Abstract
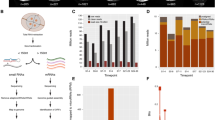
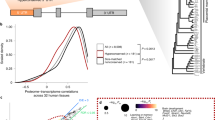
Two small RNAs regulate the timing of Caenorhabditis elegans development1,2. Transition from the first to the second larval stage fates requires the 22-nucleotide lin-4 RNA1,3,4, and transition from late larval to adult cell fates requires the 21-nucleotide let-7 RNA2. The lin-4 and let-7 RNA genes are not homologous to each other, but are each complementary to sequences in the 3′ untranslated regions of a set of protein-coding target genes that are normally negatively regulated by the RNAs1,2,5,6. Here we have detected let-7 RNAs of ∼21 nucleotides in samples from a wide range of animal species, including vertebrate, ascidian, hemichordate, mollusc, annelid and arthropod, but not in RNAs from several cnidarian and poriferan species, Saccharomyces cerevisiae, Escherichia coli or Arabidopsis. We did not detect lin-4 RNA in these species. We found that let-7 temporal regulation is also conserved: let-7 RNA expression is first detected at late larval stages in C. elegans and Drosophila , at 48 hours after fertilization in zebrafish, and in adult stages of annelids and molluscs. The let-7 regulatory RNA may control late temporal transitions during development across animal phylogeny.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Lee, R. C., Feinbaum, R. L. & Ambros, V. The C. elegans heterochronic gene lin-4 encodes small RNAs with antisense complementarity to lin-14. Cell 75, 843–854 ( 1993).
Reinhart, B. et al. The 21-nucleotide let-7 RNA regulates developmental timing in C. elegans. Nature 403, 901–906 (2000).
Chalfie, M., Horvitz, H. R. & Sulston, J. E. Mutations that lead to reiterations in the cell lineages of C. elegans. Cell 24, 59– 69 (1981).
Ambros, V. & Horvitz, H. R. The lin-14 locus of Caenorhabditis elegans controls the time of expression of specific postembryonic developmental events. Genes Dev. 1, 398– 414 (1987).
Wightman, B., Ha, I. & Ruvkun, G. Posttranscriptional regulation of the heterochronic gene lin-14 by lin-4 mediates temporal pattern formation in C. elegans. Cell 75, 855–862 ( 1993).
Slack, F. et al. The lin-41 RBCC gene acts in the C. elegans heterochronic pathway between the let-7 regulatory RNA and the LIN-29 transcription factor. Mol. Cell 5, 659– 669 (2000).
Aguinaldo, A. M. et al. Evidence for a clade of nematodes, arthropods and other moulting animals. Nature 387, 489– 493 (1997).
Zamore, P. D., Tuschl, T., Sharp, P. A. & Bartel, D. P. RNAi: double-stranded RNA directs the ATP-dependent cleavage of mRNA at 21 to 23 nucleotide intervals. Cell 101, 25–33 (2000).
Hammond, S. M., Bernstein, E., Beach, D. & Hannon, G. J. An RNA-directed nuclease mediates post-transcriptional gene silencing in Drosophila cells. Nature 404, 293– 296 (2000).
Tuschl, T., Zamore, P. D., Lehmann, R., Bartel, D. P. & Sharp, P. A. Targeted mRNA degradation by double-stranded RNA in vitro. Genes Dev. 13, 3191 –3197 (1999).
Montgomery, M. K., Xu, S. & Fire, A. RNA as a target of double-stranded RNA-mediated genetic interference in Caenorhabiditis elegans. Proc. Natl Acad. Sci. USA 95, 15502–15507 (1998).
Fire, A. et al. Potent and specific genetic interference by double-stranded RNA in Caenorhbditis elegans. Nature 391, 806–810 (1998).
Olsen, P. H. & Ambros, V. The lin-4 regulatory RNA controls developmental timing in Caenorhabditis elegans by blocking LIN-14 protein synthesis after the initiation of translation. Dev. Biol. 216, 671–680 (1999).
Tabara, H. et al. The rde-1 gene, RNA interference, and transposon silencing in C. elegans. Cell 99, 123– 132 (1999).
Acknowledgements
We thank the following people for RNA and tissue samples: T. Heanue, R. Pearse and C. Tabin for chick; J. Gerhart and M. Kirschner for Xenopus and acorn worm; S. Agarwal for Xenopus; N. Stavropoulos for mouse; C. Unabia and K. del Carmen for annelid and mollusc; H. Bode for Hydra; J. Nardone and S. Ferrari for Arabidopsis; P. Sudarsanam for yeast; and D. Selinger for E. coli. The phylogenetic survey in this work was inspired by the NASA Evolution and Development meetings organized by E. Davidson and C. Golden. This work was supported by an NIH grant from NIGMS to G.R. and a grant from the MGH Fund for Medical Discovery to A.E.P.
Author information
Authors and Affiliations
Rights and permissions
About this article
Cite this article
Pasquinelli, A., Reinhart, B., Slack, F. et al. Conservation of the sequence and temporal expression of let-7 heterochronic regulatory RNA. Nature 408, 86–89 (2000). https://doi.org/10.1038/35040556
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1038/35040556
This article is cited by
-
Unraveling the microRNAs, key players in folliculogenesis and ovarian diseases
Middle East Fertility Society Journal (2024)
-
Complete male-to-female sex reversal in XY mice lacking the miR-17~92 cluster
Nature Communications (2024)
-
MicroRNA cross-talk between Monilinia fungal pathogens and peach host
Phytoparasitica (2024)
-
Curcumin targets miR-134-5p to suppress the progression of colorectal cancer through regulating the CDCA3/CDK1 pathway
Naunyn-Schmiedeberg's Archives of Pharmacology (2024)
-
Regulation of gene expression by modulating microRNAs through Epigallocatechin-3-gallate in cancer
Molecular Biology Reports (2024)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.