below use the initial mouse gene catalogue.
The extended mouse gene catalogue contains 29,201 predicted transcripts, corresponding to 22,011 predicted genes that contain about 213,500 distinct exons. These include 12,226 transcripts corresponding to cDNAs in the public databases, with 7,481 of these in the well-curated RefSeq collection151. There are 9,785 predicted transcripts that do not correspond to known cDNAs, but these are built on the basis of similarity to known proteins.
The new mouse and human gene catalogues contain many new genes not previously identified in either genome. These include new paralogues for genes responsible for at least five diseases: RFX5, responsible for a type of severe combined immunodeficiency resulting from lack of expression of human leukocyte antigen (HLA) antigens on certain haematopoietic cells152; bestrophin, responsible for a form of muscular degeneration153; otoferlin, responsible for a non-syndromic prelingual deafness154; Crumbs1, mutated in two inherited eye disorders155,156; and adiponectin, a deficiency of which leads to diet-induced insulin resistance in mice157. The RFX5 case is interesting, because disruption of the known mouse homologue alone does not reproduce the human disease, but may do so in conjunction with disruption of the newly identified paralogue158.
Recently, Mural and colleagues45 analysed the sequence of mouse chromosome 16 and reported 731 gene predictions (compared with 756 gene predictions in our set for chromosome 16). Our gene catalogue contains 656 of these gene predictions, indicating extensive agreement between these two independent analyses. Most of the remaining 75 genes reported by ref. 45 seem to be systematic errors (common to all such programs), such as relatively short gene predictions arising from protein matches to low-complexity regions.
It should be emphasized that the human and mouse gene catalogues, although increasingly complete, remain imperfect. Both genome sequences are still incomplete. Some authentic genes are missing, fragmented or otherwise incorrectly described, and some predicted genes are pseudogenes or are otherwise spurious. We describe below further analysis of these challenges.
Pseudogenes
An important issue in annotating mammalian genomes is distinguishing real genes from pseudogenes, that is, inactive gene copies. Processed pseudogenes arise through retrotransposition of spliced or partially spliced mRNA into the genome; they are often recognized by the loss of some or all introns relative to other copies of the gene. Unprocessed pseudogenes arise from duplication of genomic regions or from the degeneration of an extant gene that has been released from selection. They sometimes contain all exons, but often have suffered deletions and rearrangements that may make it difficult to recognize their precise parentage. Over time, pseudogenes of either class tend to accumulate mutations that clearly reveal them to be inactive, such as multiple frameshifts or stop codons. More generally, they acquire a larger ratio of non-synonymous to synonymous substitutions (KA/KS ratio; see section on proteins below) than functional genes. These features can sometimes be used to recognize pseudogenes, although relatively recent pseudogenes may escape such filters.
The well-studied Gapdh gene and its pseudogenes illustrate the challenges159. The mouse genome contains only a single functional Gapdh gene (on chromosome 7), but we find evidence for at least 400 pseudogenes distributed across 19 of the mouse chromosomes. Some of these are readily identified as pseudogenes, but 118 have retained enough genic structure that they appear as predicted genes in our gene catalogue. They were identified as pseudogenes only after manual inspection. The Gapdh pseudogenes typically have no orthologous human gene in the corresponding region of conserved synteny.
To assess the impact of pseudogenes on gene prediction, we focused on two classes of gene predictions: (1) those that lack a corresponding gene prediction in the region of conserved synteny in the human genome (2,705); and (2) those that are members of apparent local gene clusters and that lack a reciprocal best match in the human genome (5,143). A random sample of 100 such predicted genes was selected, and the predictions were manually reviewed. We estimate that about 76% of the first class and about 30% of the second class correspond to pseudogenes. Overall, this would correspond to roughly 4,000 of the predicted genes in mouse. (A similar proportion of gene predictions on chromosome 16 by Mural and colleagues45 seem, by the same criteria, to be pseudogenes.) These two classes contain relatively few exons (average 3), and thus comprise only about 12,000 exons of the 213,562 in the mouse gene catalogue. Pseudogenes similarly arise among human gene predictions and are greatly enriched in the two classes above. This analysis shows the benefit of comparative genome analysis and suggests ways to improve gene prediction.
We also sought to identify the many additional pseudogenes that had been correctly excluded during the gene prediction process. To do so, we searched the genomic regions lying outside the predicted genes in the current catalogue for sequence with significant similarity to known proteins. We identified about 14,000 intergenic regions containing such putative pseudogenes. Most (>95%) appear to be clear pseudogenes (on the basis of such tests as ratio of non-synonymous to synonymous substitutions; see Supplementary Information and the section on proteins below), with more than half being processed pseudogenes. This is surely an underestimate of the total number of pseudogenes, owing to the limited sensitivity of the search.
Further refinement
We analysed the mouse gene predictions further, focusing on those whose best human match fell outside the region of conserved synteny and those without clear orthologues in the human genome. Two suspicious classes were identified. The first (0.4%) consists of 63 predicted genes that seem to encode Gag/Pol proteins from mouse-specific retrovirus elements. The second (about 2.5%) consists of 591 predicted genes for which the only supporting evidence comes from a single collection of mouse cDNAs (the initial RIKEN cDNAs41). These cDNAs are very short on average, with few exons (median 2) and small ORFs (average length of 85 amino acids); whereas some of these may be true genes, most seem unlikely to reflect true protein-coding genes, although they may correspond to RNA genes or other kinds of transcripts. Both groups were omitted in the comparative analysis below.
Comparison of mouse and human gene sets
We then sought to assess the extent of correspondence between the mouse and human gene sets. Approximately 99% of mouse genes have a homologue in the human genome. For 96% the homologue lies within a similar conserved syntenic interval in the human genome. For 80% of mouse genes, the best match in the human genome in turn has its best match against that same mouse gene in the conserved syntenic interval. These latter cases probably represent genes that have descended from the same common ancestral gene, termed here 1:1 orthologues.
Comprehensive identification of all orthologous gene relationships, however, is challenging. If a single ancestral gene gives rise to a gene family subsequent to the divergence of the species, the family members in each species are all orthologous to the corresponding gene or genes in the other species. Accordingly, orthology need not be a 1:1 relationship and can sometimes be difficult to discern from paralogy (see protein section below concerning lineage-specific gene family expansion).
There was no homologous predicted gene in human for less than 1% (118) of the predicted genes in mouse. In all these cases, the mouse gene prediction was supported by clear protein similarity in other organisms, but a corresponding homologue was not found in the human genome. The homologous genes may have been deleted in the human genome for these few cases, or they could represent the creation of new lineage-specific genes in the rodent lineage—this seems unlikely, because they show protein similarity to genes in other organisms. There are, however, several other possible reasons why this small set of mouse genes lack a human homologue. The gene predictions themselves or the evidence on which they are based may be incorrect. Genes that seem to be mouse-specific may correspond to human genes that are still missing owing to the incompleteness of the available human genome sequence. Alternatively, there may be true human homologues present in the available sequence, but the genes could be evolving rapidly in one or both lineages and thus be difficult to recognize. The answers should become clear as the human genome sequence is completed and other mammalian genomes are sequenced. In any case, the small number of possible mouse-specific genes demonstrates that de novo gene addition in the mouse lineage and gene deletion in the human lineage have not significantly altered the gene repertoire.
Mammalian gene count
To re-estimate the number of mammalian protein-coding genes, we studied the extent to which exons in the new set of mouse cDNAs sequenced by RIKEN132 were already represented in the set of exons contained in our initial mouse gene catalogue, which did not use this set as evidence in gene prediction. This cDNA collection is a much broader and deeper survey of mammalian cDNAs than previously available, on the basis of sampling of diverse embryonic and adult tissues150. If the RIKEN cDNAs are assumed to represent a random sampling of mouse genes, the completeness of our exon catalogue can be estimated from the overlap with the RIKEN cDNAs. We recognize this assumption is not strictly valid but nonetheless is a reasonable starting point.
The initial mouse gene catalogue of 191,290 predicted exons included 79% of the exons revealed by the RIKEN set. This is an upper bound of sensitivity as some RIKEN cDNAs are probably less than full length and many tissues remain to be sampled. On the basis of the fraction of mouse exons with human counterparts, the percentage of true exons among all predicted exons or the specificity of the initial mouse gene catalogue is estimated to be 93%. Together, these estimates suggest a count of about 225,189 exons in protein-coding genes in mouse (191,290 × 0.93/0.79).
To estimate the number of genes in the genome, we used an exon-level analysis because it is less sensitive to artefacts such as fragmentation and pseudogenes among the gene predictions. One can estimate the number of genes by dividing the estimated number of exons by a good estimate of the average number of exons per gene. A typical mouse RefSeq transcript contains 8.3 coding exons per gene, and alternative splicing adds a small number of exons per gene. The estimated gene count would then be about 27,000 with 8.3 exons per gene or about 25,000 with 9 exons per gene. If the sensitivity is only 70% (rather than 79%), the exon count rises to 254,142, yielding a range of 28,000–30,500.
In the next section, we show that gene predictions that avoid many of the biases of evidence-based gene prediction result in only a modest increase in the predicted gene count (in the range of about 1,000 genes). Together, these estimates suggest that the mammalian gene count may fall at the lower end of (or perhaps below) our previous prediction of 30,000–40,000 based on the human draft sequence1. Although small, single-exon genes may add further to the count, the total seems unlikely to greatly exceed 30,000. This lower estimate for the mammalian gene number is consistent with other recent extrapolations141. However, there are important caveats. It is possible that the genome contains many additional small, single-exon genes expressed at relatively low levels. Such genes would be hard to detect by our various techniques and would also decrease the average number of exons per gene used in the analysis above.
De novo gene prediction
The gene predictions above have the strength of being based on experimental evidence but the weakness of being unable to detect new exons without support from known transcripts or homology to known cDNAs or ESTs in some organism. In particular, genes that are expressed at very low levels or that are evolving very rapidly are less likely to be present in the catalogue (R. Guigó, unpublished data).
Ideally, one would like to perform de novo gene prediction directly from genomic sequence by recognizing statistical properties of coding regions, splice sites, introns and other gene features. Although this approach works relatively well for small genomes with a high proportion of coding sequence, it has much lower specificity when applied to mammalian genomes in which coding sequences are sparser. Even the best de novo gene prediction programs (such as GENSCAN145) predict many apparently false-positive exons.
In principle, de novo gene prediction can be improved by analysing aligned sequences from two related genomes to increase the signal-to-noise ratio135. Gene features (such as splice sites) that are conserved in both species can be given special credence, and partial gene models (such as pairs of adjacent exons) that fail to have counterparts in both species can be filtered out. Together, these techniques can increase sensitivity and specificity.
We developed three new computer programs for dual-genome de novo gene prediction: TWINSCAN160,325, SGP2 (refs 161, 326) and SLAM162. We describe here results from the first two programs. The results of the SLAM analysis can be viewed at http://bio.math.berkeley.edu/slam/mouse/. To predict genes in the mouse genome, these two programs first find the highest-scoring local mouse–human alignment (if any) in the human genome. They then search for potential exonic features, modifying the probability scores for the features according to the presence and quality of these human alignments. We filtered the initial predictions of these programs, retaining only multi-exon gene predictions for which there were corresponding consecutive exons with an intron in an aligned position in both species327.
After enrichment based on the presence of introns in aligned locations, TWINSCAN identified 145,734 exons as being part of 17,271 multi-exon genes. Most of the gene predictions (about 94%) were present in the above evidence-based gene catalogue. Conversely, about 78% of the predicted genes and about 81% of the exons in this catalogue were at least partially represented by TWINSCAN predictions. TWINSCAN predicted an extra 4,558 (3%) new exons not predicted by the evidence-based methods. SGP2 produced qualitatively similar results. The total number of predicted exons was 168,492 contained in 18,056 multi-exon genes, with 86% of the predicted genes in the evidence-based gene catalogue at least partially represented. Approximately 83% of the exons in the catalogue were detected by SGP2, which predicted an additional 9,808 (6%) new exons. There is considerable overlap between the two sets of new predicted exons, with the TWINSCAN predictions largely being a subset of the SGP2 predictions; the union of the two sets contains 11,966 new exons.
We attempted to validate a sample of 214 of the new predictions by performing PCR with reverse transcription (RT) between consecutive exons using RNA from 12 adult mouse tissues163 and verifying resulting PCR products by direct DNA sequencing. Our sampling involved selecting gene predictions without nearby evidence-based predictions on the same strand and with an intron of at least 1 kb. The validation rate was approximately 83% for TWINSCAN and about 44% for SGP2 (which had about twice as many new exons; see above). Extrapolating from these success rates, we estimate that the entire collection would yield about 788 validated gene predictions that do not overlap with the evidence-based catalogue.
The second step of filtering de novo gene predictions (by requiring the presence of adjacent exons in both species) turns out to greatly increase prediction specificity. Predicted genes that were removed by this criterion had a very low validation rate. In a sample of 101 predictions that failed to meet the criteria, the validation rate was 11% for genes with strong homology to human sequence and 3% for those without. The filtering process thus removed 24-fold more apparent false positives than true positives. Extrapolating from these results, testing the entire set of such predicted genes (that is, those that fail the test of having adjacent homologous exons in the two species) would be expected to yield only about 231 additional validated predictions.
Overall, we expect that about 1,000 (788+231) of the new gene predictions would be validated by RT–PCR. This probably corresponds to a smaller number of actual new genes, because some of these may belong to the same transcription unit as an adjacent de novo or evidence-based prediction. Conversely, some true genes may fail to have been detected by RT–PCR owing to lack of sensitivity or tissue, or developmental stage selection327.
An example of a new gene prediction, validated by RT–PCR, is a homologue of dystrophin (Fig. 16). Dystrophin is encoded by the DMD gene, which is mutated in individuals with Duchenne muscular dystrophy164. A gene prediction was found on mouse chromosome 1 and human chromosome 2, showing 38% amino acid identity over 36% of the dystrophin protein (the carboxy terminal portion, which interacts with the transmembrane protein β-dystroglycan). Other new gene predictions include homologues of aquaporin. These gene predictions were missed by the evidence-based methods because they were below various thresholds. These and other examples are described in a companion paper327.
Mouse and human gene structures are shown in blue on the chromosomes (pink). The mouse intron marked with an asterisk was verified by RT–PCR from primers complementary to the flanking exons followed by direct product sequencing327. Regions of high-scoring alignment to the entire other genome (computed before gene predictions and identification of predicted orthologues) are shown in yellow. Note the weak correspondence between predicted exons and blocks of high-scoring whole-genome alignment. Nonetheless, the predicted proteins considered in isolation show good alignment across several splice sites.
The overall results of the de novo gene prediction are encouraging in two respects. First, the results show that de novo gene prediction on the basis of two genome sequences can identify (at least partly) most predicted genes in the current mammalian gene catalogues with remarkably high specificity and without any information about cDNAs, ESTs or protein homologies from other organisms. It can also identify some additional genes not detected in the evidence-based analysis. Second, the results suggest that methods that avoid some of the inherent biases of evidence-based gene prediction do not identify more than a few thousand additional predicted exons or genes. These results are thus consistent with an estimate in the vicinity of 30,000 genes, subject to the uncertainties noted above.
RNA genes
The genome also encodes many RNAs that do not encode proteins, including abundant RNAs involved in mRNA processing and translation (such as ribosomal RNAs and tRNAs), and more recently discovered RNAs involved in the regulation of gene expression and other functions (such as micro RNAs)165,166. There are probably many new RNAs not yet discovered, but their computational identification has been difficult because they contain few hallmarks. Genomic comparisons have the potential to significantly increase the power of such predictions by using conservation to reveal relatively weak signals, such as those arising from RNA secondary structure167. We illustrate this by showing how comparative genomics can improve the recognition of even an extremely well understood gene family, the tRNA genes.
In our initial analysis of the human genome1, the program tRNAscan-SE168 predicted 518 tRNA genes and 118 pseudogenes. A small number (about 25 of the total) were filtered out by the RepeatMasker program as being fossils of the MIR transposon, a long-dead SINE element that was derived from a tRNA169,170.
The analysis of the mouse genome is much more challenging because the mouse contains an active SINE (B2) that is derived from a tRNA and thus vastly complicates the task of identifying true tRNA genes. The tRNAscan-SE program predicted 2,764 tRNA genes and 22,314 pseudogenes in mouse, but the RepeatMasker program classified 2,266 of the ‘genes’ and 22,136 of the ‘pseudogenes’ as SINEs. After eliminating these, the remaining set contained 498 putative tRNA genes. Close analysis of this set suggested that it was still contaminated with a substantial number of pseudogenes. Specifically, 19 of the putative tRNA genes violated the wobble rules that specify that only 45 distinct anticodons are expected to decode the 61 standard sense codons, plus a selenocysteine tRNA species complementary to the UGA stop codon171. In contrast, the initial analysis of the human genome identified only three putative tRNA genes that violated the wobble rules172,173.
To improve discrimination of functional tRNA genes, we exploited comparative genomic analysis of mouse and human. True functional tRNA genes would be expected to be highly conserved. Indeed, the 498 putative mouse tRNA genes differ on average by less than 5% (four differences in about 75 bp) from their nearest human match, and nearly half are identical. In contrast, non-genic tRNA-related sequences (those labelled as pseudogenes by tRNAscan-SE or as SINEs by RepeatMasker) differ by an average of 38% and none is within 5% divergence. Notably, the 19 suspect predictions that violate the wobble rules show an average of 26% divergence from their nearest human homologue, and none is within 5% divergence.
On the basis of these observations, we identified the set of tRNA genes having cross-species homologues with <5% sequence divergence. The set contained 335 tRNA genes in mouse and 345 in human. In both cases, the set represents all 46 expected anti-codons and exactly satisfies the expected wobble rules. The sets probably more closely represent the true complement of functional tRNA genes.
Although the excluded putative genes (163 in mouse and 167 in human) may include some true genes, it seems likely that our earlier estimate of approximately 500 tRNA genes in human is an overestimate. The actual count in mouse and human is probably closer to 350.
