Abstract
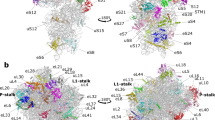
Genetic information encoded in messenger RNA is translated into protein by the ribosome, which is a large nucleoprotein complex comprising two subunits, denoted 30S and 50S in bacteria. Here we report the crystal structure of the 30S subunit from Thermus thermophilus, refined to 3 Å resolution. The final atomic model rationalizes over four decades of biochemical data on the ribosome, and provides a wealth of information about RNA and protein structure, protein–RNA interactions and ribosome assembly. It is also a structural basis for analysis of the functions of the 30S subunit, such as decoding, and for understanding the action of antibiotics. The structure will facilitate the interpretation in molecular terms of lower resolution structural data on several functional states of the ribosome from electron microscopy and crystallography.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout









Similar content being viewed by others
References
Garrett, R. A. et al. (eds) The Ribosome. Structure, Function, Antibiotics and Cellular Interactions (ASM, Washington DC, 2000).
von Böhlen, K. et al. Characterization and preliminary attempts for derivatization of crystals of large ribosomal subunits from Haloarcula marismortui diffracting to 3 Å resolution. J. Mol. Biol. 222, 11–15 (1991).
Clemons, W. M. et al. Structure of a bacterial 30S ribosomal subunit at 5.5 Å resolution. Nature 400, 833– 840 (1999).
Trakhanov, S. D. Crystallization of 70S ribosomes and 30S ribosomal subunits from Thermus thermophilus. FEBS Lett. 220, 319– 322 (1987).
Glotz, C. et al. Three-dimensional crystals of ribosomes and their subunits from eu- and archaebacteria. Biochem. Int. 15, 953–960 (1987).
Yonath, A. et al. Characterization of crystals of small ribosomal subunits. J. Mol. Biol. 203, 831–834 (1988).
Yusupov, M. M., Tischenko, S. V., Trakhanov, S. D., Ryazantsev, S. N. & Garber, M. B. A new crystalline form of 30S ribosomal subunits from Thermus thermophilus. FEBS Lett. 238, 113–115 (1988).
Yonath, A. Crystallographic studies on the ribosome, a large macromolecular assembly exhibiting severe nonisomorphism, extreme beam sensitivity and no internal symmetry. Acta Crystallogr. A 54, 945– 955 (1998).
Tocilj, A. et al. The small ribosomal subunit from Thermus thermophilus at 4.5 Å resolution: pattern fittings and the identification of a functional site. Proc. Natl Acad. Sci. USA 96, 14252– 14257 (1999).
Carter, A. P. et al. Functional insights from the structure of the 30S ribosomal subunit and its interaction with antibiotics. Nature 407, 340–348 (2000).
Hartmann, R. K. & Erdmann, V. A. Thermus thermophilus 16S rRNA is transcribed from an isolated transcription unit. J. Bacteriol. 171, 2933– 2941 (1989).
Choli, T., Franceschi, F., Yonath, A. & Wittmann-Liebold, B. Isolation and characterization of a new ribosomal protein from the thermophilic eubacteria, Thermus thermophilus, T. aquaticus and T. flavus . Biol. Chem. Hoppe Seyler 374, 377– 383 (1993).
Mueller, F. & Brimacombe, R. A new model for the three-dimensional folding of Escherichia coli 16S ribosomal RNA. I. Fitting the RNA to a 3D electron microscopic map at 20 Å. J. Mol. Biol. 271, 524–544 (1997).
Gabashvili, I. S., Agrawal, R. K., Grassucci, R. & Frank, J. Structure and structural variations of the Escherichia coli 30S ribosomal subunit as revealed by three-dimensional cryo-electron microscopy. J. Mol. Biol. 286, 1285–1291 (1999).
Gabashvili, I. S. et al. Solution structure of the E. coli 70S ribosome at 11.5 Å resolution. Cell 100, 537– 549 (2000).
Stern, S., Weiser, B. & Noller, H. F. Model for the three-dimensional folding of 16S ribosomal RNA. J. Mol. Biol. 204, 447– 481 (1988).
