Abstract
Panels of immunomarkers can provide greater information than single markers, but the problem of how to optimally interpret data from multiple immunomarkers is unresolved. We examined the expression profile of 12 immunomarkers in 200 endometrial carcinomas using a tissue microarray. The outcomes of groups of patients were analyzed by using the Kaplan–Meier method, using the log-rank statistic for comparison of survival curves. Correlation between clustering results and traditional prognosticators of endometrial carcinoma was examined by either Fisher's exact test or χ2-test. Multivariate analysis was performed using a proportional hazards method (Cox regression modeling). Seven of the 12 immunomarkers studied showed prognostic significance in univariate analysis (P<0.05) and 1 marker showed a trend toward significance (P=0.06). These eight markers were used in unsupervised hierarchical clustering of the cases, and resulted in identification of three cluster groups. There was a statistically significant difference in patient survival between these cluster groups (P=0.0001). The prognostic significance of the cluster groups was independent of tumor stage and patient age on multivariate analysis (P=0.014), but was not of independent significance when either tumor grade or cell type was added to the model. The cluster group designation was strongly correlated with tumor grade, stage, and cell type (P<0.0001 for each). Interlaboratory reproducibility of subclassification of endometrial adenocarcinoma by hierarchical clustering analysis was verified by showing highly reproducible assignment of individual cases to specific cluster groups when the immunostaining was performed, interpreted, and clustered in a second laboratory (κ=0.79, concordance rate=89.6%). Unsupervised hierarchical clustering of immunostaining data identifies prognostically relevant subsets of endometrial adenocarcinoma. Such analysis is reproducible, showing less interobserver variability than histopathological assessment of tumor cell type or grade.
Similar content being viewed by others
Main
Endometrial carcinoma is the most common invasive carcinoma of the female genital tract and the fourth most frequently diagnosed cancer in North American women.1 Endometrial carcinoma is not a single disease process and a range of clinical behaviors are encountered.2 For example, low-grade endometrioid carcinomas are indolent tumors arising in pre- and peri-menopausal women in the setting of estrogen excess, while serous carcinomas are aggressive, high-grade tumors, unrelated to estrogenic stimulation. Current treatment recommendations for patients with endometrial carcinoma vary widely and are based on tumor stage, grade, and cell type.3, 4
Morphology alone has proven insufficient for subclassification of some human tumors, such as breast carcinoma and non-Hodgkin's lymphoma, where immunostaining or other molecular markers are routinely used. The subclasses identified through these adjuvant studies influence patient management and outcomes. While adjuvant studies are not routinely used for gynecological cancers, they offer the only realistic hope of significant progress in subclassification, given the limitations of routine morphological examination. An important limitation of routine histopathological examination of endometrial carcinoma is the suboptimal reproducibility of both cell-type assignment and tumor grade, both of which are used to guide patient therapy type.5, 6, 7, 8, 9, 10, 11, 12 No single immunomarker has been found to be sufficiently powerful to merit routine use in the subclassification of endometrial carcinoma, so we have sought to use a panel of immunomarkers in this study.
The use of multiple markers for subclassification raises the question of how to optimally evaluate the information provided by these markers. Clustering analysis, which is widely used in marketing and the social sciences, is one approach to the analysis of complex data sets with multiple markers per case, grouping cases based on the overall relatedness of marker expression.13, 14, 15 The TNM classification is a form of supervised clustering analysis combining tumor size, lymph node status, and distant metastasis. Unsupervised hierarchical clustering analysis has been used to classify tumors based on mRNA expression levels of thousands of genes; prognostically relevant cluster groups have been identified for breast cancer, large B-cell lymphoma, and lung adenocarcinoma using this approach.16, 17, 18, 19 While global gene expression profiling is a powerful tool, significant problems related to cost and quality assurance must be overcome before it is widely used clinically. Tissue can be sent for RT-PCR of multiple markers20 but this approach is very expensive and, as with global expression profiling, it is not possible to be certain if the tissue submitted for analysis reflects the tumor as a whole, as morphological correlation is not possible. Immunohistochemical staining performed on formalin-fixed, paraffin-embedded sections is an attractive alternative approach to generating molecular profiles of tumors, with the advantage of being readily available technology that allows assessment of morphology at the same time as biomarkers. We and others have shown that it is possible to subclassify breast carcinoma using hierarchical clustering analysis of a limited panel of immunomarkers.15, 21, 22, 23 We also demonstrated that the cluster groups so identified are highly prognostically significant, and that the combination of multiple markers is more powerful than any of the individual markers used.15
In this study we examined the expression profile of a panel of molecular markers in a large set of endometrial carcinomas by using immunostaining of tissue microarrays. These tumors were then subclassified based on the relatedness of their immunostaining profiles using unsupervised hierarchical clustering analysis. The aims of this study were (1) to test the ability of unsupervised hierarchical clustering analysis to identify prognostically significant subsets of patients with endometrial carcinoma (cluster groups) and (2) to examine the correlation of the cluster groups with patient outcome, and other clinical and pathological features. To demonstrate the feasibility of this approach in a clinical setting, we examined interlaboratory reproducibility of assigning individual cases to cluster groups.