We also analysed the mouse genome for other known classes of non-coding RNAs. Because many of these classes also seem to have given rise to many pseudogenes, we conservatively considered only those loci that are identical or that are highly similar to RNAs that have been published as ‘true’ genes. We identified a total of 446 non-coding RNA genes, which includes 121 small nucleolar RNAs, 78 micro RNAs, and 247 other non-coding RNA genes, including rRNAs, spliceosomal RNAs, and telomerase RNA. We also classified 2,030 other loci with significant similarities to known RNA genes as probable pseudogenes.
Mouse proteome
Eukaryotic protein invention appears to have occurred largely through two important mechanisms. The first is the combination of protein domains into new architectures. (Domains are compact structures serving as evolutionarily conserved functional building blocks that are often assembled in various arrangements (architectures) in different proteins174.) The second is lineage-specific expansions of gene families that often accompany the emergence of lineage-specific functions and physiologies175 (for example, expansions of the vertebrate immunoglobulin superfamily reflecting the invention of the immune system1, receptor-like kinases in A. thaliana associated with plant-specific self-incompatibility and disease-resistance functions49, and the trypsin-like serine protease homologues in D. melanogaster associated with dorsal–ventral patterning and innate immune response176,177).
The availability of the human and mouse genome sequences provides an opportunity to explore issues of protein evolution that are best addressed through the study of more closely related genomes. The great similarity of the two proteomes allows extensive comparison of orthologous proteins (those that descended by speciation from a single gene in the common ancestor rather than by intragenome duplication), permitting an assessment of the evolutionary pressures exerted on different classes of proteins. The differences between the mouse and human proteomes, primarily in gene family expansions, might reveal how physiological, anatomical and behavioural differences are reflected at the genome level.
Overall proteome comparison
We compared the largest transcript for each gene in the mouse gene catalogue to the National Center for Biotechnology Information (NCBI) database (‘nr’ set; ftp://ftp.ncbi.nih.gov/blast/db/nr.z) using the BLASTP program178. Mouse proteins predicted to be homologues (E < 10-4) of other proteins were classified into one of six taxonomic groupings: (1) rodent-specific; (2) mammalian-specific; (3) chordate-specific; (4) metazoan-specific; (5) eukaryote-specific; and (6) other (Fig. 17). The results were similar to those from an analysis of human proteins1.
Note that only a small fraction of genes are possibly rodent-specific (<1%) as compared with those shared with other mammals (14%, not rodent-specific); shared with chordates (6%, not mammalian-specific); shared with metazoans (27%, not chordate-specific); shared with eukaryotes (29%, not metazoan-specific); and shared with prokaryotes and other organisms (23%, not eukaryotic-specific).
We annotated the current sets of mouse and human proteins with respect to the InterPro classification of domains, motifs and proteins using the InterProScan computer resource179. In this way, the proteins were assigned Gene Ontology (GO) codes180, which describe biological process, cellular compartment and molecular function. Comparisons of GO annotations between the two mammals showed no large-scale differences in molecular and cellular functions between the two protein sets (Fig. 18) that were not accountable by imperfections in gene prediction and annotation. Overall, about 72% of proteins contained at least one InterPro domain.
The GO terms assigned to mouse (blue) and human (red) proteins based on sequence matches to InterPro domains are grouped into approximately a dozen categories. These categories fell within each of the larger ontologies of cellular component (a) molecular function (b) and biological process (c) (D. Hill, personal communication). In general, mouse has a similar percentage of proteins compared with human in most categories. The apparently significant difference between the number of mouse and human proteins in the translational apparatus category of the cellular component ontology may be due to ribosomal protein pseudogenes incorrectly assigned as genes in mouse.
As expected, most of the protein or domain families have similar sizes in human and mouse (Table 11). However, 12 of the 50 most populous InterPro families in mouse show significant differences in numbers between the two proteomes, most notably high mobility group HMG1/2 box and ubiquitin domains. On close analysis, the differences for six of these families can be accounted for by differential expansion of endogenous retroviral sequences in the genomes. We return below to the issue of expansion of gene families.
Evolution of orthologues
To study the evolutionary forces that conserve proteins, we examined the set of 12,845 1:1 orthologues between human and mouse described above, expanding by nearly an order of magnitude the set of 1:1 orthologues used for evolutionary analysis14,181. These are genes for which lineage-specific duplications seem not to have occurred in either lineage.
For each orthologous gene pair, we aligned the cDNA sequences in accordance with their pairwise amino acid alignments and calculated two measures of sequence evolution: the percentage of amino acid identities and the KA/KS ratio182. The latter quantity reflects the ratio between the rates of non-synonymous (amino-acid replacing) mutations per non-synonymous site and synonymous (silent) mutations per synonymous site (see ref. 183). Non-synonymous mutations are typically subject to strong selective pressure, whereas synonymous changes are thought typically to be neutral. Orthologue pairs generally have low values of KA/KS (for example, <0.05), which implies that the proteins are subject to relatively strong purifying selection184. Proteins with KA/KS > 1 are formally defined as being subject to positive selection; that is, amino acid changes are accumulating faster than would be expected given the underlying silent substitution rate. However, proteins with KA/KS < 1 may still contain sites under positive selection, but the contribution of those sites to the KA/KS for the whole protein is offset by purifying selection at other sites185. Some care is needed, however, to exclude pseudogenes in such analyses. Because pseudogenes do not encode functional proteins, the distinction between synonymous and non-synonymous mutations is irrelevant and the apparent KA/KS ratio will converge towards 1.
For the 12,845 pairs of mouse–human 1:1 orthologues, 70.1% of the residues were identical. The median amino acid identity was 78.5% and the median KA/KS ratio was 0.115 (Fig. 19 and Table 12). Most mouse and human orthologue pairs thus have a high degree of sequence identity and are under strong-to-moderate purifying selection. One consequence of the strong sequence similarity is that computer programs such as PSI-BLAST178, that use iterative alignment to detect distant homologues, gain little by using both mouse and human sequence compared with using either genome singly. For 4,344 human proteins for which no non-primate homologue could be recognized on the basis of the human sequence, the addition of a mouse orthologue added nothing new.
The figure shows percentage residue identity and cumulative non-synonymous to synonymous codon rate ratios for total proteins and for regions with and without predicted InterPro domains, predicted SMART domains with or without known enzymatic activity, and SMART domains specific to three different subcellular compartments. The 12,845 orthologous gene pairs referred to in Table 12 were used for analysis. a, Proteins were divided into regions with and without InterPro domains, and per cent identity was calculated for total proteins (black) and for domain-containing (red line) and domain-free (grey line) regions. The higher conservation of domain-containing regions, relative to domain-free regions, is consistent with their greater functional conservation. The protein sequences are plotted in bins of 4% identity. In calculating the per cent amino acid identity between two sequences, the number of identical residues was divided by the total number of alignment positions, including positions where one sequence was aligned with a gap. b, Cumulative KA/KS ratios for total proteins (black line) and for regions with (red line) and without (grey line) predicted Interpro domains. Protein-domain-containing regions have low KA/KS ratios (<0.15), suggesting that they may be subject to greater degrees of purifying selection than are the domain-free regions. The differences in functional constraints between predicted domain regions and the rest of the protein may be found to be even more pronounced, as a significant proportion of sequences may contain as yet unpredicted protein domains. c, Cumulative KA/KS ratios for SMART domain predictions with (red line) or without (black line) known enzymatic activity. The higher proportion of catalytic domains with low KA/KS ratios is an indication of the greater purifying selection acting on these sequences. d, Cumulative KA/KS ratios for predicted SMART domains that are specific to one of three different subcellular compartments. Compared with intracellular (cytoplasmic (red) and nuclear (black)) domains, a greater proportion of secreted domains (grey) possess higher KA/KS values. This indicates that secreted, often extracellular domains are subject, on average, to greater positive diversifying selection.
We sought to quantify the relative selective pressures on protein regions containing known domains. About 65% of gene pairs encode transcripts that contain at least one InterPro domain prediction (we considered only predicted domains present in corresponding positions in both orthologues). Regions containing predicted domains had higher average percentage identities and lower KA/KS values than regions without predicted domains or than full-length proteins (Fig. 19 and Table 12). Thus, domains are under greater purifying selection than are regions not containing domains. This is consistent with the hypothesis that domains are under greater structural and functional constraints than unstructured, domain-free regions. In this analysis (as in those below), the differences in KA/KS were largely due to variations in KA (Table 12). Median KS values clustered around 0.6 synonymous substitutions per synonymous site (Table 12), indicating that each of the sets of proteins has a similar neutral substitution rate. (These results are broadly consistent with measures of neutral substitution rate provided in the repeat and evolution sections, although the precise methodologies used and categories of sites examined affect the magnitude of estimates (see Supplementary Information).)
We next considered how the molecular functions of domains affect their evolution. Domain families with enzymatic activity were found to have a lower KA/KS ratio than non-enzymatic domains (Fig. 19 and Table 12). Fewer substitutions are thus tolerated in catalytic regions, suggesting that a larger proportion of amino acids contribute to substrate binding, specificity and catalysis in enzymes. Although enzymatic domains are significantly larger than non-enzymatic domains (189 compared with 47 amino acids on average), analysis indicates that there is no significant correlation between domain length and KA/KS (r2 = 0.002).
We also examined how rates of evolution correlate with the cellular compartments in which a protein functions. We partitioned 521 of the 649 domain families in the SMART database186 into secreted, cytoplasmic or nuclear classes on the basis of published data187. The KA/KS values for the three classes showed that domains in the secreted class typically are under less purifying selection than are either nuclear or cytoplasmic domains (Fig. 19 and Table 11).
Of eight domain families with the highest (>0.15) median KA/KS values, six are specific to the secreted portions of proteins and are implicated in the mammalian defence and immune response system (Table 13). The fact that these proteins have the highest KA/KS values indicates that they are under reduced purifying selection, increased positive selection, or both. Increased positive selection may be the result of antagonistic coevolution between a mammalian host and its pathogens in a ‘genetic arms race’188, where each is under strong pressure to respond to innovations in the other genome.
Mouse orthologues of human disease genes are of particular interest to biomedical research. We examined 687 human disease genes having clear orthologues in mouse189. A total of 7,293 amino acid variants reported to be disease-associated190 were mapped to corresponding positions in the mouse sequence. The mouse sequence was identical to the normal human sequence for 90.3% of these positions, and it differed from both the normal and disease-associated sequence in human for 7.5% of the positions. To our surprise, the mouse sequence was identical to the human disease-associated sequence in a small number of cases (160, 2.2%). Although the causal connection with disease has not yet been proven in every one of these cases, there are at least 23 instances where the link between disease and mutation has been documented (Table 14). These include mutations in the cystic fibrosis transmembrane conductance regulator gene and the α-synuclein gene, which is associated with a familial form of Parkinson's disease191. In such cases, the mouse may not provide the most appropriate model system for direct study of the mutation, although understanding the basis for the species difference may prove enlightening.
We performed a similar analysis with SNPs in coding regions of human genes. We found the location of 8,322 high-quality, coding-region SNPs from HGVbase192 within human genes using the tBLASTn computer program178 and, in turn, within the corresponding positions in mouse orthologues. The mouse sequence encoded the identical amino acid as the major (more common) human allele in 67.1% of cases and as the minor human allele in 13.6% of cases. The former proportion is similar to the 70.1% of human amino acids that are conserved in mouse orthologues, indicating that most of such coding-region SNPs are not under strong selective constraint.
Evolution of gene families in mouse
As noted above, 80% of mouse proteins seem to have strict 1:1 orthologues in the human genome. Many of the remainder belong to gene families that have undergone differential expansion in at least one of the two genomes, resulting in the lack of a strict 1:1 relationship. Such gene family changes represent an insight into aspects of physiology that have emerged since the last common ancestor.
A well-documented example of family expansion is the olfactory receptor gene family, which represents a branch of the larger G-protein-coupled receptor superfamily tree193,194. Duplication of olfactory receptor genes seems to have occurred frequently in both rodent and primate lineages, and differences in number and sequence have been seen as distinguishing the degrees and repertoires of odorant detection between mice and humans. Moreover, an estimated 20% of the mouse olfactory receptor homologues194 and a higher percentage of human homologues195,196 are pseudogenes, indicating that there is a dynamic interplay between gene birth and gene death in the recent evolution of this family. The importance of these genes in reproductive behaviour is evident from defects in pheromone responses that result from deletion of the VR1 vomeronasal olfactory receptor gene cluster197.
Another example is the cytochrome P450 gene family, which is of considerable pharmacological and clinical interest. P450 cytochromes are normally terminal oxidases in multicomponent electron transfer chains, which metabolize large numbers of xenobiotic as well as endogenous compounds. Their numbers often vary among different species198. This gene family is moderately but significantly expanded in mouse (84 genes) relative to human (63 genes). By comparing the cytochrome P450 gene families from mouse, human and pufferfish (Takifugu rubripes), we found clear expansions in four subfamilies (Cyp2b, Cyp2c, Cyp2d and Cyp4a) in mouse relative to human (Fig. 20). These occur in local gene clusters that also contain unprocessed pseudogenes. The expansions appear to be associated, in part, with gender differences in the metabolism of androgens and xenobiotics (see below).
a, Phylogenetic tree, based on the neighbour-joining method297, applied to the alignment of the whole P450 protein family. Each triangle represents a cytochrome P450 family cluster. Asterisks next to a triangle represent mouse pseudogenes defined by the presence of either an in-frame stop codon or a frameshift. The colour codes are indicated in the lower-right panel. When the family presents one member in each of the studied organisms, the triangle is labelled in orange. The height of the triangle is proportional to the number of proteins, which is indicated by white-line subdivisions. Chromosomal location in mouse is shown on each of the branches for each subfamily. The lengths of the branches are not drawn to scale. Branches with significant nodes (bootstrapping value >0.7) are in black, with the remainder in blue. In 6 out of the 15 CYP2C family cases, the localization of the genomic region from which they are derived remains unassigned. b, Detailed phylogenetic tree of the CYP2C family based on the neighbour-joining method. The root of the tree was determined using a CYP2A sequence as out-group. Eight out of the 15 mouse CYP2C sequences are excluded in this tree as they are very short. Sequence identifiers are coloured on the basis of their source: red, mouse; green, human. Sequence identifiers followed by an asterisk indicate that the sequences contain either a premature in-frame stop codon or frameshift. Bootstrap values are shown at the branches. Colour codes of branches are as for a. Although the bootstrap value for the branch containing CYP2C pseudogene2 and ENSP00000285979 is rather low (0.579), it might seem that CYP2C pseudogene2 has only recently lost its function, as a putative orthologue in human (ENSP00000285979) is still clustered with it.
To explore systematically recent evolution of the mouse proteome, we searched for mouse-specific gene clusters. We identified genomic regions containing four or more homologous mouse genes that descended from a single gene in the human–mouse common ancestor; these represent local expansions in the mouse lineage. To detect such clusters, we compared all transcripts of each gene with those of five genes on either side (using the BLAST-2-Sequences program with a threshold of E < 10-4). A total of 4,563 mouse genes were found to have at least one such homologue within this window. A total of 147 such clusters containing at least four homologues was identified, of which 47 contained multiple olfactory receptor genes, which have been studied elsewhere193,199 and are not discussed further here. For the remaining 100 clusters, we then constructed dendrograms to examine the evolutionary relationship among the mouse proteins and their human homologues. This allowed us to identify those clusters containing mouse genes that are descendants of a single ancestral gene or for which multiple gene deletions had occurred in the human lineage.
In total, 25 such mouse-specific clusters were identified (Table 15; see Supplementary Information). In most cases (16), the mouse-specific cluster corresponds to only a single gene in the human genome. Among these 25 clusters, two major functional themes emerge: 14 contain genes involved in rodent reproduction and 5 contain genes involved in host defence and immunity. Each of the 14 ‘reproduction’ clusters contains at least one gene whose expression is modulated by androgens, is involved in the biosynthesis or metabolism of hormones, has an established role in the placenta, gonads or spermatozoa, or has documented roles in mate selection, including pheromone olfaction (Table 15). The fact that so many of the 25 clusters are related to reproduction is unlikely to be coincidental. Many of the most pronounced physiological differences between rodents and primates relate to reproduction, including substantial variations in placental structures, litter sizes, oestrous cycles and gestation periods. It seems likely that reproductive traits have been responsible for some of the most powerful evolutionary pressures on the mouse genome, and that the demand for innovation has been met through gene family expansions. Examination of the human genome in this way may similarly reveal gene clusters that reflect particular aspects of human reproduction.
Some of the clusters may be related to the principal differences between mice and humans in placental structure. Although both mouse and human have discoid placentae200,201, they differ in the number and types of cell layers between the maternal and fetal blood. Of the expanded gene families, the cathepsin cluster on chromosome 13 and cystatins on chromosome 16 are expressed in the placenta202,203 and may affect its development. A non-canonical homeobox cluster on chromosome X includes Pem, Psx1 and Gpbox (Psx2), which are all expressed in the placenta204,205,206,207,208.
Another cluster is related to a different specialized aspect of reproductive physiology. This cluster, on chromosome 2, contains seminal vesicle secretory proteins that are rapidly evolving, androgen-regulated proteins involved in the formation of the copulatory plug and influence the survival and efficacy of spermatozoa209,210,211.
Other clusters are closely related to hormone metabolism and response. These include clusters of prolactin-like genes on chromosome 13 (ref. 212), prolactin-inducible genes on chromosome 6 (refs 213, 214), 3-β-hydroxysteroid dehydrogenases on chromosome 3 (refs 215, 216), and cytochrome P450 Cypd genes on chromosome 15 (refs 217, 218; see Table 15).
Several of the clusters are related to olfactory cues, which have crucial roles in rodent reproduction. For example, the lipocalin-like gene cluster on chromosome X encodes proteins that are proposed to bind odorant molecules in the mucous layer overlying the receptors of the vomeronasal organ219,220.
