Abstract
Some marine worms, such as Thelepus crispus and Notomastus lobatus, secrete brominated aromatic molecules and other halogenated metabolites as repellants1. Other species, such as Amphitrite ornata, do not produce repellants but are adapted to the chemical warfare of N. lobatus and cohabit with them in estuarine mudflats. The halocompounds are tolerated by A. ornata as they are degraded by dehaloperoxidase (DHP)2. We have determined the amino-acid sequence and crystal structure of DHP and find that its fold is typical of the globin family, indicating that the enzyme evolved from an oxygen carrier protein. Residues at the dimer interface do not correspond to those in tetrameric and dimeric haemoglobins and the spatial arrangement of the dimers is different. The complete amino-acid sequence is most similar to that of myoglobin from the sea hare, with 20.6% identity among 126 overlapping amino acids.
Similar content being viewed by others
Main
The halophenols bind in the distal pocket of DHP, indicating that DHP removes the halogen by directly attacking an activated oxygen atom. If this is the case, then DHP represents a new class of enzyme: a dehaloperoxygenase. Its mechanism of action is probably similar to that of other peroxidases, given the important role of the high-valency iron–oxo intermediate that is formed when hydrogen peroxide is added to the enzyme and the subsequent heterolytic cleavage of the O–O bond3. The ability of peroxidases to form such an intermediate, which globins cannot, is consistent with the idea4 that the proximal histidine and polar residues in the distal pocket are crucial.
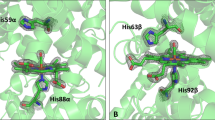
Peroxidases form a strong hydrogen bond between the Nδ1 nitrogen atom of the proximal histidine and a nearby glutamate, so the proximal histidine takes on a partial histidinate character. In contrast, the proximal pocket of globins lacks a glutamate, so they form only a weak hydrogen bond4. In DHP, a hydrogen bond forms that is stronger than in globins but, rather than bonding to a carboxylate, it is generated by a roughly 60° rotation of the imidazole ring. Its hydrogen points directly into the oxygen atom of the peptide carbonyl group of the leucine at residue 83. The hydrogen bond is shorter than that of myoglobin, in which it is bifurcated (Fig. 1). The reorientation of the axial ligand is facilitated by a shift of the proximal histidine in the sequence by two residues.
In peroxidases, the distal histidine functions as an acid or base in the transfer of protons to the leaving water molecule, and the developing negative charge is stabilized by an arginine side chain4. This arrangement is possible because peroxidases do not use the distal cavity as an organic-substrate-binding pocket, as the reaction takes place at the haem edge5.
We propose that DHP binds peroxide and uses the distal histidine to cleave the O–O bond. When the high-valency iron–oxo intermediate is ready, the distal histidine swings out of the cavity, enabling the substrate to enter the distal pocket and undergo oxidation. This hypothesis is supported by the observation that the distal histidine in the native DHP is disordered between two positions, with one outside the distal pocket and the other inside (Fig. 1), whereas it is outside the pocket in the complex with 4-iodophenol.
The distance between the Nε2 atom of the distal histidine and the haem iron in globins is 4.1 to 4.6 Å, whereas in peroxidases it is 5.5 to 6.0 Å (ref. 6). In DHP, this distance is 5.4 Å for the position inside the distal cavity, which is related to a 2.4 Å shift of its α-carbon compared with myoglobin. The shift appears to be generated by the replacement of the glycine that follows the distal histidine in myoglobin by threonine in DHP.
DHP serves primarily to dehalogenate biologically generated bromophenols, but it can also dehalogenate trichlorophenols and other anthropogenic pollutants. The efficiency of the enzyme is not significantly affected by the position and number of halogen atoms at the phenol ring2. The enzyme simply binds its organic substrate close to a highly reactive oxygen atom and so it does not need a system involving hydrogen bonds and other interactions that are features of enzymes with more sophisticated mechanisms. This mechanism means that its specificity can be altered by modifying the residues lining the distal cavity, making it potentially useful as a tool for bioremediation projects.
References
Woodin, S. A., Marinelli, R. L. & Lincoln, D. E. J. Chem. Ecol. 19, 517–530 (1993).
Chen, Y. P., Woodin, S. A., Lincoln, D. E. & Lovell, C. R. J. Biol. Chem. 271, 4609–4612 (1996).
Dawson, J. H. Science 240, 433–439 (1988).
Poulos, T. L. Adv. Inorg. Biochem. 7, 1–36 (1988).
Ator, M. A. & Ortiz de Montellano, P. R. J. Biol. Chem. 262, 1542–1551 (1987).
Matsui, T., Ozaki, S., Liong, E., Phillips, G. N. & Watanabe, Y. J. Biol. Chem. 274, 2838–2844 (1999).
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Lebioda, L., LaCount, M., Zhang, E. et al. An enzymatic globin from a marine worm. Nature 401, 445 (1999). https://doi.org/10.1038/46728
Issue Date:
DOI: https://doi.org/10.1038/46728
This article is cited by
-
Selective tuning of activity in a multifunctional enzyme as revealed in the F21W mutant of dehaloperoxidase B from Amphitrite ornata
JBIC Journal of Biological Inorganic Chemistry (2018)
-
Degradation of sulfide by dehaloperoxidase-hemoglobin from Amphitrite ornata
JBIC Journal of Biological Inorganic Chemistry (2011)
-
The haemoglobin enzyme
Nature (1999)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.