Abstract
Genome and expressed sequence tag projects are rapidly cataloguing and cloning the genes of higher organisms, including humans. An emerging challenge is to rapidly uncover the functions of genes and to identify gene products with desired properties. We have developed a microarray-driven gene expression system for the functional analysis of many gene products in parallel. Mammalian cells are cultured on a glass slide printed in defined locations with different DNAs. Cells growing on the printed areas take up the DNA, creating spots of localized transfection within a lawn of non-transfected cells. By printing sets of complementary DNAs cloned in expression vectors, we make microarrays whose features are clusters of live cells that express a defined cDNA at each location. Here we demonstrate two uses for our approach: as an alternative to protein microarrays for the identification of drug targets, and as an expression cloning system for the discovery of gene products that alter cellular physiology. By screening transfected cell microarrays expressing 192 different cDNAs, we identified proteins involved in tyrosine kinase signalling, apoptosis and cell adhesion, and with distinct subcellular distributions.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
References
Schena, M., Shalon, D., Davis, R. W. & Brown, P. O. Quantitative monitoring of gene expression patterns with a complementary DNA microarray. Science 270, 467–470 (1995).
Lockhard, D. J. et al. Expression monitoring by hybridization to high-density oligonucleotide arrays. Nature Biotechnol. 14, 1675–1678 (1996).
Uetz, P. et al. A comprehensive analysis of protein–protein interactions in Saccharomyces cerevisiae. Nature 403, 623–627 (2000).
Kino, T. et al. FK-506, a novel immunosuppressant isolated from a Streptomyces. II. Immunosuppressive effect of FK-506 in vitro. J. Antibiot. (Tokyo) 40, 1256–1265 (1987).
Harding, M. W., Galat, A., Uehling, D. E. & Schreiber, S. L. A receptor for the immunosuppressant FK506 is a cis-trans peptidyl-prolyl isomerase. Nature 341, 758–760 (1989).
Siekierka, J. J., Hung, S. H., Poe, M., Lin, C. S. & Sigal, N. H. A cytolsolic binding protein for the immunosuppressant FK506 has peptidyl-prolyl isomerase activity but is distinct from cyclophilin. Nature 341, 755–757 (1989).
Schreiber, S. L. Chemistry and biology of the immunophilins and their immunosuppressive ligands. Science 251, 283–287 (1991).
Jacinto, E., Werlen, G. & Karin, M. Cooperation between Syk and Rac1 leads to synergistic JNK activation in T lymphocytes. Immunity 8, 31–41 (1998).
Gavrieli, Y., Sherman, Y. & Ben-Sasson, S. A. Identification of programmed cell death in situ via specific labeling of nuclear DNA fragmentation. J. Cell Biol. 119, 493–501 (1992).
Walczak, H. et al. TRAIL-R2: a novel apoptosis-mediating receptor for TRAIL. EMBO J. 16, 5386–5397 (1997).
Tandon, N. N., Kralisz, U. & Jamieson, G. A. Identification of glycoprotein IV (CD36) as a primary receptor for platelet-collagen adhesion. J. Biol. Chem. 264, 7576–7583 (1989).
Simonsen, H. & Lodish, H. F. Cloning by function: expression cloning in mammalian cells. Trends Pharmacol. Sci. 15, 437–441 (1994).
Strausberg, R. L., Feingold, E. A., Klausner, R. D. & Collins, F. S. The mammalian gene collection. Science 286, 455–457 (1999).
Kawai, J. et al. Functional annotation of a full-length mouse cDNA collection. Nature 409, 685–690 (2001).
Yudate, H. T. et al. HUNT: launch of a full-length cDNA database from the Helix Research Institute. Nucleic Acids Res. 29,185–188 (2001).
Wiemann, S. et al. Toward a catalog of human genes and proteins: sequencing and analysis of 500 novel complete protein coding human cDNAs. Genome Res. 11,422–435 (2001).
MacBeath, G. & Schreiber, S. L. Printing proteins as microarrays for high-throughput function determination. Science 289, 1760–1763 (2000).
Lueking, A. et al. Protein microarrays for gene expression and antibody screening. Anal. Biochem. 270, 103–111 (1999).
de Wildt, R. M., Mundy, C. R., Gorick, B. D. & Tomlinson, I. M. Antibody arrays for high-throughput screening of antibody-antigen interactions. Nature Biotechnol. 9, 989–994 (2000).
Morgenstern, J. P. & Land, H. Advanced mammalian gene transfer: high titre retroviral vectors with multiple drug selection markers and a complementary helper-free packaging cell line. Nucleic Acids Res. 18, 3587–3596 (1990).
Sabatini, D. M. et al. Interaction of RAFT1 with gephyrin required for rapamycin-sensitive signaling. Science 284, 1161–1164 (1999).
Acknowledgements
We thank C. Wilson and T. Volkert for advice on use of the microarrayer, and K. McKernan and P. McEwan for the neurotransmitter receptor constructs. We also thank R. Young, N. Hacohen and members of the Sabatini laboratory for their support and suggestions; P. Kim and M. Gerlach for help with the manuscript; and Invitrogen for the full-length clones from the Genestorm collection. This research was supported by the G. Harold and Leila Y. Mathers Charitable Foundation, the Whitehead Institute Fellows Program and Corning. Microscopy work was conducted at the W. M. Keck Foundation Biological Imaging facility at the Whitehead Institute.
Author information
Authors and Affiliations
Corresponding author
Supplementary information
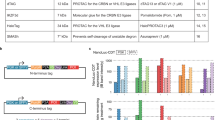
Clones from the Invitrogen Genestorm collection were used to create transfected cell microarrays expressing 192 cDNAs.
Genestorm cDNAs are in pcDNA3.1/GS vector.
Rights and permissions
About this article
Cite this article
Ziauddin, J., Sabatini, D. Microarrays of cells expressing defined cDNAs. Nature 411, 107–110 (2001). https://doi.org/10.1038/35075114
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1038/35075114
This article is cited by
-
Droplet microarray platforms for high-throughput drug screening
Microchimica Acta (2023)
-
Accelerated transgene expression of pDNA/polysaccharide complexes by solid-phase reverse transfection and analysis of the cell transfection mechanism
Polymer Journal (2022)
-
Multiplexed Optical Sensors in Arrayed Islands of Cells for multimodal recordings of cellular physiology
Nature Communications (2020)
-
Novel upregulation of amyloid-β precursor protein (APP) by microRNA-346 via targeting of APP mRNA 5′-untranslated region: Implications in Alzheimer’s disease
Molecular Psychiatry (2019)
-
A mixed antagonistic/synergistic miRNA repression model enables accurate predictions of multi-input miRNA sensor activity
Nature Communications (2018)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.