The salivary androgen-binding protein alpha (Abpα) pheromone gene lies within a cluster on mouse chromosome 7 that contains numerous highly related genes and pseudogenes. Males apply Abpα to their pelts by licking and then deposit it on their surroundings within their territory. In laboratory behavioural experiments, female mice have been shown to have a mating preference for males with a similar Abpα genotype, possibly to avoid inter-subspecies breeding221,222. Consequently, Abpα has been proposed to have a key role in the sexual isolation between M. musculus subspecies. The hitherto unknown Abpα paralogues on chromosome 7 may represent evolutionary vestiges of previously functioning Abpα-like molecules and/or additional functional Abpα-like pheromones.
Another notable cluster of probable pheromone genes was found on chromosome X. Aphrodisin is an aphrodisiac pheromone of the female hamster Cricetus cricetus that elicits copulatory behaviour from males223. The mouse chromosome X cluster contains predicted genes that are highly sequence-similar to aphrodisin and might possess similar behavioural functions.
The five mouse clusters that encode genes involved in immunity suggest that another major evolutionary force is acting on host defence genes. The five clusters include the major histocompatibility complex (MHC) class Ib genes, two clusters of antimicrobial β-defensins, a cluster of WAP domain antimicrobial proteins and a cluster of type A ribonucleases. Ribonuclease A genes appear to have been under strong positive selection, possibly due to their significant role in host-defence mechanisms224. The mouse genome also contains other interesting examples of recently expanded gene clusters involved in immunity, which fall short of our strict definition of mouse-specific clusters because small families consisting of a few genes appear to have been present in the common ancestor. Examples include the Ly6 and Ly49 gene families, which are greatly expanded on chromosomes 15 and 6. The Ly49 genes are of particular interest because equivalent functional niches are occupied in humans and primates by a different gene family (the non-homologous KIR family of natural killer cell receptors), an instance of convergent functional evolution225,226.
The two major themes—reproduction and immunity—may not be entirely unrelated; that is, the MHC class Ib genes have roles in both pregnancy and immunity. MHC genotype is also known from ethological studies to influence mate selection, although the molecular mechanisms underlying this effect remain unknown. Within the MHC complex, the class I genes are the most divergent, having arisen after the rodent–human divergence227.
The 25 mouse-specific clusters have been generated predominantly by local gene duplication. For 74% of genes in these clusters, the most similar homologue in the mouse genome can be found either in the same cluster or within five genes from that cluster. As well as gene birth, the clusters bear witness to gene death: the Abpα, P450 Cyp4a and Cyp4d cytochrome P450, and carboxylesterase families all contain one or more predicted pseudogene.
Members of the clusters also seem to be undergoing rapid sequence evolution, as measured by the KA/KS ratio (Fig. 21). The relatively high values of KA/KS may reflect both positive selection (as genes diverge to take up new function) and the accumulation of mutations in moribund or dead genes. Previous studies have documented rapid evolution for a number of these clusters, including eosinophil-associated ribonucleases224, MHC class I227, class Cyp2d cytochromes P450 (ref. 228), Abpα subunits221, the Gpbox homeobox cluster204,206 and submandibular gland secretory and proline-rich proteins229.
a, Cumulative histogram of KA/KS values for locally duplicated, paralogous mouse-specific gene clusters (black boxes) in comparison with mouse–human orthologues (red boxes). The mouse-specific paralogues are more likely to be under positive diversifying selection. b, Box plot of KA/KS values for different locally duplicated, paralogous mouse-specific gene clusters. Full descriptions are found in Table 15. The chromosome on which the clusters are found is indicated in brackets after the abbreviated cluster name. The KA/KS values for each sequence pair in the cluster was calculated from sequences aligned using ClustalW (see Supplementary Information). The red horizontal line represents the median and the box indicates the middle 67% of the data between the 16th and 83rd percentiles. All of the paralogous clusters have median KA/KS values that are higher than the mouse–human orthologue median KA/KS (0.115), and 22 out of 25 have values greater than the 83rd percentile orthologue KA/KS (0.275). The Cyp2d category includes KA/KS values calculated separately over two sequence-similar regions in the alignment.
Genome evolution: selection
Investigation of the two principal forces that shape the evolution of the mouse and human genomes—mutation and selection—requires looking beyond coarse-scale identification of regions of conserved synteny and purely codon-based analysis of orthologues, to fine-scale alignment of the two genomes at the nucleotide level.
The substantial sequence divergence between the mouse and human genomes is still low enough that orthologous sequences undergoing neutral drift remain conserved enough for them to be aligned reliably. The challenge then is to use such alignments to tease apart the effects of neutral drift, which can teach us about underlying mutational processes, and selection, which can inform us about functionally important elements. It should be emphasized that sequence similarity alone does not imply functional constraint.
In this section, we use whole-genome alignments to explore the extent of sequence conservation in neutral sites (such as ancestral repeat sequences), known functional elements (such as coding regions) and the genome as a whole. By comparing these, we are able to estimate the proportion of regions of the mammalian genome under evolutionary selection (about 5%), which far exceeds the amount attributable to protein-coding sequences. In the next section, we then use the neutral sites to study how mutational forces vary across the genome.
Fine-scale alignment of genomes
We began by creating a catalogue of sequence alignments between the mouse and human genomes. The alignments were produced by the BLASTZ328 program by comparing all non-repeat sequences across the genome to identify all high-scoring matches (see Supplementary Information; available for download at http://genome.ucsc.edu/downloads.html), then, using these as seeds, we extended the alignments into the surrounding regions, including into repeat sequences. To make the catalogue as comprehensive as possible, a given region in one genome was allowed to align to multiple, possibly non-syntenically conserved regions in the other genome. In fact, only a small proportion of the genome aligned to multiple regions (about 3.3%) or to non-syntenic regions (about 3.2%); the conclusions below are not significantly altered if we restrict attention to sequences that match uniquely in syntenic regions. Within the regions forming alignments, about 88.4% of individual human bases were aligned to bases in mouse, with the remainder aligned to indels (insertions or deletions). The alignments included approximately 98% of known coding regions, indicating that they correctly captured known, well-conserved sequence.
Regions that could be aligned clearly at the nucleotide level totalled about 1.1 Gb, corresponding to roughly 40% of the human genome (Fig. 22). This proportion may seem high if one imagines that all such sequence conservation reflects biological function, but it does not. Simulation experiments show that DNA sequences subjected to random mutation at the neutral rate that has occurred between the human and mouse genomes (see below) can still be readily aligned by computer. In other words, most of the non-functional orthologous sequences should still be alignable. Consistent with this analysis, the alignable portion of the genomes contains a vast number of ancestral repeats, primarily relics of transposons that were present in the genome of our common ancestor with mouse and most of which are non-functional.
Typically, 40% of the human genome sequence aligns to mouse. a, b, Approximately 98% of a 2,050-bp region on human chromosome 20 aligns to the orthologous region on mouse chromosome 2 (a), and 56% of a 5,250-bp region on human chromosome 2 aligns to the orthologous region on mouse chromosome 1 (b). In both cases, the alignment skips over young/lineage-specific repeats (red boxes), but aligns through most of the ancestral repeats (blue boxes) and non-repetitive sequence (no colour). The ancestral repeats that do align are, not unexpectedly, identified as the same repeat category. Mouse also has a larger number of simple-sequence repeats (green boxes).
But if orthologous sequences should be readily alignable, the question becomes: why isn't the alignable portion much higher than 40%? In fact, the proportion is broadly consistent with what would be expected given the probable rate of turnover of sequence in the mouse and human genomes.
As a starting point, let us assume that the genome size of the last common ancestor was about 2.9 Gb (similar to the modern genomes of human and most other mammals) and let us focus only on large-scale insertions and deletions, ignoring nucleotide-level indels within aligned regions and lineage-specific duplications. These assumptions will be relaxed below.
This would imply no net change in genome size in the human lineage despite the accumulation of about 700 Mb of lineage-specific repeat sequence since the common ancestor (see section on repeats). This would require approximately 700 Mb of deletions, implying that about 24% (700 out of 2,900) of the ancestral genome was deleted and about 76% retained in the human lineage. It would also imply a net loss of about 400 Mb in the mouse lineage, despite the probable addition of about 900 Mb of lineage-specific repeat sequences, an estimate about 10% higher than that given by the RepeatMasker program to allow for incomplete sensitivity in the more rapidly changing mouse genome. This would imply roughly 1,300 Mb of deletions, corresponding to the deletion of about 45% (1,330 out of 2,900) and retention of 55% of the ancestral genome.
If there was no correlation in the fixation of deletions in the two lineages, the expected proportion of the ancestral genome retained in both lineages would be about 42% (76% × 55%). Complete independence is unlikely because deletions of functional sequences would have been selectively disadvantageous. However, deletions of modest size may largely be neutral given the relatively low proportion of functional sequence in the genome.
The estimates can be adjusted (see Supplementary Information) to account for nucleotide-level insertions and deletions and lineage-specific duplications (the expectation remains roughly the same), or to allow for different assumptions about ancestral genome size (the expectation increases by 3–4% for an intermediate size of about 2.7 Gb). This simple analysis suggests that the observed proportion of alignable genome (about 40%) is not surprising, but rather it probably reflects the actual proportion of orthologous genome remaining after the deletion in the two lineages.
In a preliminary test of this hypothesis, we identified ancestral repeats in the mouse that lay in intervals defined by orthologous landmarks. Examination of the corresponding interval in the human genome showed a rate of loss of these elements, broadly consistent with the 24% deletion rate in the human lineage assumed above (see Supplementary Information).
Such a deletion rate in the human lineage over about 75 million years is also roughly compatible with the observation that roughly 6% has been deleted over about 22 million years since the divergence from baboon, an estimate derived from the sequencing of specific regions in human and baboon (E. Green, unpublished data). Although we do not have a corresponding direct estimate of large-scale deletions in the mouse lineage, the predicted rate of about 45% is roughly twice as high as for the human lineage, which is similar to the ratio seen for nucleotide substitutions.
Rate of neutral substitution
The genome-wide alignments can be used to measure divergence rates for different types of sequence. The neutral substitution rate, for example, can be estimated from the alignment of non-functional DNA. We believe that the best representative of this class is ancestral repeat sequence, representing transposable elements inserted and fixed before the mouse–human divergence. Such ancestral repeats are more likely than any other sequence in the genome to have been under no functional constraint.
The human–mouse alignment catalogue contains approximately 165 Mb of ancestral repeat sequences, with most being clearly orthologous by alignment of adjacent non-repetitive DNA. These alignments show 66.7% sequence identity. The observed base changes can be used to infer the underlying substitution rate, which includes back mutations, by using various continuous-time Markov models230. Applying the REV model231 to the ancestral repeat sites, we estimate that neutral divergence has led to between 0.46 and 0.47 substitutions per site (see Supplementary Information). Similar results are obtained for any of the other published continuous-time Markov models that distinguish between transitions and transversions (D. Haussler, unpublished data). Although the model does not assign substitutions separately to the mouse and human lineages, as discussed above in the repeat section, the roughly twofold higher mutation rate in mouse (see above) implies that the substitutions distribute as 0.31 per site (about 4 × 10-9 per year) in the mouse lineage and 0.16 (about 2 × 10-9 per year) in the human lineage.
Having established the neutral substitution rate by examining aligned ancestral repeats, we then investigated a second class of potentially neutral sites: fourfold degenerate sites in codons of genes. Fourfold degenerate sites are subject to selection in invertebrates, such as Drosophila, but the situation is unclear for mammals. We examined alignments between fourfold degenerate codons in orthologous genes. The fourfold degenerate codons were defined as GCX (Ala), CCX (Pro), TCX (Ser), ACX (Thr), CGX (Arg), GGX (Gly), CTX (Leu) and GTX (Val). Thus for Leu, Ser and Arg, we used four of their six codons. Only fourfold degenerate codons in which the first two positions were identical in both species were considered, so that the encoded amino acid was identical. Slightly fewer than 2 million such sites were studied, defined in the human genome from about 9,600 human RefSeq cDNAs and aligned to their mouse orthologues. The observed sequence identity in fourfold degenerate sites was 67%, and the estimated number of substitutions per site, between 0.46 and 0.47, was similar to that in the ancestral repeat sites (see Supplementary Information).
Conservation in gene-related features
We used the genome-wide alignments to examine the extent of conservation in gene-related features, including coding regions, introns, untranslated regions, upstream regions and CpG islands.
For each type of feature, we characterized the nature of sequence conservation (including typical percentage identity, inferred substitution rates and insertion/deletion rate). We also defined a conservation score S that measures the extent to which a given window (typically 50 or 100 bp, in applications below) shows higher conservation than expected by chance. The conservation score S for an aligned region R is the normalized fraction of aligned bases that are identical (obtained by subtracting the mean and dividing by the standard deviation) and is given by:

where n is the number of sites within the window that are aligned, p is the fraction of aligned sites that are identical in the two genomes, and µ, is the average fraction of sites that are identical in aligned ancestral repeats in the surrounding region (µ, = 0.667 as a genome-wide average, but, as discussed below, fluctuates locally). When the conservation score S is calculated for the set of all ancestral repeats, it has a mean of 0 (by definition) and a standard deviation of 1.19 and 1.23 for windows of 50 and 100 bp, respectively (Fig. 23). This defines the typical fluctuation in conservation score in neutral sequences. The properties of the alignments are shown in Table 16 and the distribution of conservation scores relative to neutral substitution is shown in Fig. 24.
The empirical distribution of S(R) for all 1.9 million non-overlapping 50-bp windows (blue) containing at least 45 aligned ancestral repeat sites (standard deviation 1.19) and 1.7 million non-overlapping 100-bp windows (green) containing at least 50 aligned ancestral repeat sites (standard deviation 1.23). Both curves are bell-shaped, with a mean of zero, but the standard deviations are higher than would be expected if the sites in each window were independent and conserved with (locally estimated) probability µ, . In that case the distribution of S would be approximately normal with a standard deviation of 1. Thus, these data show that there is some dependency between the substitutions within the window.
We required that at least 50 bp be aligned in each window. a–d, Comparisons with coding exons (blue) and introns (green) (a), 5′ UTR (blue) and 3′ UTR (green) (b), 200-bp upstream of transcription start (blue) and 200 bp downstream of transcription end (green) (c), and CpG islands (blue) and known regulatory regions (green) (d) are shown. The RefSeq database was used to define gene features. CpG islands were determined as discussed in the text, and known regulatory regions were collected as discussed in the text.
Coding regions are distinctive in many ways. They show the highest degree of conservation (85% sequence identity or 0.165 substitutions per nucleotide site). Alignment gaps are tenfold less common than in non-coding regions. In addition, 52% of coding regions have highly significant alignments to more than one genomic region (typically, paralogues and pseudogenes), whereas only 3.3% of the genome shows such multiple alignments.
Introns are very similar, in most respects, to the genome as a whole in terms of percentage identity, gaps and multiple alignment statistics.
Conservation levels in 5′ and 3′ UTRs are similar to one another and intermediate between levels in coding regions and introns. The sequence identity of 75–76% is well above the intronic level of 69%. Note that our estimate of sequence identity is higher than the 70–71% reported previously181, in large part because that study used a global rather than a local alignment programme. The insertion and deletion characteristics of the UTRs are very similar to those of introns. Overall, 5′ UTRs are slightly better conserved than 3′ UTRs; however, significantly more of 3′-UTR sequence is covered by multiple alignments than 5′-UTR sequence (21% compared with 16%). This may reflect the fact that pseudogene insertion tends to proceed from the 3′ end and often terminates before completion.
Promoter regions are of considerable interest. We analysed the regions located 200 bp upstream of transcription start because they were likely to contain important promoter and regulatory signals. However, such analysis is necessarily limited by the fact that transcriptional start sites remain poorly defined for many genes. With this caveat, the upstream regions share many characteristics of 5′ UTRs but have a lower percentage identity, a significantly lower proportion covered by multiple alignments, and a higher (G+C) content.
CpG islands show a conservation level similar to those of promoter and UTR regions (Fig. 24).
We also observed that levels of conservation were not uniform across these features (coding regions, introns, UTRs, upstream regions and CpG islands)232. Figure 25 shows how conservation levels vary regionally within the features of a ‘typical’ gene. Sequence identity rises gradually from a background level to 78% near the approximate transcription start site, where the level reaches a plateau. It is possible that sharper definitions of transcriptional start sites would allow the footprint of the TATA box and other common structures near the transcription start site to emerge. Conversely, many human promoters lack a TATA box, and transcription start at such promoters is not typically sharply defined233. Sequence identity falls slowly across the 5′ UTR, and then starts to rise again near the start codon. As expected, conservation levels rise sharply at the translation start site234, remain high throughout the coding regions, and have sharp peaks at splice sites. After the stop codon, the per cent identity is relatively low for most of the 3′ UTR, but then begins to increase about 200 bases before the polyadenylation site. The main polyadenylation signal is AATAAA or ATTAAA positioned 10–30 bases upstream of polyadenylation235. The region of increased conservation is considerably longer than can be explained by the polyadenylation signal alone, suggesting that other 3′-UTR regulatory signals, such as those that affect mRNA stability and localization, may frequently occur near the end of the mRNA. After the polyadenylation site, there is a 30-base plateau of moderate conservation, corresponding to the weaker (T)-rich or (G+T)-rich downstream region following the polyadenylation signal.
a, Conservation across a generic gene, on the basis of 3,165 human RefSeq mRNAs with known position in the genome. We sampled 200 evenly spaced bases across each of the variable-length regions labelled, resampling completely from regions shorter than 200 bp. The graph shows the average percentage of bases aligning and the average base identity when there is an alignment over each sample. There are peaks of conservation at the transition from one region to another. Here, in contrast to Table 16, only reviewed RefSeq mRNAs were used, and only those having at least 40 bases of annotated 5′ and 3′ UTRs. The resulting picture, however, is nearly indistinguishable from that obtained by using all RefSeq genes with at least 40 base UTRs. b, Conservation near translation start site using the same data set as in a. The bars show per cent identity of the 15 bases to either side of translation start. Note the extreme conservation of the first codon. After this, there is substantially less conservation at the third codon position. The peak at position -3 corresponds to a purine in the Kozak consensus sequence. c, Conservation near the 5′ splice site. The peak of conservation corresponds to the AG/GT consensus at this location, with the first G in the intron being nearly invariant. A G in the fifth base of the intron is also found in a large majority of 5′ splice sites. An echo of the variation in the third codon position occurs here because it is common for exons to begin and end at codon boundaries. d, Conservation near the 3′ splice site. Conservation in the last two bases of the intron—always AG for introns processed by the major spliceosome—is very apparent. The polypyrimidine tract beginning five bases into the intron is also visibly conserved. Once again, an echo of the variation in the third codon position can be seen.
