Abstract
Arbuscular mycorrhizas are the most common non-pathogenic symbioses in the roots of plants. It is generally assumed that this symbiosis facilitated the colonization of land by plants1. In arbuscular mycorrhizas, fungal hyphae often extend between the root cells and tuft-like branched structures (arbuscules) form within the cell lumina that act as the functional interface for nutrient exchange. In the mutualistic arbuscular-mycorrhizal symbiosis the host plant derives mainly phosphorus from the fungus, which in turn benefits from plant-based glucose2. The molecular basis of the establishment and functioning of the arbuscular-mycorrhizal symbiosis is largely not understood. Here we identify the phosphate transporter gene StPT3 in potato (Solanum tuberosum). Functionality of the encoded protein was confirmed by yeast complementation. RNA localization and reporter gene expression indicated expression of StPT3 in root sectors where mycorrhizal structures are formed. A sequence motif in the StPT3 promoter is similar to transposon-like elements, suggesting that the mutualistic symbiosis evolved by genetic rearrangements in the StPT3 promoter.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Remy, W., Taylor, T. N., Hass, H. & Kerp, H. Four-hundred-million-year-old vesicular arbuscular mycorrhizae. Proc. Natl Acad. Sci. USA 91, 11841–11843 (1994).
Smith, S. E. & Read, D. J. Mycorrhizal Symbiosis (Academic, San Diego, California, 1997).
Gianinazzi-Pearson, V., Arnould, C., Oufattole, M., Arango, M. & Gianinazzi, S. Differential activation of H+-ATPase genes by an arbuscular mycorrhizal fungus in root cells of transgenic tobacco. Planta 211, 609–613 (2000).
Chiou, T. J., Liu, H. & Harrison, M. J. The spatial expression patterns of a phosphate transporter (MtPT1) from Medicago truncatula indicate a role in phosphate transport at the root/soil interface. Plant J. 25, 281–293 (2001).
Daram, P., Brunner, S., Persson, B. L., Amrhein, N. & Bucher, M. Functional analysis and cell-specific expression of a phosphate transporter from tomato. Planta 206, 225–233 (1998).
Harrison, M. J. & Van Buuren, M. L. A phosphate transporter from the mycorrhizal fungus Glomus versiforme. Nature 378, 626–629 (1995).
Bucher, M., Rausch, C. & Daram, P. Molecular and biochemical mechanisms of phosphorus uptake into plants. J. Plant Nutr. Soil Sci. 164, 209–217 (2001).
Rosewarne, G., Barker, S., Smith, S., Smith, F. & Schachtman, D. A Lycopersicon esculentum phosphate transporter (LePT1) involved in phosphorus uptake from a vesicular-arbuscular mycorrhizal fungus. New Phytol. 144, 507–516 (1999).
Liu, H., Trieu, A. T., Blaylock, L. A. & Harrison, M. J. Cloning and characterization of two phosphate transporters from Medicago truncatula roots: Regulation in response to phosphate and to colonization by arbuscular mycorrhizal (AM) fungi. Mol. Plant–Microbe Interact. 11, 14–22 (1998).
Leggewie, G., Willimitzer, L. & Riesmeier, J. W. Two cDNAs from potato are able to complement a phosphate uptake-deficient yeast mutant: Identification of phosphate transporters from higher plants. Plant Cell 9, 381–392 (1997).
Martinez, P. & Persson, B. L. Identification, cloning and characterization of a derepressible Na+-coupled phosphate transporter in Saccharomyces cerevisiae. Mol. Gen. Genet. 258, 628–638 (1998).
Jefferson, R. A., Kavanagh, T. A. & Bevan, M. W. GUS fusions: β-glucuronidase as a sensitive and versatile gene fusion marker in higher plants. EMBO J. 3901–3907 (1987).
Wessler, S. R., Bureau, T. E. & White, S. E. LTR-retrotransposons and MITES—Important players in the evolution of plant genomes. Curr. Opin. Genet. Dev. 5, 814–821 (1995).
Hibbett, D. S., Gilbert, L. B. & Donoghue, M. J. Evolutionary instability of ectomycorrhizal symbioses in basidiomycetes. Nature 407, 506–508 (2000).
Selinger, D. A. & Chandler, V. L. Major recent and independent changes in levels and patterns of expression have occurred at the b gene, a regulatory locus in maize. Proc. Natl Acad. Sci. USA 96, 15007–15012 (1999).
Varma, A. in Endomycorrhiza (eds Varma, A. & Hock, B.) 521–556 (Springer, Berlin, 1999).
Johnson, N. C., Graham, J. H. & Smith, F. A. Functioning of mycorrhizal associations along the mutualism–parasitism continuum. New Phytol. 135, 575–585 (1997).
Grime, J. P., Mackey, J. M. L., Hillier, S. H. & Read, D. J. Floristic diversity in a model system using experimental microcosms. Nature 328, 420–422 (1987).
van der Heijden, M. et al. Mycorrhizal fungal diversity determines plant biodiversity, ecosystem variability and productivity. Nature 396, 69–72 (1998).
Lockett, T. A bacteriophage lambda DNA purification procedure suitable for the analysis of DNA from either large or multiple small lysates. Anal. Biochem. 185, 230–234 (1990).
Depicker, A., Stachel, S., Dhaese, P., Zambryski, P. & Goodman, H. M. Nopaline synthase: transcript mapping and DNA sequence. J. Mol. Appl. Genet. 1, 561–573 (1982).
Bariola, P. et al. The Arabidopsis ribonuclease gene RNS1 is tightly controlled in response to phosphate limitation. Plant J. 6, 673–685 (1994).
Bevan, M. Binary Agrobacterium vectors for plant transformation. Nucleic Acids Res. 12, 8711–8721 (1984).
Dohmen, R. J., Strasser, A. W. M., Höner, C. B. & Hollenberg, C. P. An efficient transformation procedure enabling long-term storage of competent yeast cells of various yeast genera. Yeast 7, 691–692 (1991).
Daram, P. et al. Pht2;1 encodes a low-affinity phosphate transporter from Arabidopsis. Plant Cell 11, 2153–2166 (1999).
Deblaere, R. et al. Efficient octopine Ti plasmid-derived vectors for Agrobacterium-mediated gene transfer to plants. Nucleic Acids Res. 13, 4777–4788 (1985).
Rocha-Sosa, M. et al. Both developmental and metabolic signals activate the promoter of a class I patatin gene. EMBO J. 8, 23–29 (1989).
Hoagland, D. & Broyer, T. General nature of the process of salt accumulation by roots with description of experimental methods. Plant Physiol. 11, 471–507 (1936).
Brundrett, M. C., Piché, Y. & Peterson, R. L. A new method for observing the morphology of vesicular-arbuscular mycorrhizae. Can. J. Bot. 62, 2128–2134 (1984).
Merryweather, J. W. & Fitter, A. H. A modified procedure for elucidating the structure of the fungal partner in a vesicular-arbuscular mycorrhiza. Myc. Res. 95, 1435–1437 (1991).
Acknowledgements
We thank R. Ruh for support with microscopic analysis; B. Regierer and J. Kossmann for the potato genomic library; and T. Mandel and C. Kuhlemeier for the 35S minimal promoter. This work was supported by the Swiss National Science Foundation, and Hoffmann-La Roche, Switzerland.
Author information
Authors and Affiliations
Corresponding author
Supplementary information
StPT3 gene structure and predicted topology of the encoded protein.