Malhotra, A. & Harvey, S. C. A quantitative model of the Escherichia coli 16S RNA in the 30S ribosomal subunit. J. Mol. Biol. 240, 308–340 ( 1994).
Ramakrishnan, V. Distribution of protein and RNA in the 30S ribosomal subunit. Science 231, 1562–1564 ( 1986).
Wimberly, B., Varani, G. & Tinoco, I. The conformation of loop E of eukaryotic 5S ribosomal RNA. Biochemistry 32, 1078– 1087 (1993).
Lodmell, J. S. & Dahlberg, A. E. A conformational switch in Escherichia coli 16S ribosomal RNA during decoding of messenger RNA. Science 277, 1262– 1267 (1997).
Gutell, R. R. in Ribosomal RNA Structure, Evolution, Processing and Function in Protein Biosynthesis (eds Dahlberg, A. E. &&&&& Zimmermann, R. A.) 111–128 (CRC, Boca Raton, 1996).
Nikulin, A. et al. Crystal structure of the S15-rRNA complex. Nature Struct. Biol. 7, 273–277 (2000).
Agalarov, S. C., Sridhar Prasad, G., Funke, P. M., Stout, C. D. & Williamson, J. R. Structure of the S15,S6,S18-rRNA complex: assembly of the 30S ribosome central domain. Science 288, 107–113 (2000).
Powers, T. & Noller, H. F. Hydroxyl radical footprinting of ribosomal proteins on 16S rRNA. RNA 1, 194–209 (1995).
Mueller, F. & Brimacombe, R. A new model for the three-dimensional folding of Escherichia coli 16S ribosomal RNA. II. The RNA-protein interaction data. J. Mol. Biol. 271, 545 –565 (1997).
Capel, M. S. et al. A complete mapping of the proteins in the small ribosomal subunit of Escherichia coli. Science 238, 1403–1406 (1987).
Nowotny, V. & Nierhaus, K. H. Assembly of the 30S subunit from Escherichia coli ribosomes occurs via two assembly domains which are initiated by S4 and S7. Biochemistry 27, 7051–7055 (1988).
Stern, S., Powers, T., Changchien, L. -M. & Noller, H. F. RNA-protein interactions in 30S ribosomal subunits: folding and function of 16S rRNA. Science 244, 783– 790 (1989).
Held, W. A., Ballou, B., Mizushima, S. & Nomura, M. Assembly mapping of 30S ribosomal proteins from Escherichia coli. Further studies. J. Biol. Chem. 249, 3103– 3111 (1974).
Otwinowski, Z. & Minor, W. in Methods in Enzymology (eds Carter, C. W. & Sweet, R. M.) 307–325 (Academic, New York, 1997).
Terwilliger, T. & Berendzen, J. Automated MAD and MIR structure determination. Acta Crystallogr. D 55, 849–861 (1999).
Abrahams, J. P. Bias reduction in phase refinement by modified interference functions: introducing the gamma correction. Acta Crystallogr. D 53, 371–376 (1997).
de la Fortelle, E. & Bricogne, G. in Methods in Enzymology (eds Carter, C. W. & Sweet, R. M.) 472– 493 (Academic, New York, 1997).
Cowtan, K. & Main, P. Miscellaneous algorithms for density modification. Acta Crystallogr. D 54, 487 –493 (1998).
Jones, T. A. & Kjeldgaard, M. Electron-density map interpretation. Methods Enzymol. 277B, 173– 207 (1997).
Brünger, A. T. et al. Crystallography & NMR system: A new software suite for macromolecular structure determination. Acta Crystallogr. D 54, 905–921 (1998).
Carson, M. Ribbons 2.0. J. Appl. Cryst. 24, 958– 961 (1991).
Davies, C., Gerstner, R. B., Draper, D. E., Ramakrishnan, V. & White, S. W. The crystal structure of ribosomal protein S4 reveals a two-domain molecule with an extensive RNA-binding surface: one domain shows structural homology to the ETS DNA-binding motif. EMBO J. 17, 4545–4558 (1998).