Conservation of gene structure
We also examined the conservation of exon structure and splice signals in more detail using 1,506 pairs of human–mouse RefSeq genes confidently assigned to be orthologous (http://www.ncbi.nlm.nih.gov/HomoloGene/). As previously reported using smaller data sets236, overall gene structures are highly conserved between orthologous pairs: 86% of the cases (1,289 out of 1,506) have the identical number of coding exons, and 46% (692 out of 1,506) have the identical coding sequence length. When we consider all exons rather than just coding exons, we find that 941 pairs (62%) have the same number of exons. The true concordance of gene structure between the two species is probably higher, because differences will be exaggerated by differential representation of alternative splice forms between the two data sets, difficulties in mapping the cDNA sequences back to the genome, and the absence of true 5′ and 3′ ends.
The set of 1,289 genes with an identical number of coding exons contains 10,061 pairs of orthologous exons (plus 124 intronless genes). Exon length between orthologous exons is highly conserved: 9,131 (91%) of these human–mouse exon pairs have identical exon length. When exon pairs do have different lengths, the differences are predominantly multiples of three (858 out of the 930 with different lengths), as expected from coding-frame constraints. Nearly all orthologous exons conserve phase (10,015 or 99.5%). In contrast, only 90 out of 8,896 orthologous introns (1%) have identical length, although there is strong correlation between the lengths of orthologous introns. Consistent with the smaller size of the mouse genome overall, orthologous mouse introns tend to be shorter. Excluding outliers, the average human intron in this data set is 4,661 bp, whereas the average mouse intron is 3,888 bp.
Within the set of 1,506 orthologous human–mouse gene pairs, there are 22 cases in which the overall coding length is identical between the gene pairs, but they differ in the number of exons. Most of these cases can be explained by a single intron insertion/deletion (Fig. 26)237, demonstrating the dynamic (but slow) evolution of gene structure.
We also found several non-canonical splice sites in the set of 8,896 orthologous introns, including RTATCCTY 5′ splice signals characteristic of U12 introns, which are singularly conserved (see ref. 238 for review). We found this 5′ splice signal in 20 human and 22 mouse introns from the set of 8,896, and 19 of these cases correspond to orthologous introns, indicating high levels of conservation of this distinct splicing mechanism. Also conserved are the non-canonical GC-AG introns (mechanistically identical to the GT-AG canonical introns): in the set there are 23 non-canonical GC-AG introns in human and 23 in mouse, including 19 orthologous pairs.
Conservation in known regulatory regions
We similarly sought to study the extent of conservation in regulatory control regions of genes232,239,240. So far, relatively few regulatory elements have been studied extensively. We compiled a list of 95 well-characterized regulatory regions, including some liver-specific241, muscle-specific242 and general regulatory regions243. The sequences were carefully checked against the primary publications and trimmed to contain the smallest reported functional unit. The distribution of the elements was: 10% in introns, 85% in the immediate vicinity (<2 kb) of promoters, and 5% more distal from promoters. About 19% overlapped a CpG island.
The extent of conservation (Fig. 24 and Table 16) was considerably lower than in coding regions, but much higher than the neutral rate in ancestral repeats or than the average rate across the genome. Overall, the known regulatory regions showed a level of conservation similar to that of 5′ UTRs. The (G+C) content is also substantially higher for the regulatory elements than for the genome as a whole, a property shared with exons and 5′ UTRs.
Although the extent of conservation in regulatory regions—as measured by the score S(R)—overlaps with that in neutral DNA (Fig. 24), this does not preclude the use of this measure to identify candidate regulatory elements. An example is given by the insulin-like growth factor binding protein acid-labile subunit gene (IGFALS), where the region surrounding a well-known transcription factor binding site244,245,246 stands out as unusually conserved using this measure (Fig. 27). More sophisticated models, such as Markov models on the fine texture of the alignments (matches, transitions, transversions and gaps), may discriminate regulatory regions under selection from neutrally evolving regions with better efficiency329.
In the track near the top of figure, the two coding exons of the gene are displayed as taller blue rectangles, UTRs as shorter rectangles, and the intron, which separates the coding exons, is shown as a barbed line indicating direction of transcription (the gene is on the reverse strand). Log probability scores (L-scores) for all 50-bp windows are shown below the gene. The L-score is -log10(p), where p is the probability under the neutral density, Sneutral, of getting a conservation score as high as is observed in the window. Many windows in the coding region get L-scores greater than 3, indicating less than a 1/1,000 chance of occurring under neutral evolution (Pselected(S) > 0.94; see Fig. 28), and some in a local peak in the upstream region of the gene on the right show L-scores greater than 2, indicating less than a 1/100 chance of occurring (Pselected(S) > 0.75). The red bar shows the location of the interferon-γ-activated sequence-like element (GLE), which is bound by transcription factors from the STAT5a and STAT5b protein family to control expression of this gene244,245. Additional regulatory elements may be located in the other peaks of conservation. This figure is taken with permission from the UCSC browser (http://genome.ucsc.edu).
Proportion of genome under selection
We then set out to investigate the fraction of a mammalian genome under evolutionary selection for biological function.
To do this, we estimated the proportion of the genome that is better conserved than would be expected given the underlying neutral rate of substitution. We compared the overall distribution Sgenome of conservation scores for the genome to the neutral distribution Sneutral of conservation scores for ancestral repeats (Fig. 23, blue curve) using a genome-wide set of 14.3 million non-overlapping 50-bp (human) windows, each containing at least 45 bp (mean 48.67 bp) of aligned sequence. The genome-wide score distribution for these windows has a prominent tail extending to the right, reflecting a substantial excess of windows with high conservation scores relative to the neutral rate (Fig. 28). The excess can be estimated by decomposing the genome-wide distribution Sgenome as a mixture of two components: Sneutral and Sselected (reflecting windows under selection).
a, The genome-wide density of conservation scores, Sgenome (dark blue), was decomposed into a mixture of two component densities: Sneutral (red) and Sselected (light blue and grey). Sgenome is derived from the conservation scores S(R) for all windows of 50 bp in the human genome with at least 45 bases aligning to mouse. Sneutral is a scaled version of the Sneutral density from the blue curve in Fig. 23 for the 50-bp windows in ancestral repeats, representing neutrally evolving DNA. Sselected is the difference between the blue density and the red component, and thus represents a scaled version of Sselected, the predicted density for conservation scores of 50-bp windows in the human genome that are evolving under selection. The scaling factors are the estimated mixture coefficients, which are p0 = 0.792 for Sneutral, and 1 - p0 = 0.208 for Sselected. The coefficient p0 is calculated as the minimum of the ratio between Sgenome(S) and Sneutral(S) for all values of S, giving a conservative estimate that maximizes the share of the mixture attributed to Sneutral. b, The probability, Pselected(S), that a 50-bp window is under selection as a function of its conservation score S = S(R). This function is derived from the mixture decomposition by setting Pselected(S) = 1 - p0Sneutral(S)/Sgenome(S).
The mixture coefficients indicate that at least 20.8% of the windows are under selection, with the remainder consistent with neutral substitution. Because about 25.2% of all human bases are contained in the windows, this suggests that at least 5.25% (25.2% of 20.8%) of the 50-base windows in the human genome is under selection. Repeating the analysis on more stringently filtered alignments (with non-syntenic and non-reciprocal best matches removed) requiring different numbers of aligned bases per window and with 100-bp windows, yields similar estimates, ranging mostly from 4.8% to about 6.1% of windows under selection (D. Haussler, unpublished data), as does using an alternative score function that considers flanking base context effects and uses a gap penalty330. Significantly smaller window sizes, for example, 30 bp, do not provide sufficient statistical separation between the neutral and genome-wide score distributions to provide useful estimates of the share under selection.
The analysis thus suggests that about 5% of small segments (50 bp) in the human genome are under evolutionary selection for biological functions common to human and mouse. This corresponds to regions totalling about 140 Mb of human genomic DNA, although not all of the nucleotides in these windows are under selection. In addition, some bases outside these windows are likely to be under selection. In a loose sense, these regions might be regarded as containing the ‘functional’ conserved subset of the mammalian genome. Of course, it should be noted that non-conserved sequence may have important roles, for example, as a passive spacer or providing a function specific to one lineage. Notably, protein-coding regions of genes can account for only a fraction of the genome under selection. From our analysis of the number and properties of genes, coding regions comprise only about 1.5% of the human genome and account for less than half of the segments under selection.
What accounts for the remainder of the genome under selection? About 1% of the genome is contained in untranslated regions of protein-coding genes, and some of this sequence is under some functional constraint. Another main class of interest are those sequences that control gene expression, such as the control element for the IGFALS gene shown in Fig. 27; if a typical gene contains a few such regulatory sequences, there may be tens to hundreds of thousands of such elements. In addition, conserved sequences probably encode non-protein-coding RNAs (which remain difficult to discern) and chromosomal structural elements. Furthermore, some of the conserved fraction may correspond to sequences that were under selection for some period of time but are no longer functional; these could include recent pseudogenes. In an accompanying paper, Dermitzakis and colleagues show that a large number of conserved sequences on human chromosome 21 are actively conserved but are unlikely to be genes, suggesting that a large number of non-coding sequence are under selection247. Characterization of the conserved sequences should be a high priority for genomics in the years ahead.
The analysis above allows us to infer the proportion of the genome under selection by decomposing the curve Sgenome into curves Sneutral and Sselected. Importantly, it does not definitively assign an individual conserved sequence as being neutral or selected. One can calculate, for a sequence with conservation score S, the probability Pselected(S) that the window of sequence belongs to the selected subset (Fig. 28). The probability exceeds 83% for sequences with S > 3 and 93% for S > 4, but is only 52% for S = 2. In other words, some functionally important sequence cannot be separated cleanly from the tail of the distribution of neutral conservation.
How can we cleanly separate neutral and selected sequences? One solution is to extend the analysis from two species to multiple species from different branches of the mammalian radiation. Neutral sequences will tend to drift in different ways along each lineage, whereas selected sequences will tend to preserve specific sites. Multiple species comparisons should thus sharpen and separate the distributions of conservation scores, Sneutral and Sselected.
Genome evolution: mutation
Genome-wide alignments also allow us to investigate how the patterns of neutral substitution, deletion and insertion vary across the genome, providing an insight on the underlying mutational processes.
Substitution rate varies across the genome
Significant variation in the level of sequence conservation has been reported in several small-scale studies of human and mouse genomic regions10,248,249,250,251,252,253,254 and in several larger-scale studies of coding sequences255,256,257,258,259,260. It has not been clear in all cases whether the variation reflects differences in neutral substitution rates or in selection. The human–mouse genome alignments allow us to address the variation more comprehensively and to test for co-variation with the rates of other processes, such as insertions of transposable elements255 and meiotic recombination258.
We used the collection of aligned ancestral repeats and aligned fourfold degenerate sites to calculate the apparent neutral substitution rate for about 2,500 overlapping 5-Mb windows across the human genome. To accurately follow fluctuations while accounting for regional changes in base composition, the regional nucleotide substitution rate in ancestral repeat sites, tAR, was calculated separately for each 5-Mb window by maximum likelihood estimation of the parameters of the REV model using only the ancestral repeat sites in the window (average of about 280,000 sites per window). The regional nucleotide substitution rate in fourfold degenerate sites, t4D, was calculated similarly from an average of about 3,700 fourfold degenerate sites per window. Windows with fewer than 800 ancestral repeats or fourfold degenerate sites were discarded.
The mean and standard deviations across the windows were tAR = 0.467 ± 0.022 and t4D = 0.447 ± 0.067 substitutions per site. The standard deviation is much larger (over tenfold and threefold, respectively) than would be expected from sampling variance. These data clearly indicate substantial regional fluctuation. Regional variation is also evident in comparing the average rates on different chromosomes (Fig. 29). Notably, the neutral substitution rate is lowest for chromosome X. This observation is consistent with recent reports, including our initial analysis of the human genome1, that the mutation rate is about twofold lower in female meiosis than male meiosis. Because the proportion of time spent in the female germ line for chromosome X is 2/3 and for autosomes is 1/2, the predicted substitution rate for chromosome X should be about 8/9 or 89% of the genome-wide average. In fact, the observed ratio is 87% for fourfold degenerate sites and 92% for ancestral repeat sites. This would be consistent with (but does not prove) a roughly twofold lower mutation rate in the female germ line during the history of both the human and mouse lineages, and it explains a small amount of the variation in the genome-wide substitution rate. Nonetheless, the variability among autosomes is still much greater than could occur under a uniform substitution process, suggesting the existence of long-range factors that affect the mutation rate.
a, Estimates are made from the REV model using all aligned sites of the given type in the chromosome. Dashed lines show the genome-wide averages. Human chromosome 19 is a conspicuous outlier for its very large number of substitutions in fourfold degenerate sites (also noted in ref. 259); notably, its substitution rate in ancestral repeat sites is normal. Chromosome X shows lower rates of substitution in both types of sites, consistent with the observation that the male mutation rate is approximately twice the female rate1 (see text). Variability in neutral rates among autosomes is significant, as noted in ref. 13. b, Scatter plot of tAR against t4D for 2,424 5-Mb windows in the human genome with at least 800 aligning sites. The red line is the linear regression line (r2 = 0.22; P < 10-6).
Looking at a finer scale, the two measures tAR and t4D are strongly correlated across the genome (Fig. 29). They often exhibit similar behaviour across a human chromosome, as seen for human chromosome 22 (Fig. 30).
a, Variation in tAR (red) and t4D (blue) in 5-Mb windows, overlapping by 4-Mb, along human chromosome 22. Only windows with at least 800 aligned fourfold degenerate sites and 800 aligned ancestral repeat sites are shown. The position of the window is plotted at the midpoint. Horizontal dotted lines indicate the genome-wide estimates of tAR and t4D. Confidence intervals were computed on the basis of the number of ancestral repeat and fourfold degenerate sites aligning in each window; points where the confidence interval does not overlap the genome-wide estimate indicate windows with significant differences in evolutionary rate. b, Similar to a, but with t*AR and t*4D, the normalized rates obtained taking residuals of tAR and t4D from the quadratic functions of (G+C) content shown in Fig. 31. c, Fraction of DNA (blue) that is not in lineage-specific repeats identified by RepeatMasker and does not align to mouse, NAanc, and the fraction of DNA (green) contained in human lineage-specific LTR repeats identified by RepeatMasker, along with t*AR (red), calculated in overlapping 5-Mb windows as in b. d, SNP density (blue) in each overlapping 5-Mb window (average number of SNPs per 10 kb) calculated using SNPs from random reads (The SNP Consortium website; data were collected in July 2002, http://snp.cshl.org). The average recombination rate (black) in each 5-Mb window, in cM per Mb, estimated from the deCode genetic map269 is shown, as well as t*AR (red), calculated in overlapping 5-Mb windows as in b.
What properties of chromosomal DNA could account for the variation in substitution rate? One possible explanation is local (G+C) content, but previous studies disagree on whether it correlates strongly with divergence92,255,262,263. We find that tAR and t4D vary with local (G+C) content, although the dependence is nonlinear262,264 and is better fitted by regression with a quadratic curve263 (Fig. 31). In other words, the substitution rate seems to be higher in regions of extremely high or low (G+C) content, with the sign of the correlation differing in regions with high versus low (G+C) content. This pattern persists if CpG substitutions are removed from the analysis (data not shown).
Notably, tAR and t4D show different dependence on local (G+C) content. In particular, t4D increases more sharply with high (G+C) content, whereas tAR does not show as much divergence. In this and some other properties, tAR and t4D show differing patterns; hence they are not equivalent neutral sites. Differences in the nature of the dependence on local (G+C) content imply that the (G+C) content is a confounding variable in comparing tAR and t4D. Accordingly, we normalized the rates for local (G+C) content by calculating the residuals, t*AR and t*4D, with respect to the quadratic regressions above. The correspondence along chromosome 22 (a particularly (G+C)-rich chromosome) is markedly enhanced (r2 increases from 0.55 to 0.75) by this correction (Fig. 30), as is the overall genome-wide correlation (r2 increases from 0.22 to 0.33).
Substitution rate co-varies with other evolutionary rates
In addition to nucleotide substitutions, genomes evolve by insertion (primarily of transposable elements) and deletion. We examined the rate of deletion in the mouse genome, as measured by the fraction of non-aligning ancestral human DNA (NAanc). Although some of the non-alignable sequence may represent lineage-specific insertions not detected by RepeatMasker (http://ftp.genome.washington.edu/cgi-bin/RepeatMasker)177 or failure to align some orthologous sequences, the great bulk probably represents deletions in the mouse genome. The fraction NAanc varies markedly across overlapping windows of 5 Mb, with a range from 0.295 to 0.985 and mean and standard deviation 0.521 ± 0.095.
We also examined the rate of insertion (and retention) in the human genome since its divergence from mouse, as measured by the proportion of lineage-specific repeats in overlapping 5-Mb windows across the human genome. The overall level of insertion and retention showed substantial variation across the genome, ranging from 0.159 to 0.805 with a mean of 0.290 ± 0.063. To avoid complications from the tendency of some repeats, such as Alus, to be selectively removed from some regions of the genome1, we used one family of repeats, the LTRs, to monitor the relative frequency of insertion and retention. Similar to repeats as a whole, the fraction of each window occupied by lineage-specific LTRs varies substantially across the human genome, ranging from 0 to 0.378, with a mean of 0.0598 ± 0.0197.
All three forces that alter the genome (nucleotide substitution, deletion and insertion) thus vary substantially across the genome. Moreover, they are significantly correlated and tend to co-vary along chromosomes (Fig. 30 and Table 17). Notably, these three measures of interspecies divergence are also correlated with recent substitutions in the human genome, as measured by the density of SNPs identified by the SNP Consortium265 (Fig. 30).