Figure A
(GIF 17.2 KB)
a, The structure of the StPT3 gene is presented. The diagram depicts the intron-less open reading frame (open box) which is terminated by a TAA in-frame stop codon and 5’ upstream and 3’ non-translated regions (thin lines). TATA box, putative binding site for HD-ZIP transcription factors, and MRR element are indicated in the 5’ upstream region. Sizes are given in base pairs with 1 indicating the first nucleotide of the intitiation codon ATG. Distances are drawn to scale. b, Predicted topology of the StPT3 polypeptide with encircled numbers specifying the 12 transmembrane domains embedded in the membrane bilayer. Putative sites for posttranslational modifications are indicated with P for phosphorylation and a schematic oligosaccharide moiety for N-linked glycosylation.
Alignment of the deduced amino acid sequence of StPT3 with that of plant Pht1 proteins.

Figure B
(GIF 29.5 KB)
Alignment of the deduced amino acid sequence of StPT3 with the Pht1 transporters from potato and tomato (see Fig. 1). Identical amino acids are shaded in black, similar amino acids are shaded in grey. The membrane spanning domains are underlined in the StPT3 sequence (-). (*) Consensus site for N-linked glycosylation, and (+) sites for phosphorylation conserved in all proteins aligned in Figure 1.
A MITE-like element in the StPT3 promoter region exhibits homology to resistance (R) genes.

Figure C
(GIF 17.5 KB)
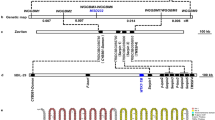
a, Sequence of a 431 base pairs MITE-like transposable element in the StPT3 upstream genic region. Shaded regions designate TIRs. The MRR element is underlined. b, alignment of StPT3 MRR element with sequences from solanaceous resistance (R) genes as obtained by a BLAST search. The highly conserved boxes MRR1 and MRR2 are designated by thin lines. Gene or cluster names are followed by GenBank accession numbers: rclust-a and rclust-b from the S. tuberosum resistance gene cluster (AF265664); VFNT-Pto from the L. esculentum VFNT Cherry Pto locus (AF220603); rx from the S. tuberosum rx gene (AJ011801); Pto-a and Pto-b from the L. pimpinellifolium Rio Grande 76R Pto locus (AF220602); Mi-copy2 from the L. esculentum disease resistance gene homologue Mi-copy2 (U81378). c, alignment of MRR1 and MRR2 elements from the genomic regions specified at left. Identities between different elements are listed at the bottom of the different alignments. HCR9-4A and HCR9-9A specify genes from the L. pimpinellifolium Cf-9 resistance gene cluster (AJ002236) or the L. hirsutum Cf-4 resistance gene cluster (AJ002235), respectively.
Rights and permissions
About this article
Cite this article
Rausch, C., Daram, P., Brunner, S. et al. A phosphate transporter expressed in arbuscule-containing cells in potato. Nature 414, 462–465 (2001). https://doi.org/10.1038/35106601
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1038/35106601
This article is cited by
-
Identification of GmPT proteins and investigation of their expressions in response to abiotic stress in soybean
Planta (2024)
-
Effects of field inoculation of potato tubers with the arbuscular mycorrhizal fungus Rhizophagus irregularis DAOM 197198 are cultivar dependent
Symbiosis (2023)
-
Mycorrhizal symbiosis-induced abiotic stress mitigation through phosphate transporters in Solanum lycopersicum L
Plant Growth Regulation (2023)
-
Diverse mycorrhizal maize inbred lines differentially modulate mycelial traits and the expression of plant and fungal phosphate transporters
Scientific Reports (2022)
-
Microbial enhancement of plant nutrient acquisition
Stress Biology (2022)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.