Material and methods
Tumors and Patients
Two hundred consecutive cases of endometrial carcinoma, treated by hysterectomy, were retrieved from the archives of the Department of Pathology, Vancouver General Hospital, for the period 1983–1998. None of the included cases received preoperative radiotherapy or chemotherapy. There were 156 cases of endometrioid, 13 papillary serous, 5 clear cell, 4 small cell, and 22 mixed subtypes. Hematoxylin and eosin (H&E) stained slides and follow up data were available for all cases. For each case the slides were reviewed, diagnosis confirmed, and cell type assessed. The patients were staged and the tumors graded according to 1988 International Federation of Gynecology and Obstetrics (FIGO) criteria.24 This series of cases has been the subject of a previous study on grading of endometrial carcinoma.5
Tissue Microarray
While reviewing the H&E-stained slides, a slide with representative tumor was selected from each case, and an area of tumor on the selected H&E slide was circled. The corresponding paraffin block was retrieved from the hospital archives and the marked slide was aligned with the surface of the corresponding donor block to guide marking the selected area on the paraffin block with a felt marker. Using a tissue microarrayer (Beecher Instruments, Silver Spring, MD, USA), the area of interest from the donor block was cored twice with a 0.6-mm diameter cylinder and transferred to a recipient paraffin block. Sections from these arrays were then stained with H&E, to assess adequacy. In mixed tumors, the higher grade areas were selected for tissue microarray construction, as the prognosis in these mixed tumors is related to this component.25
Immunohistochemistry and Hierarchal Clustering Analysis
The avidin–biotin (ABC) method was used for immunostaining and applied to formalin-fixed and paraffin-embedded tissue. Serial sections of the recipient paraffin blocks were cut at 3 μm, deparaffinized with xylene, and rehydrated through a series of graded alcohols. Sections were stained with the panel of antibodies listed in Table 1. An automated stainer (Ventana, Tucson, AZ, USA) was used, according to the manufacturer's guidelines, for bcl-2, ER, p53, HER2, B72.3, p63, and CK5/6. Immunostaining with the remaining antibodies (p21, p27, E2F-1, PTEN, Gfi-1) was performed manually. The polyclonal anti-PTEN antibody was a gift from Dr Heung-Chin Cheng, University of Melbourne, Melbourne, Australia, and the monoclonal anti-Gfi-1 antibody was a gift from Dr H Leigh Grimes, Cincinnati Children's Hospital Medical Center, Cincinnati, Ohio. Antigen retrieval was carried out as indicated in Table 1. This panel of antibodies was chosen based on previous reports of their being of prognostic significance or their known involvement in cell-cycle regulation or hormonal signaling (reviewed in Prat4).
Sections stained with the various antibodies were then scored by two pathologists (AA and CBG), using a two-headed microscope, using a 4 point scale where 0=negative (<5% of cells staining), 1=weak staining (ie 5–50% of cells showing weak-to-intermediate intensity staining in an appropriate subcellular distribution), 2=strong staining (ie >50% of cells showing weak-to-moderately intense staining or >5% of cells showing strong staining, in an appropriate subcellular distribution), and 3=uninterpretable (eg because of loss of the tissue or excessive background staining). Scores were entered into a Microsoft Excel spreadsheet. Unsatisfactory results were eliminated from further consideration. Score results for duplicate cores were consolidated into one score where a higher score for positive immunostaining would always supercede a weaker, negative, or uninterpretable result. Cases that did not have interpretable staining results for more than 80% of the antibodies used in our panel were also excluded from the analysis.
For univariate survival analysis, scoring results were simplified into either negative (score of 0) or positive (score of 1 or 2); an exception was p53 staining, where we have previously shown that a cutoff of 50% best served to discriminate between favorable and unfavorable outcomes26 so that a p53 score of 0 or 1 was considered negative and 2 was considered positive, for survival analysis. For clustering analysis the unsimplified data were used.
Hierarchal clustering analysis of our tissue microarray data was performed using software tools that were originally designed for analyzing cDNA microarray data. An Excel macro, TMA-Deconvoluter, was designed and written specifically for processing of raw tissue microarray staining data into a format compatible for use with the previously developed Cluster software.14, 27 The clustered data were then graphically viewed using TreeView (Cluster and TreeView software are freely available programs that can be accessed at http://rana.lbl.gov/EisenSoftware.htm).