Furthermore, recent studies report that divergence at fourfold degenerate sites and SNP frequency are both correlated with the local rate of meiotic recombination258,266,267,268. We examined the relationship between our measures of genome-wide divergence and recombination rate using recently reported high-resolution measurements of recombination rates in the human genome269. Both measures of neutral substitution rate and SNP rate showed a significant correlation with recombination rate (Fig. 30 and Table 17).
The correlations above are not explained by co-variation with local (G+C) content. All except the correlation between SNP frequency and LTR insertion rate remain significant when dependence on underlying human (G+C) content is factored out by taking the residuals of a quadratic regression on regional human (G+C) content; indeed, the correlations are for the most part enhanced (Table 17). Similarly, correlations remain significant when the difference between the (G+C) content of orthologous mouse and human regions is also factored out261. Thus, (G+C) content changes between mouse and human, as explored previously259, do not adequately explain the correlations. Finally, to obtain more rigorous estimates of significance, the correlations were re-evaluated on non-overlapping sets of 5-Mb windows, and on non-overlapping 1-Mb windows as well, with similar results261.
Possible explanations for variation
What explains the correlation among these many measures of genome divergence? It seems unlikely that direct selection would account for variation and co-variation at such large scales (about 5 Mb) and involving abundant neutral sites taken from ancestral transposon relics. Selection against deleterious mutations can remove linked polymorphisms270,271, but it is not clear that such effects or related effects272 could extend to such large scales or to interspecies divergence over such large time periods273.
It seems more probable that these features reflect local variation in underlying mutation rate, caused by differences in DNA metabolism or chromosome physiology. The causative factors may include recombination-associated mutagenesis258,266, transcription-associated mutagenesis274, transposon-associated deletion and genomic rearrangement275,276,277,278, and replication timing279,280. Nuclear location may also be involved, including proximity to matrix attachment sites, heterochromatin, nuclear membrane, and origins of replication.
It is clear that the mammalian genome is evolving under the influence of non-uniform local forces. It remains an important challenge to unravel the mechanistic basis and evolutionary consequences of such variation.
Genetic variation among strains
To facilitate genetic mapping studies, it would be valuable to create a mouse genetic map based on SNPs. The use of SNPs would allow the generation of an even denser map, which would allow mouse geneticists to fully exploit the recombinational resolution that can be achieved in large crosses. A cross with 2,000 meioses divides the genome (with a genetic length of about 16 morgans) into approximately 32,000 distinct recombinational ‘bins’ and it would be convenient to have an even higher density of genetic markers available for fine-scale mapping. In addition, SNPs offer potential advantages in terms of automation and parallelism265,281,282.
Given a reference sequence of the B6 strain, it is straightforward to find SNPs relative to any other strain. One simply needs to generate random shotgun reads from the strain, align them to the reference sequence and search for high-quality sequence differences.
As a pilot project, we created initial SNP collections from three strains: 129S1/SvImJ (129), C3H/HeJ (C3H) and BALB/cByJ (BALB) (Table 18). So far we have identified 47,279 high-quality candidate SNPs between the 129 and B6 strains, 20,294 SNPs between C3H and B6 and 11,696 between BALB and B6. The initial SNP collection thus contains more than 79,000 SNPs. This total is expected to grow with deeper coverage and the inclusion of additional strains. We tested a random sample of 83 candidate SNPs by resequencing and found that all 83 were authentic, indicating that most of the candidate SNPs are true variants.
The average density of SNPs between B6 and each of the three strains was in the range 1 per 500–700 bp. The distribution of SNPs is highly non-uniform (consistent with earlier observations282). Some regions of the genome appear to be unusually rich in SNPs, whereas others are devoid of SNPs.
In an accompanying paper, Wade and colleagues283 analyse this non-uniform distribution of SNPs and demonstrate that genetic variation between strains occurs in a harlequin pattern of alternating blocks of either high or low SNP rate, typically extending more than 1 Mb. Genotyping of additional strains reveals that the SNPs largely represent alternative alleles from M. m. domesticus and M. m. musculus, and that the blocks probably represent the distinct segmental contributions of the two subspecies to existing laboratory mouse strains. Detailed knowledge of these blocks can thus allow reconstruction of the history and relationship among mouse strains. Furthermore, it can be used to perform association studies on mouse strains, by correlating differences in phenotype across multiple strains with the underlying block structure of genetic variation.
Implications for the laboratory mouse
The promise of genomics is the ability to connect phenotypes with genotypes for a wide variety of traits and to use the resulting molecular insights to develop new approaches for the cure and prevention of disease. The laboratory mouse occupies a central place in this vision, both as a prototype for all mammalian biology and as a well-characterized organism for modelling human disease states15,16,123. In this section, we briefly discuss ways in which the mouse genome sequence will accelerate biomedical progress in the future. Because the sequence has been made available in public databases in advance of publication, examples for many of the predictions can already be cited.
Positional cloning of genes for mendelian phenotypes
More than 1,000 spontaneously arising and radiation-induced mouse mutants causing heritable mendelian phenotypes are catalogued in the Mouse Genome Informatics (MGI) database (http://www.informatics.jax.org). Largely through positional cloning, the molecular defect is now known for about 200 of these mutants. The availability of an annotated mouse genome sequence now provides the most efficient tool yet in the gene hunter's toolkit. One can move directly from genetic mapping to identification of candidate genes, and the experimental process is reduced to PCR amplification and sequencing of exons and other conserved elements in the candidate interval. With this streamlined protocol, it is anticipated that many decades-old mouse mutants will be understood precisely at the DNA level in the near future. An example of how the draft genome sequence has already been successfully used is the recent identification of the mouse mutation ‘chocolate’ in the melanosome protein Rab38 (ref. 284).
The mouse genome sequence will be even more crucial in efforts to exploit the growing repertoire of mutant mice being generated by chemical mutagenesis with N-ethyl-N-nitrosurea (ENU) and other agents. At least ten large-scale ENU mutagenesis centres have recently been established worldwide, focusing on dominant or recessive screens for a wide variety of viable, clinically relevant phenotypes15. Hundreds of new mutants with biochemical, development and behavioural phenotypes are being generated each year. For each mutant, identification of the molecular cause will require positional cloning.
Another means of generating mutants, the so-called ‘gene trap’ approach, uses a promoterless reporter construct for random insertion into the genome of embryonic stem cells. Expression of the reporter correlates with integration into a transcriptional unit, which is disrupted by the event and confers its tissue and developmental specificity to the reporter. Several large-scale gene-trap programmes are underway worldwide15. Availability of the genome sequence now makes the determination of the precise integration site in an interesting mutant an almost trivial exercise.
Identification of quantitative trait loci
The availability of more than 50 commonly used laboratory inbred strains of mice, each with its own phenotype for multiple continuously variable traits, has provided an important opportunity to map QTLs that underlie heritable phenotypic variation. A systematic initiative is currently underway285 to define parameters such as body weight, behavioural patterns, and disease susceptibility among a standard set of inbred lines, and to make these data freely available to the scientific community in the Mouse Phenome Database (www.jax.org/phenome). Appropriate crosses between such lines, followed by genotyping, will enable the mapping of QTLs, which can then be subjected to positional cloning. The degree of difficulty is substantially greater for a QTL cloning project than for a mendelian disorder, however, as the responsible intervals are usually much larger, the boundaries more difficult to delineate precisely, and the causative variant often much more subtle286. For these reasons, only a handful of the approximately 1,000 mapped QTLs have been identified at the molecular level. The availability of the mouse sequence should greatly improve the chances for future success.
Success in QTL identification will be enhanced if genetic mapping can be combined with genomic sequence, expression array data and proteomic data. Furthermore, the use of high-density SNP maps to identify blocks of ancestral identity among mouse strains and to correlate them with phenotypes may assist in the design of QTL experiments. The availability of BAC libraries from several strains will facilitate testing candidate genes for QTLs through the construction of transgenic mice287. The combination of multiple perspectives on genome sequence, variation and function should thus provide a powerful platform for revealing molecular mechanisms of phenotypic variation.
Creation of knockout and knockin mice
The wide application of homologous recombination in embryonic stem cells has provided a remarkable abundance of ‘custom’ mice with specifically engineered loss- or gain-of-function mutations in specific genes of biological or medical interest. Yet this remains a time-consuming process. The design of recombinant DNA constructs for injection has often been delayed by incomplete knowledge of gene structure, requiring tedious restriction mapping or sequencing, and occasionally giving rise to unsatisfying outcomes due to incorrect information. The availability of the mouse genome sequence will both speed the design of such constructs and reduce the likelihood of unfortunate choices. Furthermore, the long-range continuity of the sequence should facilitate the generation of models of contiguous gene-deletion syndromes.
Creation of transgenic animals
For many transgenic experiments, it is important to maintain copy-dependent, tissue-specific expression of the transgene. This is most readily accomplished through BAC transgenesis. The availability of a deep, end-sequenced BAC library from the B6 strain mapped to the genome sequence now makes it straightforward to obtain a desired gene in a BAC for such experiments; end-sequenced BAC libraries from other strains should be available in the future. BACs also provide the ability to make mutant alleles with relative ease, by taking advantage of powerful genetic engineering techniques for custom mutagenesis in the Escherichia coli host.
Applications to cancer
The mouse genome sequence also has powerful applications to the molecular characterization of the somatic mutations that result in neoplasia. High-density SNP mapping to identify loss of heterozygosity288,289, combined with comparative genomic hybridization using cDNA or BAC arrays290,291, can be used to identify chromosomal segments showing loss or gain of copy number in particular tumour types. The combination of such approaches with expression arrays that include all mouse genes should further enhance the ability to pinpoint the molecular lesions that result in carcinogenesis. Full sequencing of all the exons and regulatory regions of known tumour suppressors, oncogenes, and other candidate genes can now be contemplated, as has been initiated in a few centres for human tumours292.
As a specific example of the use of the draft sequence for oncogene discovery, several groups recently used retroviral infection in mice to recover new cancer susceptibility loci. The ability to compare rapidly retrieved sequence tags to the draft genome sequence greatly accelerated the process of cancer gene discovery293,294,295.
Making better mouse models
Not all mouse models replicate the human phenotype in the expected way. The availability of the full human and mouse sequences provides an opportunity to anticipate these differences, and perhaps to compensate for them. In some instances, it may turn out that the murine mutation did not reside in the true orthologue of the human disease gene. Alternatively, in a circumstance where the human genome contains only a single gene family member, but the mouse genome contains a paralogue as well as the orthologue, one can anticipate that knockout of the orthologue alone may give a much milder phenotype (or none at all). Such was the case, for instance, with the occulocerebrorenal syndrome described by Lowe and colleagues296. Creating double knockout mice may then provide a closer match to the human disease phenotype.
Understanding gene regulation
Of the approximately 5% of windows of the mammalian genome that are under selection, most do not appear to code for protein. Much of this sequence is probably involved in the regulation of gene expression. It should be possible to pinpoint these regulatory elements more precisely with the availability of additional related genomes. However, mouse is likely to provide the most powerful experimental platform for generating and testing hypotheses about their function. An example is the recent demonstration, based on mouse–human sequence alignment followed by knockout manipulation, of several long-range locus control regions that affect expression of the Il4/Il13/Il5 cluster4.
Conclusion
The mouse provides a unique lens through which we can view ourselves. As the leading mammalian system for genetic research over the past century, it has provided a model for human physiology and disease, leading to major discoveries in such fields as immunology and metabolism. With the availability of the mouse genome sequence, it now provides a model and informs the study of our genome as well.
Comparative genome analysis is perhaps the most powerful tool for understanding biological function. Its power lies in the fact that evolution's crucible is a far more sensitive instrument than any other available to modern experimental science: a functional alteration that diminishes a mammal's fitness by one part in 104 is undetectable at the laboratory bench, but is lethal from the standpoint of evolution.
Comparative analysis of genomes should thus make it possible to discern, by virtue of evolutionary conservation, biological features that would otherwise escape our notice. In this way, it will play a crucial role in our understanding of the human genome and thereby help lay the foundation for biomedicine in the twenty-first century.
The initial sequence of the mouse genome reported here is merely a first step in this intellectual programme. The sequencing of many additional mammalian and other vertebrate genomes will be needed to extract the full information hidden within our chromosomes. Moreover, as we begin to understand the common elements shared among species, it may also become possible to approach the even harder challenge of identifying and understanding the functional differences that make each species unique.
Methods
Production of sequence reads
Paired-end reads from libraries with different insert sizes were produced as previously described1 using 384-well trays to ensure linkages.
Availability of sequence and assembly data
Unprocessed sequence reads are available from the NCBI trace archive (ftp://ftp.ncbi.nih.gov/pub/TraceDB/mus_musculus/). Raw assembly data (before removal of contaminants, anchoring to chromosomes, and addition of finished sequence) are available from the Whitehead Institute for Biomedical Research (WIBR) (ftp://wolfram.wi.mit.edu/pub/mouse_contigs/Mar10_02/). The released assembly MGSCv3 is available from Ensembl (http://www.ensembl.org/Mus_musculus/), NCBI (ftp://ftp.ncbi.nih.gov/genomes/M_musculus/MGSCv3_Release1/), UCSC (http://genome.ucsc.edu/downloads.html) and WIBR (ftp://wolfram.wi.mit.edu/pub/mouse_contigs/MGSC_V3/). (See Supplementary Information for detailed Methods.)
