Abstract
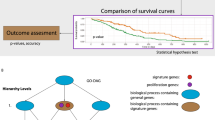
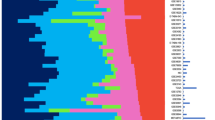
Breast cancer patients with the same stage of disease can have markedly different treatment responses and overall outcome. The strongest predictors for metastases (for example, lymph node status and histological grade) fail to classify accurately breast tumours according to their clinical behaviour1,2,3. Chemotherapy or hormonal therapy reduces the risk of distant metastases by approximately one-third; however, 70–80% of patients receiving this treatment would have survived without it4,5. None of the signatures of breast cancer gene expression reported to date6,7,8,9,10,11,12 allow for patient-tailored therapy strategies. Here we used DNA microarray analysis on primary breast tumours of 117 young patients, and applied supervised classification to identify a gene expression signature strongly predictive of a short interval to distant metastases (‘poor prognosis’ signature) in patients without tumour cells in local lymph nodes at diagnosis (lymph node negative). In addition, we established a signature that identifies tumours of BRCA1 carriers. The poor prognosis signature consists of genes regulating cell cycle, invasion, metastasis and angiogenesis. This gene expression profile will outperform all currently used clinical parameters in predicting disease outcome. Our findings provide a strategy to select patients who would benefit from adjuvant therapy.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
References
McGuire, W. L. Breast cancer prognostic factors: evaluation guidelines. J. Natl Cancer Inst. 83, 154–155 (1991).
Goldhirsch, A., Glick, J. H., Gelber, R. D. & Senn, H. J. Meeting highlights: international consensus panel on the treatment of primary breast cancer. J. Natl Cancer Inst. 90, 1601–1608 (1998).
Eifel, P. et al. National institutes of health consensus development conference statement: adjuvant therapy for breast cancer, November 1–3, 2000. J. Natl Cancer Inst. 93, 979–989 (2001).
Early Breast Cancer Trialists' Collaborative Group. Polychemotherapy for early breast cancer: an overview of the randomised trials. Lancet 352, 930–942 (1998).
Early Breast Cancer Trialists' Collaborative Group. Tamoxifen for early breast cancer: an overview of the randomised trials. Lancet 351, 1451–1467 (1998).
Perou, C. M. et al. Distinctive gene expression patterns in human mammary epithelial cells and breast cancers. Proc. Natl Acad. Sci. USA 96, 9212–9217 (1999).
Perou, C. M. et al. Molecular portraits of human breast tumours. Nature 406, 747–752 (2000).
Gruvberger, S. et al. Estrogen receptor status in breast cancer is associated with remarkably distinct gene expression patterns. Cancer Res. 61, 5979–5984 (2001).
Martin, K. J. et al. Linking gene expression patterns to therapeutic groups in breast cancer. Cancer Res. 60, 2232–2238 (2000).
Zajchowski, D. A. et al. Identification of gene expression profiles that predict the aggressive behavior of breast cancer cells. Cancer Res. 61, 5168–5178 (2001).
Sorlie, T. et al. Gene expression patterns of breast carcinomas distinguish tumor subclasses with clinical implications. Proc. Natl Acad. Sci. USA 98, 10869–10874 (2001).
West, M. et al. Predicting the clinical status of human breast cancer by using gene expression profiles. Proc. Natl Acad. Sci. USA 98, 11462–11467 (2001).
Hughes, T. R. et al. Expression profiling using microarrays fabricated by an ink-jet oligonucleotide synthesizer. Nature Biotechnol. 19, 342–347 (2001).
Roberts, C. J. et al. Signaling and circuitry of multiple MAPK pathways revealed by a matrix of global gene expression profiles. Science 287, 873–880 (2000).
Lakhani, S. R. et al. Multifactorial analysis of differences between sporadic breast cancers and cancers involving BRCA1 and BRCA2 mutations. J. Natl Cancer Inst. 90, 1138–1145 (1998).
Brenton, J. D., Aparicio, S. A. & Caldas, C. Molecular profiling of breast cancer: portraits but not physiognomy. Breast Cancer Res. 3, 77–80 (2001).
Khan, J. et al. Classification and diagnostic prediction of cancers using gene expression profiling and artificial neural networks. Naure Med. 7, 673–679 (2001).
He, Y. D. & Friend, S. H. Microarrays—the 21st century divining rod? Nature Med. 7, 658–659 (2001).
Bieche, I. et al. Genetic alterations in breast cancer. Genes Chromosomes Cancer 14, 227–251 (1995).
Steeg, P. S. & Zhou, Q. Cyclins and breast cancer. Breast Cancer Res. Treat. 52, 17–28 (1998).
Janicke, F. et al. Randomized adjuvant chemotherapy trial in high-risk, lymph node-negative breast cancer patients identified by urokinase-type plasminogen activator and plasminogen activator inhibitor type 1. J. Natl Cancer Inst. 93, 913–920 (2001).
van Diest, P. J. et al. Cyclin D1 expression in invasive breast cancer. Correlations and prognostic value. Am. J. Pathol. 150, 705–711 (1997).
Zheng, L., Annab, L. A., Afshari, C. A., Lee, W. H. & Boyer, T. G. BRCA1 mediates ligand-independent transcriptional repression of the estrogen receptor. Proc. Natl Acad. Sci. USA 98, 9587–9592 (2001).
Esteller, M. et al. Promoter hypermethylation and BRCA1 inactivation in sporadic breast and ovarian tumors. J. Natl Cancer Inst. 92, 564–569 (2000).
Chapman, M. S. & Verma, I. M. Transcriptional activation by BRCA1. Nature 382, 678–679 (1996).
Hedenfalk, I. et al. Gene-expression profiles in hereditary breast cancer. N. Engl. J. Med. 344, 539–548 (2001).
Acknowledgements
We thank D. Atsma and D. Majoor for assistance with the histological analyses and the preparation of tumour RNA; T. van der Velde, W. van Waardenburg and O. Dalesio for medical record data extraction; D. Slade, J. McDonald, J. Koch, T. Erkkila, M. Parrish and others at Rosetta's High Throughput Gene Expression Profiling Facility for microarray experiments; R. Stoughton, F. van Leeuwen, M. Rookus, P. Nederlof, F. Hogervorst and D. Voskuil for suggestions; and A. Berns, L. Hartwell, J. Radich and S. Rodenhuis for support and reading of the manuscript. This work was supported by a grant from the Center for Biomedical Genetics.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
S.H.F. is the Vice President of the MRC Merck Research Laboratories.
Supplementary information
Rights and permissions
About this article
Cite this article
van 't Veer, L., Dai, H., van de Vijver, M. et al. Gene expression profiling predicts clinical outcome of breast cancer. Nature 415, 530–536 (2002). https://doi.org/10.1038/415530a
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1038/415530a
This article is cited by
-
Clinically relevant gene signatures provide independent prognostic information in older breast cancer patients
Breast Cancer Research (2024)
-
Predictive, preventive, and personalized medicine in breast cancer: targeting the PI3K pathway
Journal of Translational Medicine (2024)
-
Novel model integrating computed tomography-based image markers with genetic markers for discriminating radiation pneumonitis in patients with unresectable stage III non-small cell lung cancer receiving radiotherapy: a retrospective multi-center radiogenomics study
BMC Cancer (2024)
-
Identification of lineage-specific epigenetic regulators FOXA1 and GRHL2 through chromatin accessibility profiling in breast cancer cell lines
Cancer Gene Therapy (2024)
-
Development of an invasion score based on metastasis-related pathway activity profiles for identifying invasive molecular subtypes of lung adenocarcinoma
Scientific Reports (2024)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.