Markus, M. A., Gerstner, R. B., Draper, D. E. & Torchia, D. A. The solution structure of ribosomal protein S4 delta41 reveals two subdomains and a positively charged surface that may interact with RNA. EMBO J. 17, 4559–4571 ( 1998).
Ramakrishnan, V. & White, S. W. Structure of ribosomal protein S5 reveals sites of interaction with 16S RNA. Nature 358, 768–771 ( 1992).
Lindahl, M. et al. Crystal structure of the ribosomal protein S6 from Thermus thermophilus. EMBO J. 13, 1249– 1254 (1994).
Wimberly, B. T., White, S. W. & Ramakrishnan, V. The structure of ribosomal protein S7 at 1.9 Å resolution reveals a beta-hairpin motif that binds double-stranded nucleic acids. Structure 5, 1187– 1198 (1997).
Hosaka, H. et al. Ribosomal protein S7: a new RNA-binding motif with structural similarities to a DNA architectural factor. Structure 5, 1199–1208 (1997).
Davies, C., Ramakrishnan, V. & White, S. W. Structural evidence for specific S8-RNA and S8-protein interactions within the 30S ribosomal subunit: ribosomal protein S8 from Bacillus stearothermophilus at 1.9 Å resolution. Structure 4, 1093–1104 ( 1996).
Nevskaya, N. et al. Crystal structure of ribosomal protein S8 from Thermus thermophilus reveals a high degree of structural conservation of a specific RNA binding site. J. Mol. Biol. 279, 233 –244 (1998).
Berglund, H., Rak, A., Serganov, A., Garber, M. & Härd, T. Solution structure of the ribosomal RNA binding protein S15 from Thermus thermophilus. Nature Struct. Biol. 4, 20–23 (1997).
Clemons, W. M., Davies, C., White, S. W. & Ramakrishnan, V. Conformational variability of the N-terminal helix in the structure of ribosomal protein S15. Structure 6, 429–438 (1998).
Allard, P. Another piece of the ribosome: Solution structure of S16 and its location in the 30S subunit. Structure 8, 875– 882 (2000).
Golden, B. L., Hoffman, D. W., Ramakrishnan, V. & White, S. W. Ribosomal protein S17: characterization of the three-dimensional structure by 1H- and 15N-NMR. Biochemistry 32, 12812–12820 ( 1993).
Helgstrand, M. et al. Solution structure of the ribosomal protein S19 from Thermus thermophilus. J. Mol. Biol. 292, 1071 –1081 (2000).
Acknowledgements
This work was supported by the Medical Research Council (UK) and a US National Institutes of Health grant to V.R. and S. W. White. Beamlines at Argonne and Brookhaven were supported by the US Department of Energy. D.E.B. was supported by an EMBO long-term postdoctoral fellowship,and W.M.C. by an NIH predoctoral fellowship. We thank B. S. Brunschwig and M. H. Chou for gifts of osmium hexammine and osmium bipyridine; T. Terwilliger for help with phasing using SOLVE; T. A. Leaf-Jones for providing us a version of O with RNA tools; and our colleagues at the LMB for their advice and encouragement. We are indebted to A. Joachimiak, S. L. Ginell, R. Ravelli, S. McSweeney, G. Leonard, A. Thompson, H. Lewis, L. Berman, M. Papiz, S. Girdwood and M. MacDonald for help and advice on synchrotron beamlines.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Wimberly, B., Brodersen, D., Clemons, W. et al. Structure of the 30S ribosomal subunit. Nature 407, 327–339 (2000). https://doi.org/10.1038/35030006
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1038/35030006
This article is cited by
-
The translating bacterial ribosome at 1.55 Å resolution generated by cryo-EM imaging services
Nature Communications (2023)
-
Annotating Macromolecular Complexes in the Protein Data Bank: Improving the FAIRness of Structure Data
Scientific Data (2023)
-
The paths to the atomic structures of proteins and nucleic acids
ChemTexts (2023)
-
Adaptation to genome decay in the structure of the smallest eukaryotic ribosome
Nature Communications (2022)
-
Fractional 2′-O-methylation in the ribosomal RNA of Dictyostelium discoideum supports ribosome heterogeneity in Amoebozoa
Scientific Reports (2022)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.