References
International Human Genome Sequencing Consortium Initial sequencing and analysis of the human genome. Nature 409, 860–921 (2001)
Venter, J. C. et al. The sequence of the human genome. Science 291, 1304–1351 (2001)
O'Brien, S. J. et al. The promise of comparative genomics in mammals. Science 286, 458–462, 479–481 (1999)
Loots, G. G. et al. Identification of a coordinate regulator of interleukins 4, 13, and 5 by cross-species sequence comparisons. Science 288, 136–140 (2000)
Pennacchio, L. A. & Rubin, E. M. Genomic strategies to identify mammalian regulatory sequences. Nature Rev. Genet. 2, 100–109 (2001)
Oeltjen, J. C. et al. Large-scale comparative sequence analysis of the human and murine Bruton's tyrosine kinase loci reveals conserved regulatory domains. Genome Res. 7, 315–329 (1997)
Ellsworth, R. E. et al. Comparative genomic sequence analysis of the human and mouse cystic fibrosis transmembrane conductance regulator genes. Proc. Natl Acad. Sci. USA 97, 1172–1177 (2000)
Mallon, A. M. et al. Comparative genome sequence analysis of the Bpa/Str region in mouse and man. Genome Res. 10, 758–775 (2000)
Dehal, P. et al. Human chromosome 19 and related regions in mouse: conservative and lineage-specific evolution. Science 293, 104–111 (2001)
DeSilva, U. et al. Generation and comparative analysis of approximately 3.3 Mb of mouse genomic sequence orthologous to the region of human chromosome 7q11.23 implicated in Williams syndrome. Genome Res. 12, 3–15 (2002)
Toyoda, A. et al. Comparative genomic sequence analysis of the human chromosome 21 down syndrome critical region. Genome Res. 12, 1323–1332 (2002)
Ansari-Lari, M. A. et al. Comparative sequence analysis of a gene-rich cluster at human chromosome 12p13 and its syntenic region in mouse chromosome 6. Genome Res. 8, 29–40 (1998)
Lercher, M. J., Williams, E. J. & Hurst, L. D. Local similarity in evolutionary rates extends over whole chromosomes in human-rodent and mouse-rat comparisons: implications for understanding the mechanistic basis of the male mutation bias. Mol. Biol. Evol. 18, 2032–2039 (2001)
Makalowski, W. & Boguski, M. S. Evolutionary parameters of the transcribed mammalian genome: an analysis of 2,820 orthologous rodent and human sequences. Proc. Natl Acad. Sci. USA 95, 9407–9412 (1998)
Rossant, J. & McKerlie, C. Mouse-based phenogenomics for modelling human disease. Trends Mol. Med. 7, 502–507 (2001)
Paigen, K. A miracle enough: the power of mice. Nature Med. 1, 215–220 (1995)
Hogan, B., Beddington, R., Costantini, F. & Lacy, E. Manipulating the Mouse Embryo: A Laboratory Manual (Cold Spring Harbor Laboratory Press, Woodbury, New York, 1994)
Joyner, A. L. Gene Targeting: A Practical Approach (Oxford Univ. Press, New York, 1999)
Copeland, N. G., Jenkins, N. A. & Court, D. L. Recombineering: a powerful new tool for mouse functional genomics. Nature Rev. Genet. 2, 769–779 (2001)
Yu, Y. & Bradley, A. Engineering chromosomal rearrangements in mice. Nature Rev. Genet. 2, 780–790 (2001)
Bucan, M. & Abel, T. The mouse: genetics meets behaviour. Nature Rev. Genet. 3, 114–123 (2002)
Silver, L. M. Mouse Genetics: Concepts and Practice (Oxford Univ. Press, New York, 1995)
Bromham, L., Phillips, M. J. & Penny, D. Growing up with dinosaurs: molecular dates and the mammalian radiation. Trends Ecol. Evol. 14, 113–118 (1999)
Nei, M., Xu, P. & Glazko, G. Estimation of divergence times from multiprotein sequences for a few mammalian species and several distantly related organisms. Proc. Natl Acad. Sci. USA 98, 2497–2502 (2001)
Kumar, S. & Hedges, S. B. A molecular timescale for vertebrate evolution. Nature 392, 917–920 (1998)
Madsen, O. et al. Parallel adaptive radiations in two major clades of placental mammals. Nature 409, 610–614 (2001)
Murphy, W. J. et al. Molecular phylogenetics and the origins of placental mammals. Nature 409, 614–618 (2001)
Keeler, C. E. The Laboratory Mouse: Its Origin, Heredity and Culture (Harvard Univ. Press, Cambridge, Massachusetts, 1931)
Morse, H. The Mouse in Biomedical Research (eds Foster, H. L., Small, J. D. & Fox, J. G.) 1–16 (Academic, New York, 1981)
Morse, H. C. Origins of Inbred Mice (ed. Morse, H. C.) 1–21 (Academic, New York, 1978)
Haldane, J. B. S., Sprunt, A. D. & Haldane, N. M. Reduplication in mice. J. Genet. 5, 133–135 (1915)
Botstein, D., White, R. L., Skolnick, M. & Davis, R. W. Construction of a genetic linkage map in man using restriction fragment length polymorphisms. Am. J. Hum. Genet. 32, 314–331 (1980)
Dietrich, W. et al. Genetic Maps (ed. O'Brien, S.) 4.110–4.142, (1992)
Dietrich, W. F. et al. A comprehensive genetic map of the mouse genome. Nature 380, 149–152 (1996)
Love, J. M., Knight, A. M., McAleer, M. A. & Todd, J. A. Towards construction of a high resolution map of the mouse genome using PCR-analysed microsatellites. Nucleic Acids Res. 18, 4123–4130 (1990)
Weber, J. L. & May, P. E. Abundant class of human DNA polymorphisms which can be typed using the polymerase chain reaction. Am. J. Hum. Genet. 44, 388–396 (1989)
Hudson, T. J. et al. A radiation hybrid map of mouse genes. Nature Genet. 29, 201–205 (2001)
Van Etten, W. J. et al. Radiation hybrid map of the mouse genome. Nature Genet. 22, 384–387 (1999)
Nusbaum, C. et al. A YAC-based physical map of the mouse genome. Nature Genet. 22, 388–393 (1999)
Marra, M. et al. An encyclopedia of mouse genes. Nature Genet. 21, 191–194 (1999)
Kawai, J. et al. Functional annotation of a full-length mouse cDNA collection. Nature 409, 685–690 (2001)
Strausberg, R. L., Feingold, E. A., Klausner, R. D. & Collins, F. S. The mammalian gene collection. Science 286, 455–457 (1999)
Osoegawa, K. et al. Bacterial artificial chromosome libraries for mouse sequencing and functional analysis. Genome Res. 10, 116–128 (2000)
Gregory, S. G. et al. A physical map of the mouse genome. Nature 418, 743–750 (2002)
Mural, R. J. et al. A comparison of whole-genome shotgun-derived mouse chromosome 16 and the human genome. Science 296, 1661–1671 (2002)
Green, E. D. Strategies for the systematic sequencing of complex genomes. Nature Rev. Genet. 2, 573–583 (2001)
Edwards, A. et al. Automated DNA sequencing of the human HPRT locus. Genomics 6, 593–608 (1990)
Huson, D. H. et al. Design of a compartmentalized shotgun assembler for the human genome. Bioinformatics 17, S132–S139 (2001)
Analysis of the genome sequence of the flowering plant Arabidopsis thaliana. Nature 408, 796–815 (2000)
Adams, M. D. et al. The genome sequence of Drosophila melanogaster. Science 287, 2185–2195 (2000)
Yu, J. et al. A draft sequence of the rice genome. Science 296, 79–92 (2002)
Battey, J., Jordan, E., Cox, D. & Dove, W. An action plan for mouse genomics. Nature Genet. 21, 73–75 (1999)
Kuroda-Kawaguchi, T. et al. The AZFc region of the Y chromosome features massive palindromes and uniform recurrent deletions in infertile men. Nature Genet. 29, 279–286 (2001)
Zhao, S. et al. Mouse BAC ends quality assessment and sequence analyses. Genome Res. 11, 1736–1745 (2001)
Ewing, B. & Green, P. Analysis of expressed sequence tags indicates 35,000 human genes. Nature Genet. 25, 232–234 (2000)
Batzoglou, S. et al. ARACHNE: a whole-genome shotgun assembler. Genome Res. 12, 177–189 (2002)
Jaffe, D. B. et al. Whole-genome sequence assembly for mammalian genomes: Arachne 2. Genome Res. (in the press)
Mullikin, J. & Ning, Z. The Phusion Assembler. Genome Res. (in the press)
Bailey, J. A. et al. Recent segmental duplications in the human genome. Science 297, 1003–1007 (2002)
Traut, W., Winking, H. & Adolph, S. An extra segment in chromosome 1 of wild Mus musculus: a C-band positive homogeneously staining region. Cytogenet. Cell Genet. 38, 290–297 (1984)
Weichenhan, D. et al. Source and component genes of a 6-200 Mb gene cluster in the house mouse. Mamm. Genome 12, 590–594 (2001)
Purmann, L., Plass, C., Gruneberg, M., Winking, H. & Traut, W. A long-range repeat cluster in chromosome 1 of the house mouse, Mus musculus, and its relation to a germline homogeneously staining region. Genomics 12, 80–88 (1992)
Wong, A. K. & Rattner, J. B. Sequence organization and cytological localization of the minor satellite of mouse. Nucleic Acids Res. 16, 11645–11661 (1988)
Joseph, A., Mitchell, A. R. & Miller, O. J. The organization of the mouse satellite DNA at centromeres. Exp. Cell Res. 183, 494–500 (1989)
Davisson, M. T. & Roderick, T. H. Genetic Variants and Strains of the Laboratory Mouse (eds Lyon, M. F. & Searle, A. G.) 416–427 (Oxford Univ. Press, Oxford, 1989)
Mouse Genome Sequencing Consortium Progress in sequencing the mouse genome. Genesis 31, 137–141 (2001)
Clark, F. H. Inheritance and linkage relations of mutant characteristics in the deermouse. Contrib. Lab. Vert. Biol. 7, 1–11 (1938)
Castle, W. W. Observations of the occurrence of linkage in rats and mice. Car. Inst. Wash. Pub. 288, 29–36 (1919)
Lalley, P. A., Minna, J. D. & Francke, U. Conservation of autosomal gene synteny groups in mouse and man. Nature 274, 160–163 (1978)
Nadeau, J. H. & Taylor, B. A. Lengths of chromosomal segments conserved since divergence of man and mouse. Proc. Natl Acad. Sci. USA 81, 814–818 (1984)
Ma, B., Tromp, J. & Li, M. PatternHunter: faster and more sensitive homology search. Bioinformatics 18, 440–445 (2002)
Ohno, S. Sex Chromosomes and Sex-Linked Genes (Springer, Berlin, 1996)
Sturtevant, A. H. & Beadle, G. W. The relations of inversions in the X chromosome of Drosophila melanogaster to crossing over and disjunction. Genetics 21, 554–604 (1936)
Ranz, J. M., Casals, F. & Ruiz, A. How malleable is the eukaryotic genome? Extreme rate of chromosomal rearrangement in the genus Drosophila. Genome Res. 11, 230–239 (2001)
Nadeau, J. H. & Sankoff, D. The lengths of undiscovered conserved segments in comparative maps. Mamm. Genome 9, 491–495 (1998)
Ferretti, V., Nadeau, J. H. & Sankoff, D. Combinatorial Pattern Matching, 7th Annual Symposium (eds Hirschberg, D. & Myers, G.) 159–167 (Springer, Berlin, 1996)
Bourque, G. & Pevzner, P. A. Genome-scale evolution: reconstructing gene orders in the ancestral species. Genome Res. 12, 26–36 (2002)
Thiery, J. P., Macaya, G. & Bernardi, G. An analysis of eukaryotic genomes by density gradient centrifugation. J. Mol. Biol. 108, 219–235 (1976)
Salinas, J., Zerial, M., Filipski, J. & Bernardi, G. Gene distribution and nucleotide sequence organization in the mouse genome. Eur. J. Biochem. 160, 469–478 (1986)
Sabeur, G., Macaya, G., Kadi, F. & Bernardi, G. The isochore patterns of mammalian genomes and their phylogenetic implications. J. Mol. Evol. 37, 93–108 (1993)
Zerial, M., Salinas, J., Filipski, J. & Bernardi, G. Gene distribution and nucleotide sequence organization in the human genome. Eur. J. Biochem. 160, 479–485 (1986)
Mouchiroud, D., Fichant, G. & Bernardi, G. Compositional compartmentalization and gene composition in the genome of vertebrates. J. Mol. Evol. 26, 198–204 (1987)
Mouchiroud, D., Gautier, C. & Bernardi, G. The compositional distribution of coding sequences and DNA molecules in humans and murids. J. Mol. Evol. 27, 311–320 (1988)
Mouchiroud, D. & Gautier, C. Codon usage changes and sequence dissimilarity between human and rat. J. Mol. Evol. 31, 81–91 (1990)
Robinson, M., Gautier, C. & Mouchiroud, D. Evolution of isochores in rodents. Mol. Biol. Evol. 14, 823–828 (1997)
Bernardi, G. et al. The mosaic genome of warm-blooded vertebrates. Science 228, 953–958 (1985)
Mouchiroud, D. et al. The distribution of genes in the human genome. Gene 100, 181–187 (1991)
Zoubak, S., Clay, O. & Bernardi, G. The gene distribution of the human genome. Gene 174, 95–102 (1996)
Saccone, S., Pavlicek, A., Federico, C., Paces, J. & Bernard, G. Genes, isochores and bands in human chromosomes 21 and 22. Chromosome Res. 9, 533–539 (2001)
Bernardi, G. Compositional constraints and genome evolution. J. Mol. Evol. 24, 1–11 (1986)
Bernardi, G., Mouchiroud, D. & Gautier, C. Compositional patterns in vertebrate genomes: conservation and change in evolution. J. Mol. Evol. 28, 7–18 (1988)
Wolfe, K. H., Sharp, P. M. & Li, W. H. Mutation rates differ among regions of the mammalian genome. Nature 337, 283–285 (1989)
Sueoka, N. Directional mutation pressure and neutral molecular evolution. Proc. Natl Acad. Sci. USA 85, 2653–2657 (1988)
Sueoka, N. On the genetic basis of variation and heterogeneity of DNA base composition. Proc. Natl Acad. Sci. USA 48, 582–592 (1962)
Bird, A. P. DNA methylation and the frequency of CpG in animal DNA. Nucleic Acids Res. 8, 1499–1504 (1980)
Larsen, F., Gundersen, G., Lopez, R. & Prydz, H. CpG islands as gene markers in the human genome. Genomics 13, 1095–1107 (1992)
Gardiner-Garden, M. & Frommer, M. CpG islands in vertebrate genomes. J. Mol. Biol. 196, 261–282 (1987)
Antequera, F. & Bird, A. Number of CpG islands and genes in human and mouse. Proc. Natl Acad. Sci. USA 90, 11995–11999 (1993)
Adams, R. L. & Eason, R. Increased G+C content of DNA stabilizes methyl CpG dinucleotides. Nucleic Acids Res. 12, 5869–5877 (1984)
Smit, A. F. Interspersed repeats and other mementos of transposable elements in mammalian genomes. Curr. Opin. Genet. Dev. 9, 657–663 (1999)
Laird, C. D., McConaughy, B. L. & McCarthy, B. J. Rate of fixation of nucleotide substitutions in evolution. Nature 224, 149–154 (1969)
Kohne, D. E. Evolution of higher-organism DNA. Q. Rev. Biophys. 3, 327–375 (1970)
Goodman, M., Barnabas, J., Matsuda, G. & Moore, G. W. Molecular evolution in the descent of man. Nature 233, 604–613 (1971)
Kumar, S. & Subramanian, S. Mutation rates in mammalian genomes. Proc. Natl Acad. Sci. USA 99, 803–808 (2002)
Easteal, S., Collet, C. & Betty, D. The Mammalian Molecular Clock (Landes, Austin, Texas, 1995)
Li, W. H., Ellsworth, D. L., Krushkal, J., Chang, B. H. & Hewett-Emmett, D. Rates of nucleotide substitution in primates and rodents and the generation-time effect hypothesis. Mol. Phylogenet. Evol. 5, 182–187 (1996)
Martin, A. P. & Palumbi, S. R. Body size, metabolic rate, generation time, and the molecular clock. Proc. Natl Acad. Sci. USA 90, 4087–4091 (1993)
Bromham, L. Molecular clocks in reptiles: life history influences rate of molecular evolution. Mol. Biol. Evol. 19, 302–309 (2002)
Wu, C. I. & Li, W. H. Evidence for higher rates of nucleotide substitution in rodents than in man. Proc. Natl Acad. Sci. USA 82, 1741–1745 (1985)
Smit, A. F., Toth, G., Riggs, A. D. & Jurka, J. Ancestral, mammalian-wide subfamilies of LINE-1 repetitive sequences. J. Mol. Biol. 246, 401–417 (1995)
Adey, N. B. et al. Rodent L1 evolution has been driven by a single dominant lineage that has repeatedly acquired new transcriptional regulatory sequences. Mol. Biol. Evol. 11, 778–789 (1994)
Mears, M. L. & Hutchison, C. A. III The evolution of modern lineages of mouse L1 elements. J. Mol. Evol. 52, 51–62 (2001)
Goodier, J. L., Ostertag, E. M., Du, K. & Kazazian, H. H. Jr A novel active L1 retrotransposon subfamily in the mouse. Genome Res. 11, 1677–1685 (2001)
Hardies, S. C. et al. LINE-1 (L1) lineages in the mouse. Mol. Biol. Evol. 17, 616–628 (2000)
Ohshima, K., Hamada, M., Terai, Y. & Okada, N. The 3′ ends of tRNA-derived short interspersed repetitive elements are derived from the 3′ ends of long interspersed repetitive elements. Mol. Cell Biol. 16, 3756–3764 (1996)
Smit, A. F. The origin of interspersed repeats in the human genome. Curr. Opin. Genet. Dev. 6, 743–748 (1996)
Quentin, Y. A master sequence related to a free left Alu monomer (FLAM) at the origin of the B1 family in rodent genomes. Nucleic Acids Res. 22, 2222–2227 (1994)
Kim, J. & Deininger, P. L. Recent amplification of rat ID sequences. J. Mol. Biol. 261, 322–327 (1996)
Lee, I. Y. et al. Complete genomic sequence and analysis of the prion protein gene region from three mammalian species. Genome Res. 8, 1022–1037 (1998)
Serdobova, I. M. & Kramerov, D. A. Short retroposons of the B2 superfamily: evolution and application for the study of rodent phylogeny. J. Mol. Evol. 46, 202–214 (1998)
Coffin, J. M., Hughes, S. H. & Varmus, H. E. (eds) Retroviruses (Cold Spring Harbor Laboratory Press, Cold Spring Harbor, New York, 1997)
Smit, A. F. Identification of a new, abundant superfamily of mammalian LTR- transposons. Nucleic Acids Res. 21, 1863–1872 (1993)
Hamilton, B. A. & Frankel, W. N. Of mice and genome sequence. Cell 107, 13–16 (2001)
Turner, G. et al. Insertional polymorphisms of full-length endogenous retroviruses in humans. Curr. Biol. 11, 1531–1535 (2001)
Kidwell, M. G. Horizontal transfer. Curr. Opin. Genet. Dev. 2, 868–873 (1992)
Feng, Q., Moran, J. V., Kazazian, H. H. Jr & Boeke, J. D. Human L1 retrotransposon encodes a conserved endonuclease required for retrotransposition. Cell 87, 905–916 (1996)
Jurka, J. Sequence patterns indicate an enzymatic involvement in integration of mammalian retroposons. Proc. Natl Acad. Sci. USA 94, 1872–1877 (1997)
Bernardi, G. The isochore organization of the human genome. Annu. Rev. Genet. 23, 637–661 (1989)
Holmquist, G. P. Chromosome bands, their chromatin flavors, and their functional features. Am. J. Hum. Genet. 51, 17–37 (1992)
Korenberg, J. R. & Rykowski, M. C. Human genome organization: Alu, lines, and the molecular structure of metaphase chromosome bands. Cell 53, 391–400 (1988)
Boyle, A. L., Ballard, S. G. & Ward, D. C. Differential distribution of long and short interspersed element sequences in the mouse genome: chromosome karyotyping by fluorescence in situ hybridization. Proc. Natl Acad. Sci. USA 87, 7757–7761 (1990)
Lyon, M. F. X-chromosome inactivation: a repeat hypothesis. Cytogenet. Cell Genet. 80, 133–137 (1998)
Bailey, J. A., Carrel, L., Chakravarti, A. & Eichler, E. E. Molecular evidence for a relationship between LINE-1 elements and X chromosome inactivation: the Lyon repeat hypothesis. Proc. Natl Acad. Sci. USA 97, 6634–6639 (2000)
Boissinot, S. & Furano, A. V. Adaptive evolution in LINE-1 retrotransposons. Mol. Biol. Evol. 18, 2186–2194 (2001)
Beckman, J. S. & Weber, J. L. Survey of human and rat microsatellites. Genomics 12, 627–631 (1992)
Toth, G., Gaspari, Z. & Jurka, J. Microsatellites in different eukaryotic genomes: survey and analysis. Genome Res. 10, 967–981 (2000)
Kruglyak, S., Durrett, R. T., Schug, M. D. & Aquadro, C. F. Equilibrium distributions of microsatellite repeat length resulting from a balance between slippage events and point mutations. Proc. Natl Acad. Sci. USA 95, 10774–10778 (1998)
Santibanez-Koref, M. F., Gangeswaran, R. & Hancock, J. M. A relationship between lengths of microsatellites and nearby substitution rates in mammalian genomes. Mol. Biol. Evol. 18, 2119–2123 (2001)
Dunham, I. et al. The DNA sequence of human chromosome 22. Nature 402, 489–495 (1999)
Hattori, M. et al. The DNA sequence of human chromosome 21. Nature 405, 311–319 (2000)
Roest Crollius, H. et al. Estimate of human gene number provided by genome-wide analysis using Tetraodon nigroviridis DNA sequence. Nature Genet. 25, 235–238 (2000)
Hubbard, T. et al. The Ensembl genome database project. Nucleic Acids Res. 30, 38–41 (2002)
Kulp, D., Haussler, D., Reese, M. G. & Eeckman, F. H. Integrating database homology in a probabilistic gene structure model. Pac. Symp. Biocomput. 232–244 (1997)
Birney, E. & Durbin, R. Using GeneWise in the Drosophila annotation experiment. Genome Res. 10, 547–548 (2000)
Burge, C. & Karlin, S. Prediction of complete gene structures in human genomic DNA. J. Mol. Biol. 268, 78–94 (1997)
Hogenesch, J. B. et al. A comparison of the Celera and Ensembl predicted gene sets reveals little overlap in novel genes. Cell 106, 413–415 (2001)
Saha, S. et al. Using the transcriptome to annotate the genome. Nature Biotechnol. 20, 508–512 (2002)
Daly, M. J. Estimating the human gene count. Cell 109, 283–284 (2002)
Kapranov, P. et al. Large-scale transcriptional activity in chromosomes 21 and 22. Science 296, 916–919 (2002)
The FANTOM Consortium and the RIKEN Genome Exploration Research Group Phase I & II Team. Analysis of the mouse transcriptome based on functional annotation of 60,770 full-length cDNAs. Nature 420, 563–573 (2002)
Pruitt, K. D. & Maglott, D. R. RefSeq and LocusLink: NCBI gene-centered resources. Nucleic Acids Res. 29, 137–140 (2001)
Steimle, V. et al. A novel DNA-binding regulatory factor is mutated in primary MHC class II deficiency (bare lymphocyte syndrome). Genes Dev. 9, 1021–1032 (1995)
Sun, H., Tsunenari, T., Yau, K. W. & Nathans, J. The vitelliform macular dystrophy protein defines a new family of chloride channels. Proc. Natl Acad. Sci. USA 99, 4008–4013 (2002)
Yasunaga, S. et al. A mutation in OTOF, encoding otoferlin, a FER-1-like protein, causes DFNB9, a nonsyndromic form of deafness. Nature Genet. 21, 363–369 (1999)
den Hollander, A. I. et al. Leber congenital amaurosis and retinitis pigmentosa with Coats-like exudative vasculopathy are associated with mutations in the crumbs homologue 1 (CRB1) gene. Am. J. Hum. Genet. 69, 198–203 (2001)
den Hollander, A. I. et al. Mutations in a human homologue of Drosophila crumbs cause retinitis pigmentosa (RP12). Nature Genet. 23, 217–221 (1999)
Maeda, N. et al. Diet-induced insulin resistance in mice lacking adiponectin/ACRP30. Nature Med. 8, 731–737 (2002)
Clausen, B. E. et al. Residual MHC class II expression on mature dendritic cells and activated B cells in RFX5-deficient mice. Immunity 8, 143–155 (1998)
Garcia-Meunier, P., Etienne-Julan, M., Fort, P., Piechaczyk, M. & Bonhomme, F. Concerted evolution in the GAPDH family of retrotransposed pseudogenes. Mamm. Genome 4, 695–703 (1993)
Korf, I., Flicek, P., Duan, D. & Brent, M. R. Integrating genomic homology into gene structure prediction. Bioinformatics 17, S140–S148 (2001)
Wiehe, T., Gebauer-Jung, S., Mitchell-Olds, T. & Guigo, R. SGP-1: prediction and validation of homologous genes based on sequence alignments. Genome Res. 11, 1574–1583 (2001)
Alexandersson, M., Cawley, S. & Pachter, L. SLAM—cross-species GeneFinding and alignment with a generalized pair hidden Markov model. Genome Res. (in the press)
Reymond, A. et al. Human chromosome 21 gene expression atlas in the mouse. Nature 420, 582–586 (2002)
Blake, D. J., Weir, A., Newey, S. E. & Davies, K. E. Function and genetics of dystrophin-related proteins in muscle. Physiol. Rev. 82, 291–329 (2002)
Eddy, S. R. Non-coding RNA genes and the modern RNA world. Nature Rev. Genet. 2, 919–929 (2001)
Storz, G. An expanding universe of noncoding RNAs. Science 296, 1260–1263 (2002)
Eddy, S. R. Computational genomics of noncoding RNA genes. Cell 109, 137–140 (2002)
Lowe, T. M. & Eddy, S. R. tRNAscan-SE: a program for improved detection of transfer RNA genes in genomic sequence. Nucleic Acids Res. 25, 955–964 (1997)
Daniels, G. R. & Deininger, P. L. Repeat sequence families derived from mammalian tRNA genes. Nature 317, 819–822 (1985)
Lawrence, C., McDonnell, D. & Ramsey, W. Analysis of repetitive sequence elements containing tRNA-like sequences. Nucleic Acids Res. 13, 4239–4252 (1985)
Baron, C. & Bock, A. tRNA: Structure, Biosynthesis, and Function (eds Soll, D. & RajBhandary, U. L.) 529–544 (Am. Soc. Microbiol., Washington DC, 1995)
Crick, F. H. Codon–anticodon pairing: the wobble hypothesis. J. Mol. Biol. 19, 548–555 (1966)
Guthrie, C. & Abelson, J. The Molecular Biology of the Yeast Saccharomyces: Metabolism and Gene Expression (eds Strathern, J. N., Jones, E. W. & Broach, J. R.) 487–528 (Cold Spring Harbor Laboratory Press, Woodbury, New York, 1982)
Ponting, C. P. & Russell, R. R. The natural history of protein domains. Annu. Rev. Biophys. Biomol. Struct. 31, 45–71 (2002)
Lespinet, O., Wolf, Y. I., Koonin, E. V. & Aravind, L. The role of lineage-specific gene family expansion in the evolution of eukaryotes. Genome Res. 12, 1048–1059 (2002)
Ponting, C. P., Mott, R., Bork, P. & Copley, R. R. Novel protein domains and repeats in Drosophila melanogaster: insights into structure, function, and evolution. Genome Res. 11, 1996–2008 (2001)
Rubin, G. M. et al. Comparative genomics of the eukaryotes. Science 287, 2204–2215 (2000)
Altschul, S. F. et al. Gapped BLAST and PSI-BLAST: a new generation of protein database search programs. Nucleic Acids Res. 25, 3389–3402 (1997)
Zdobnov, E. M. & Apweiler, R. InterProScan—an integration platform for the signature-recognition methods in InterPro. Bioinformatics 17, 847–848 (2001)
Creating the gene ontology resource: design and implementation. Genome Res. 11, 1425–1433 (2001)
Makalowski, W. & Boguski, M. S. Synonymous and nonsynonymous substitution distances are correlated in mouse and rat genes. J. Mol. Evol. 47, 119–121 (1998)
Hughes, A. L. & Nei, M. Pattern of nucleotide substitution at major histocompatibility complex class I loci reveals overdominant selection. Nature 335, 167–170 (1988)
Yang, Z. & Nielsen, R. Estimating synonymous and nonsynonymous substitution rates under realistic evolutionary models. Mol. Biol. Evol. 17, 32–43 (2000)
Nekrutenko, A., Makova, K. D. & Li, W. H. The K(A)/K(S) ratio test for assessing the protein-coding potential of genomic regions: an empirical and simulation study. Genome Res. 12, 198–202 (2002)
Sharp, P. M. In search of molecular darwinism. Nature 385, 111–112 (1997)
Letunic, I. et al. Recent improvements to the SMART domain-based sequence annotation resource. Nucleic Acids Res. 30, 242–244 (2002)
Mott, R., Schultz, J., Bork, P. & Ponting, C. P. Predicting protein cellular localization using a domain projection method. Genome Res. 12, 1168–1174 (2002)
Hurst, L. D. & Smith, N. G. Do essential genes evolve slowly? Curr. Biol. 9, 747–750 (1999)
Goodstadt, L. & Ponting, C. P. Sequence variation and disease in the wake of the draft human genome. Hum. Mol. Genet. 10, 2209–2214 (2001)
Bairoch, A. & Apweiler, R. The SWISS-PROT protein sequence database and its supplement TrEMBL in 2000. Nucleic Acids Res. 28, 45–48 (2000)
Polymeropoulos, M. H. et al. Mutation in the alpha-synuclein gene identified in families with Parkinson's disease. Science 276, 2045–2047 (1997)
Fredman, D. et al. HGVbase: a human sequence variation database emphasizing data quality and a broad spectrum of data sources. Nucleic Acids Res. 30, 387–391 (2002)
Young, J. M. et al. Different evolutionary processes shaped the mouse and human olfactory receptor gene families. Hum. Mol. Genet. 11, 535–546 (2002)
Zhang, X. & Firestein, S. The olfactory receptor gene superfamily of the mouse. Nature Neurosci. 5, 124–133 (2002)
Glusman, G., Yanai, I., Rubin, I. & Lancet, D. The complete human olfactory subgenome. Genome Res. 11, 685–702 (2001)
Rouquier, S. et al. Distribution of olfactory receptor genes in the human genome. Nature Genet. 18, 243–250 (1998)
Del Punta, K. et al. Deficient pheromone responses in mice lacking a cluster of vomeronasal receptor genes. Nature 419, 70–74 (2002)
Nelson, D. R. Cytochrome P450 and the individuality of species. Arch. Biochem. Biophys. 369, 1–10 (1999)
Lane, R. P. et al. Genomic analysis of orthologous mouse and human olfactory receptor loci. Proc. Natl Acad. Sci. USA 98, 7390–7395 (2001)
Rossant, J. & Cross, J. C. Placental development: lessons from mouse mutants. Nature Rev. Genet. 2, 538–548 (2001)
Georgiades, P., Ferguson-Smith, A. C. & Burton, G. J. Comparative developmental anatomy of the murine and human definitive placentae. Placenta 23, 3–19 (2002)
Deussing, J. et al. Identification and characterization of a dense cluster of placenta- specific cysteine peptidase genes and related genes on mouse chromosome 13. Genomics 79, 225–240 (2002)
Afonso, S., Tovar, C., Romagnano, L. & Babiarz, B. Control and expression of cystatin C by mouse decidual cultures. Mol. Reprod. Dev. 61, 155–163 (2002)
Sutton, K. A. & Wilkinson, M. F. The rapidly evolving Pem homeobox gene and Agtr2, Ant2, and Lamp2 are closely linked in the proximal region of the mouse X chromosome. Genomics 45, 447–450 (1997)
Wilkinson, M. F., Kleeman, J., Richards, J. & MacLeod, C. L. A novel oncofetal gene is expressed in a stage-specific manner in murine embryonic development. Dev. Biol. 141, 451–455 (1990)
Han, Y. J., Park, A. R., Sung, D. Y. & Chun, J. Y. Psx, a novel murine homeobox gene expressed in placenta. Gene 207, 159–166 (1998)
Chun, J. Y., Han, Y. J. & Ahn, K. Y. Psx homeobox gene is X-linked and specifically expressed in trophoblast cells of mouse placenta. Dev. Dyn. 216, 257–266 (1999)
Takasaki, N., McIsaac, R. & Dean, J. Gpbox (Psx2), a homeobox gene preferentially expressed in female germ cells at the onset of sexual dimorphism in mice. Dev. Biol. 223, 181–193 (2000)
Lundwall, A. & Lazure, C. A novel gene family encoding proteins with highly differing structure because of a rapidly evolving exon. FEBS Lett. 374, 53–56 (1995)
Simon, A. M., Veyssiere, G. & Jean, C. Structure and sequence of a mouse gene encoding an androgen-regulated protein: a new member of the seminal vesicle secretory protein family. J. Mol. Endocrinol. 15, 305–316 (1995)
Morel, L. et al. Mouse seminal vesicle secretory protein of 99 amino acids (MSVSP99): characterization and hormonal and developmental regulation. J. Androl. 22, 549–557 (2001)
Linzer, D. I. & Fisher, S. J. The placenta and the prolactin family of hormones: regulation of the physiology of pregnancy. Mol. Endocrinol. 13, 837–840 (1999)
Huang, Y. H., Chu, S. T. & Chen, Y. H. A seminal vesicle autoantigen of mouse is able to suppress sperm capacitation-related events stimulated by serum albumin. Biol. Reprod. 63, 1562–1566 (2000)
Yoshida, M., Kaneko, M., Kurachi, H. & Osawa, M. Identification of two rodent genes encoding homologues to seminal vesicle autoantigen: a gene family including the gene for prolactin-inducible protein. Biochem. Biophys. Res. Commun. 281, 94–100 (2001)
Bain, P. A., Yoo, M., Clarke, T., Hammond, S. H. & Payne, A. H. Multiple forms of mouse 3 beta-hydroxysteroid dehydrogenase/delta 5-delta 4 isomerase and differential expression in gonads, adrenal glands, liver, and kidneys of both sexes. Proc. Natl Acad. Sci. USA 88, 8870–8874 (1991)
Payne, A. H., Abbaszade, I. G., Clarke, T. R., Bain, P. A. & Park, C. H. The multiple murine 3 beta-hydroxysteroid dehydrogenase isoforms: structure, function, and tissue- and developmentally specific expression. Steroids 62, 169–175 (1997)
Blume, N. et al. Characterization of Cyp2d22, a novel cytochrome P450 expressed in mouse mammary cells. Arch. Biochem. Biophys. 381, 191–204 (2000)
Lakso, M., Masaki, R., Noshiro, M. & Negishi, M. Structures and characterization of sex-specific mouse cytochrome P-450 genes as members within a large family. Duplication boundary and evolution. Eur. J. Biochem. 195, 477–486 (1991)
Tegoni, M. et al. Mammalian odorant binding proteins. Biochim. Biophys. Acta 1482, 229–240 (2000)
Miyawaki, A., Matsushita, F., Ryo, Y. & Mikoshiba, K. Possible pheromone-carrier function of two lipocalin proteins in the vomeronasal organ. EMBO J. 13, 5835–5842 (1994)
Karn, R. C. & Nachman, M. W. Reduced nucleotide variability at an androgen-binding protein locus (Abpa) in house mice: evidence for positive natural selection. Mol. Biol. Evol. 16, 1192–1197 (1999)
Karn, R. C., Orth, A., Bonhomme, F. & Boursot, P. The complex history of a gene proposed to participate in a sexual isolation mechanism in house mice. Mol. Biol. Evol. 19, 462–471 (2002)
Singer, A. G., Macrides, F., Clancy, A. N. & Agosta, W. C. Purification and analysis of a proteinaceous aphrodisiac pheromone from hamster vaginal discharge. J. Biol. Chem. 261, 13323–13326 (1986)
Zhang, J., Dyer, K. D. & Rosenberg, H. F. Evolution of the rodent eosinophil-associated RNase gene family by rapid gene sorting and positive selection. Proc. Natl Acad. Sci. USA 97, 4701–4706 (2000)
Natarajan, K., Dimasi, N., Wang, J., Margulies, D. H. & Mariuzza, R. A. MHC class I recognition by Ly49 natural killer cell receptors. Mol. Immunol. 38, 1023–1027 (2002)
Natarajan, K., Dimasi, N., Wang, J., Mariuzza, R. A. & Margulies, D. H. Structure and function of natural killer cell receptors: multiple molecular solutions to self, nonself discrimination. Annu. Rev. Immunol. 20, 853–885 (2002)
Yeager, M. & Hughes, A. L. Evolution of the mammalian MHC: natural selection, recombination, and convergent evolution. Immunol. Rev. 167, 45–58 (1999)
Ichikawa, T., Itakura, T. & Negishi, M. Functional characterization of two cytochrome P-450s within the mouse, male-specific steroid 16 alpha-hydroxylase gene family: expression in mammalian cells and chimeric proteins. Biochemistry 28, 4779–4784 (1989)
Miao, Y. J., Subramaniam, N. & Carlson, D. M. cDNA cloning and characterization of rat salivary glycoproteins. Novel members of the proline-rich-protein multigene families. Eur. J. Biochem. 228, 343–350 (1995)
Whelan, S., Lio, P. & Goldman, N. Molecular phylogenetics: state-of-the-art methods for looking into the past. Trends Genet. 17, 262–272 (2001)
Taveré, S. Some probabilistic and statistical problems on the analysis of DNA sequences. Lec. Math. Life Sci. 17, 57–86 (1986)
Jareborg, N., Birney, E. & Durbin, R. Comparative analysis of noncoding regions of 77 orthologous mouse and human gene pairs. Genome Res. 9, 815–824 (1999)
Suzuki, Y. et al. Diverse transcriptional initiation revealed by fine, large-scale mapping of mRNA start sites. EMBO Rep. 2, 388–393 (2001)
Kozak, M. Do the 5′untranslated domains of human cDNAs challenge the rules for initiation of translation (or is it vice versa)? Genomics 70, 396–406 (2000)
Zhao, J., Hyman, L. & Moore, C. Formation of mRNA 3′ ends in eukaryotes: mechanism, regulation, and interrelationships with other steps in mRNA synthesis. Microbiol. Mol. Biol. Rev. 63, 405–445 (1999)
Batzoglou, S., Pachter, L., Mesirov, J. P., Berger, B. & Lander, E. S. Human and mouse gene structure: comparative analysis and application to exon prediction. Genome Res. 10, 950–958 (2000)
Ogata, H., Fujibuchi, W. & Kanehisa, M. The size differences among mammalian introns are due to the accumulation of small deletions. FEBS Lett. 390, 99–103 (1996)
Burge, C. B., Padgett, R. A. & Sharp, P. A. Evolutionary fates and origins of U12-type introns. Mol. Cell 2, 773–785 (1998)
Wasserman, W. W., Palumbo, M., Thompson, W., Fickett, J. W. & Lawrence, C. E. Human-mouse genome comparisons to locate regulatory sites. Nature Genet. 26, 225–228 (2000)
Loots, G. G., Ovcharenko, I., Pachter, L., Dubchak, I. & Rubin, E. M. rVista for comparative sequence-based discovery of functional transcription factor binding sites. Genome Res. 12, 832–839 (2002)
Krivan, W. & Wasserman, W. W. A predictive model for regulatory sequences directing liver-specific transcription. Genome Res. 11, 1559–1566 (2001)
Wasserman, W. W. & Fickett, J. W. Identification of regulatory regions which confer muscle-specific gene expression. J. Mol. Biol. 278, 167–181 (1998)
Dermitzakis, E. & Clark, A. Evolution of transcription factor binding sites in mammalian gene regulatory regions: conservation and turnover. Mol. Biol. Evol. 19, 1114–1121 (2002)
Ooi, G. T., Hurst, K. R., Poy, M. N., Rechler, M. M. & Boisclair, Y. R. Binding of STAT5a and STAT5b to a single element resembling a gamma-interferon-activated sequence mediates the growth hormone induction of the mouse acid-labile subunit promoter in liver cells. Mol. Endocrinol. 12, 675–687 (1998)
Suwanichkul, A., Boisclair, Y. R., Olne, R. C., Durham, S. K. & Powell, D. R. Conservation of a growth hormone-responsive promoter element in the human and mouse acid-labile subunit genes. Endocrinology 141, 833–838 (2000)
Campbell, S. M., Rosen, J. M., Hennighausen, L. G., Strech-Jurk, U. & Sippel, A. E. Comparison of the whey acidic protein genes of the rat and mouse. Nucleic Acids Res. 12, 8685–8697 (1984)
Dermitzakis, E. T. et al. Numerous potentially functional but non-genic conserved sequences on human chromosome 21. Nature 420, 578–582 (2002)
Koop, B. F. Human and rodent DNA sequence comparisons: a mosaic model of genomic evolution. Trends Genet. 11, 367–371 (1995)
DeBry, R. W. & Seldin, M. F. Human/mouse homology relationships. Genomics 33, 337–351 (1996)
Gottgens, B. et al. Long-range comparison of human and mouse SCL loci: localized regions of sensitivity to restriction endonucleases correspond precisely with peaks of conserved noncoding sequences. Genome Res. 11, 87–97 (2001)
Shiraishi, T. et al. Sequence conservation at human and mouse orthologous common fragile regions, FRA3B/FHIT and Fra14A2/Fhit. Proc. Natl Acad. Sci. USA 98, 5722–5727 (2001)
Wilson, M. D. et al. Comparative analysis of the gene-dense ACHE/TFR2 region on human chromosome 7q22 with the orthologous region on mouse chromosome 5. Nucleic Acids Res. 29, 1352–1365 (2001)
Hardison, R. C. Conserved noncoding sequences are reliable guides to regulatory elements. Trends Genet. 16, 369–372 (2000)
Chiaromonte, F. et al. Association between divergence and interspersed repeats in mammalian noncoding genomic DNA. Proc. Natl Acad. Sci. USA 98, 14503–14508 (2001)
Matassi, G., Sharp, P. M. & Gautier, C. Chromosomal location effects on gene sequence evolution in mammals. Curr. Biol. 9, 786–791 (1999)
Williams, E. J. & Hurst, L. D. The proteins of linked genes evolve at similar rates. Nature 407, 900–903 (2000)
Chen, F. C., Vallender, E. J., Wang, H., Tzeng, C. S. & Li, W. H. Genomic divergence between human and chimpanzee estimated from large-scale alignments of genomic sequences. J. Hered. 92, 481–489 (2001)
Lercher, M. J. & Hurst, L. D. Human SNP variability and mutation rate are higher in regions of high recombination. Trends Genet. 18, 337–340 (2002)
Castresana, J. Genes on human chromosome 19 show extreme divergence from the mouse orthologs and a high GC content. Nucleic Acids Res. 30, 1751–1756 (2002)
Smith, N. G. C., Webster, M. & Ellegren, H. Deterministic mutation rate variation in the human genome. Genome Res. 12, 1350–1356 (2002)
Hardison, R. et al. Co-variation in frequencies of substitution, deletion, transposition and recombination during eutherian evolution. Genome Res. (in the press)
Bernardi, G. The human genome: organization and evolutionary history. Ann. Rev. Genet. 23, 637–661 (1995)
Hurst, L. D. & Willliams, E. J. B. Covarication of GC content and the silent site substitution rate in rodents: implications for methodology and for the evolution of isochores. Gene 261, 107–114 (2000)
Bernardi, G. Misunderstandings about isochores. Part 1. Gene 276, 3–13 (2001)
The SNP Consortium An SNP map of the human genome generated by reduced representation shotgun sequencing. Nature 407, 513–516 (2000)
Perry, J. & Ashworth, A. Evolutionary rate of a gene affected by chromosomal position. Curr. Biol. 9, 987–989 (1999)
Begun, D. J. & Aquadro, C. F. Levels of naturally occurring DNA polymorphism correlate with recombination rates in D. melanogaster. Nature 356, 519–520 (1992)
Nachman, M. W. Single nucleotide polymorphisms and recombination rate in humans. Trends Genet. 17, 481–485 (2001)
Kong, A. et al. A high-resolution recombination map of the human genome. Nature Genet. 31, 241–247 (2002)
Charlesworth, B. The effect of background selection against deleterious mutations on weakly selected, linked variants. Genet. Res. 63, 213–227 (1994)
Hudson, R. R. & Kaplan, N. L. Deleterious background selection with recombination. Genetics 141, 1605–1617 (1995)
Maynard Smith, J. & Haigh, J. The hitch-hiking effect of a favourable gene. Genet. Res. 23, 23–35 (1974)
Birky, C. W. & Walsh, J. B. Effects of linkage on rates of molecular evolution. Proc. Natl Acad. Sci. USA 85, 6414–6418 (1988)
Francino, M. P. & Ochman, H. Strand asymmetries in DNA evolution. Trends Genet. 13, 240–245 (1997)
Gilbert, N., Lutz-Prigge, S. & Moran, J. Genomic deletions created upon LINE-1 retrotransposition. Cell 110, 315–325 (2002)
Symer, D. et al. Human l1 retrotransposition is associated with genetic instability in vivo. Cell 110, 327–338 (2002)
Moran, J. et al. High frequency retrotransposition in cultured mammalian cells. Cell 87, 917–927 (1996)
Hughes, J. F. & Coffin, J. M. Evidence for genomic rearrangements mediated by human endogenous retroviruses during primate evolution. Nature Genet. 29, 487–489 (2001)
Wolfe, K. H. Mammalian DNA replication: mutation biases and the mutation rate. J. Theor. Biol. 149, 441–451 (1991)
Gu, X. & Li, W. H. A model for the correlation of mutation rate with GC content and the origin of GC-rich isochores. J. Mol. Evol. 38, 468–475 (1994)
Gabriel, S. B. et al. The structure of haplotype blocks in the human genome. Science 296, 2225–2229 (2002)
Lindblad-Toh, K. et al. Large-scale discovery and genotyping of single-nucleotide polymorphisms in the mouse. Nature Genet. 24, 381–386 (2000)
Wade, C. M. et al. The mosaic structure of variation in the laboratory mouse genome. Nature 420, 574–578 (2002)
Loftus, S. K. et al. Mutation of melanosome protein RAB38 in chocolate mice. Proc. Natl Acad. Sci. USA 99, 4471–4476 (2002)
Paigen, K. & Eppig, J. T. A mouse phenome project. Mamm. Genome 11, 715–717 (2000)
Doerge, R. W. Mapping and analysis of quantitative trait loci in experimental populations. Nature Rev. Genet. 3, 43–52 (2002)
Cormier, R. T. et al. The Mom1AKR intestinal tumour resistance region consists of Pla2g2a and a locus distal to D4Mit64. Oncogene 19, 3182–3192 (2000)
Mei, R. et al. Genome-wide detection of allelic imbalance using human SNPs and high- density DNA arrays. Genome Res. 10, 1126–1137 (2000)
Lindblad-Toh, K. et al. Loss-of-heterozygosity analysis of small-cell lung carcinomas using single-nucleotide polymorphism arrays. Nature Biotechnol. 18, 1001–1005 (2000)
Heiskanen, M. et al. CGH, cDNA and tissue microarray analyses implicate FGFR2 amplification in a small subset of breast tumors. Anal. Cell Pathol. 22, 229–234 (2001)
Cai, W. W. et al. Genome-wide detection of chromosomal imbalances in tumors using BAC microarrays. Nature Biotechnol. 20, 393–396 (2002)
Davies, H. et al. Mutations of the BRAF gene in human cancer. Nature 417, 949–954 (2002)
Mikkers, H. et al. High-throughput retroviral tagging to identify components of specific signaling pathways in cancer. Nature Genet. 32, 153–159 (2002)
Hwang, H. C. et al. Identification of oncogenes collaborating with p27Kip1 loss by insertional mutagenesis and high-throughput insertion site analysis. Proc. Natl Acad. Sci. USA 99, 11293–11298 (2002)
Lund, A. et al. Genome-wide retroviral insertional tagging of genes involved in cancer in Cdkn2a-deficient mice. Nature Genet. 32, 160–165 (2002)
Janne, P. A. et al. Functional overlap between murine Inpp5b and Ocrl1 may explain why deficiency of the murine ortholog for OCRL1 does not cause Lowe syndrome in mice. J. Clin. Invest. 101, 2042–2053 (1998)
Saitou, N. & Nei, M. The neighbour-joining method: a new method for reconstructing phylogenetic trees. Mol. Biol. Evol. 4, 406–425 (1987)
Sokal, R. & Rohlf, F. Biometry: The Principles and Practice of Statistics in Biological Research (Freeman, New York, 1995)
Sutton, K. A. & Wilkinson, M. F. Rapid evolution of a homeodomain: evidence for positive selection. J. Mol. Evol. 45, 579–588 (1997)
Kasper, S. & Matusik, R. J. Rat probasin: structure and function of an outlier lipocalin. Biochim. Biophys. Acta 1482, 249–258 (2000)
Briand, L. et al. Odorant and pheromone binding by aphrodisin, a hamster aphrodisiac protein. FEBS Lett. 476, 179–185 (2000)
Gow, A. et al. CNS myelin and sertoli cell tight junction strands are absent in Osp/claudin-11 null mice. Cell 99, 649–659 (1999)
Kollmar, R., Nakamura, S. K., Kappler, J. A. & Hudspeth, A. J. Expression and phylogeny of claudins in vertebrate primordia. Proc. Natl Acad. Sci. USA 98, 10196–10201 (2001)
Ashcroft, G. S. et al. Secretory leukocyte protease inhibitor mediates non-redundant functions necessary for normal wound healing. Nature Med. 6, 1147–1153 (2000)
Henderson, C. J., Bammler, T. & Wolf, C. R. Deduced amino acid sequence of a murine cytochrome P-450 Cyp4a protein: developmental and hormonal regulation in liver and kidney. Biochim. Biophys. Acta. 1200, 182–190 (1994)
Simpson, A. E. The cytochrome P450 4 (CYP4) family. Gen. Pharmacol. 28, 351–359 (1997)
Sundseth, S. S. & Waxman, D. J. Sex-dependent expression and clofibrate inducibility of cytochrome P450 4A fatty acid omega-hydroxylases. Male specificity of liver and kidney CYP4A2 mRNA and tissue-specific regulation by growth hormone and testosterone. J. Biol. Chem. 267, 3915–3921 (1992)
Myal, Y. et al. Tissue-specific androgen-inhibited gene expression of a submaxillary gland protein, a rodent homolog of the human prolactin-inducible protein/GCDFP-15 gene. Endocrinology 135, 1605–1610 (1994)
Huang, Y. H., Chu, S. T. & Chen, Y. H. Seminal vesicle autoantigen, a novel phospholipid-binding protein secreted from luminal epithelium of mouse seminal vesicle, exhibits the ability to suppress mouse sperm motility. Biochem. J. 343, 241–248 (1999)
Ann, D. K., Smith, M. K. & Carlson, D. M. Molecular evolution of the mouse proline-rich protein multigene family. Insertion of a long interspersed repeated DNA element. J. Biol. Chem. 263, 10887–10893 (1988)
Rosinski-Chupin, I. & Rougeon, F. A new member of the glutamine-rich protein gene family is characterized by the absence of internal repeats and the androgen control of its expression in the submandibular gland of rats. J. Biol. Chem. 265, 10709–10713 (1990)
Rajkovic, A., Yan, C., Yan, W., Klysik, M. & Matzuk, M. M. Obox, a family of homeobox genes preferentially expressed in germ cells. Genomics 79, 711–717 (2002)
Talley, H. M., Laukaitis, C. M. & Karn, R. C. Female preference for male saliva: implications for sexual isolation of Mus musculus subspecies. Evol. Int. J. Org. Evol. 55, 631–634 (2001)
Dlouhy, S. R., Taylor, B. A. & Karn, R. C. The genes for mouse salivary androgen-binding protein (ABP) subunits alpha and gamma are located on chromosome 7. Genetics 115, 535–543 (1987)
Jia, H. P. et al. A novel murine beta-defensin expressed in tongue, esophagus, and trachea. J. Biol. Chem. 275, 33314–33320 (2000)
Peters, J. Nonspecific esterases of Mus musculus. Biochem. Genet. 20, 585–606 (1982)
Abou-Haila, A., Orgebin-Crist, M. C., Skudlarek, M. D. & Tulsiani, D. R. Identification and androgen regulation of egasyn in the mouse epididymis. Biochim. Biophys. Acta. 1401, 177–186 (1998)
Lin, J., Toft, D. J., Bengtson, N. W. & Linzer, D. I. Placental prolactins and the physiology of pregnancy. Recent Prog. Horm. Res. 55, 37–51 (2000)
Goffin, V., Binart, N., Touraine, P. & Kelly, P. A. Prolactin: the new biology of an old hormone. Annu. Rev. Physiol. 64, 47–67 (2002)
Batten, D., Dyer, K. D., Domachowske, J. B. & Rosenberg, H. F. Molecular cloning of four novel murine ribonuclease genes: unusual expansion within the ribonuclease A gene family. Nucleic Acids Res. 25, 4235–4239 (1997)
Cormier, S. A. et al. Mouse eosinophil-associated ribonucleases: a unique subfamily expressed during hematopoiesis. Mamm. Genome 12, 352–361 (2001)
Tsui, F. W. et al. Molecular characterization and mapping of murine genes encoding three members of the stefin family of cysteine proteinase inhibitors. Genomics 15, 507–514 (1993)
Parham, P. Virtual reality in the MHC. Immunol. Rev. 167, 5–15 (1999)
Ning, Z., Cox, A. J. & Mullikin, J. C. SSAHA: a fast search method for large DNA databases. Genome Res. 11, 1725–1729 (2001)
Flicek, P. et al. Leveraging the mouse genome for gene prediction in human: From the whole-genome shotgun reads to a global synteny map. Genome Res. (in the press)
Parra, G. et al. Comparative gene prediction in human and mouse. Genome Res. (in the press)
Guigó, R. et al. Comparison of mouse and human genomes followed by experimental verification yields an estimated 1,019 additional genes. Proc. Natl Acad. Sci. USA (in the press)
Schwartz, S. et al. Human-mouse alignments with Blastz. Genome Res. (in the press)
Elnitski, L. et al. Distinguishing regulatory DNA from neutral sites. Genome Res. (in the press)
Roskin, K. M. Score Functions for Assessing Conservation in Locally Aligned Regions of DNA from Two Species. UCSC Tech Report UCSC-CRL-02-30, School of Engineering, Univ. California (2002)
Acknowledgements
We thank J. Takahashi and M. Johnston for comments on the manuscript; the Mouse Liaison Group for strategic advice; L. Gaffney, D. Leja and K.-S. Toh for graphical help; B. Graham and G. Roberts for administrative work on sequencing of individual mouse BACs; and P. Kassos and M. McMurtry for secretarial assistance. We thank D. Hill and L. Corbani of the Mouse Genome Informatics Group for their contributions to the GO analysis for mouse and human, and the members of the Bork group at EMBL for discussions. Funding was provided by the National Institutes of Health (National Human Genome Research Institute, National Cancer Institute, National Institute of Dental and Craniofacial Research, National Institute of Diabetes and Digestive and Kidney Diseases, National Institute of General Medical Sciences, National Eye Institute, National Institute of Environmental Health Sciences, National Institute of Aging, National Institute of Arthritis and Musculoskeletal and Skin Diseases, National Institute on Deafness and Other Communication Disorders, National Institute of Mental Health, National Institute on Drug Abuse, National Center for Research Resources, the National Heart Lung and Blood Institute and The Fogarty International Center); the Wellcome Trust; the Howard Hughes Medical Institute; the United States Department of Energy; the National Science Foundation; the Medical Research Council; NSERC; BMBF (German Ministry for Research and Education); the European Molecular Biology Laboratory; Plan Nacional de I + D and Instituto Carlos III; Swiss National Science Foundation, NCCR Frontiers in Genetics, the Swiss Cancer League and the ‘Childcare’ and ‘J. Lejeune’ Foundations; and the Ministry of Education, Culture, Sports, Science and Technology of Japan. The initial threefold sequence coverage was partly supported by the Mouse Sequencing Consortium (GlaxoSmithKline, Merck and Affymetrix) through the Foundation for the National Institutes of Health. We acknowledge A. Holden for coordinating the Mouse Sequencing Consortium. We thank the Sanger Institute systems group for maintenance and provision of the computer resource. The MGSC also used Hewlett-Packard Company's BioCluster, a configuration of 27 HP AlphaServer ES40 systems with 100 CPUs and 1 terabyte of storage. The BioCluster is housed in Hewlett-Packard's IQ Solutions Center, and was accessed remotely. The computing resource greatly accelerated the analysis.
Author information
Authors and Affiliations
Consortia
Corresponding authors
Ethics declarations
Competing interests
The authors declare that they have no competing financial interests.
Supplementary information
Rights and permissions
About this article
Cite this article
Mouse Genome Sequencing Consortium. Initial sequencing and comparative analysis of the mouse genome. Nature 420, 520–562 (2002). https://doi.org/10.1038/nature01262
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1038/nature01262
This article is cited by
-
Differential gene expression (DGE) analysis in persons with a history of giardiasis
AMB Express (2024)
-
Decryption of sequence, structure, and functional features of SINE repeat elements in SINEUP non-coding RNA-mediated post-transcriptional gene regulation
Nature Communications (2024)
-
How chemokines organize the tumour microenvironment
Nature Reviews Cancer (2024)
-
Whole‑exome sequencing reveals Lewis lung carcinoma is a hypermutated Kras/Nras–mutant cancer with extensive regional mutation clusters in its genome
Scientific Reports (2024)
-
Long-run real-time PCR analysis of repetitive nuclear elements as a novel tool for DNA damage quantification in single cells: an approach validated on mouse oocytes and fibroblasts
Journal of Applied Genetics (2024)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.