Testing Interlaboratory Reproducibility
To test interlaboratory reproducibility, unstained sections of this tissue microarray were stained in a second laboratory (Foothills Hospital, Calgary, Alberta) in the case of antibodies in routine diagnostic use in that laboratory (ER, bcl-2, p53, HER2) using a Ventana immunostainer. The anti-ER and anti-bcl-2 antibodies were purchased pre-diluted from Ventanna (clones 6F11 and bcl-2/100/D5, respectively), and used after heat antigen retrieval. The anti-p53 and anti-HER2 antibodies used were the same as those listed in Table 1, at dilutions of 1:100 and 1:200, respectively, after heat antigen retrieval. Fresh sections were stained with the antibodies not in routine use in that laboratory (PTEN, p21, p27, E2F-1) by the same manual method described previously (see above). The slides were then interpreted using the scoring system described previously, by two different pathologists (MA and RP) using a two-headed microscope, and the immunostaining results analyzed by hierarchical clustering analysis, as described previously. There was no training of the pathologists at the second laboratory before analysis of the immunostaining results. Cluster group assignment based on this analysis was compared to the original cluster group designations.
Statistical Analysis
The Kaplan–Meier method was used to construct disease-specific survival curves for patients based on expression results for each molecular marker used and also for each of the cluster groups identified by clustering analysis of the expanded expression profile based on eight immunomarkers. Comparison of curves was carried out using log-rank statistic. Time to event was defined as disease-specific survival from initial hysterectomy to date of death due to endometrial carcinoma, with all others considered censored. Correlation between clustering result and traditional prognosticators of endometrial carcinoma was examined by either Fisher's exact or χ2-test. Multivariate analysis was performed using Cox's proportional hazards method. For all analyses, two-sided tests of significance were used with α of 0.05. Interobserver variation was tested by calculation of κ-statistics, as described previously.5 All analyses were performed using SPSS software version 11.0 (Chicago, USA).
Results
The median follow-up was 77 months (range, 10 days to 224 months). Forty-six patients died of endometrial carcinoma (23%), and 36 (18%) died of unrelated causes. The median patient age was 66 years (range, 28–95 years). There were 152 FIGO stage I, 22 stage II, 25 stage III, and 1 stage IV tumors. Ninety nine tumors were FIGO grade 1, 27 grade 2, and 74 grade 3, reflecting a referral bias, with higher grade and stage tumors more likely to be referred to our center for subspecialist care.
Table 2 shows the expression profile and prognostic value, in univariate analysis, of all antibodies used in tissue microarray sections. Only seven of these markers were found to be of prognostic significance in endometrial carcinoma (p53, ER, bcl-2, HER2, p27, E2F-1, PTEN). p21 approached significance as a prognostic indicator in univariate analysis (P=0.06). Expression of these eight markers in tumors of different cell type and grade is shown in Table 3. Staining of the mixed tumors is not presented in this table. Although there were significant differences in expression of these immunomarkers between the groups of tumor, stratified based on cell type and grade, none of the markers were completely sensitive or specific for either cell type or grade.
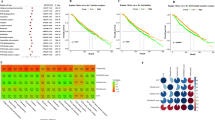
The clustering analysis of study cases was performed based on the expression profile of eight molecular markers (p53, ER, bcl-2, HER2, p27, E2F-1, PTEN, p21). In the graphical display of the clustering results using the Treeview program (Figure 1) the rows represent individual cases and the columns are the results of immunostaining with individual antibodies. The green color indicates negative staining, red indicates positive staining (with light red for weak positivity and dark red for strong positivity), while gray represents missing data (absent tumor core or uninterpretable staining result). The clustering analysis divided the tumors into three groups based on the relatedness of their immunostaining profiles, which we have designated as cluster groups I, II, and III. As can be seen from the Treeview output (Figure 1), the first cluster group was characterized by ER expression (114 out of 114 cases), with only 20% showing bcl-2 expression, and with most tumors being negative for the remaining markers. All cases in cluster group III were ER negative; they were positive for PTEN and p53 in most cases, and showed variable expression of p27, p21, and E2F-1. All cases in cluster group II showed ER positivity (33 out of 33 cases) and most tumors in this group were also positive for p53, p21, and/or p27. Three tumors were outliers and failed to cluster into any of these groups.
Results of hierarchical clustering analysis, with identification of three cluster groups (designated I, II, and III). Each row represents a single case and each column a single immunomarker. Green indicates negative immunostaining, light red indicates weak immunostaining, and dark red strong immunoreactivity. The dendrogram on the left indicates the relatedness of the immunoprofiles of individual cases, while the dendrogram at the top indicates the relatedness of cases stained by each of the markers; the longer the dendrogram arm, the greater the differences.
The distribution of tumor cell type and FIGO grade in the three cluster groups is illustrated in Figure 2. Eighty-three percentage of tumors in cluster group III were of FIGO grade 3, and only 31% of tumors in this cluster were of endometrioid type. On the other hand, 90% of tumors in cluster group I were of endometrioid type and 80% were grade 1 or 2. Cluster group II was made up of predominantly grade 2 and grade 3 endometrioid carcinomas and grade 3 mixed tumors. The cluster group designation was strongly significantly correlated with tumor grade, stage, and cell type (P<0.0001 for each).
Kaplan–Meier survival curves were constructed based on the cluster group designation as determined by clustering analysis (Figure 3). There was a statistically significant difference in the patient survival based on cluster group assignment (P=0.0001). The difference in outcome between patients in cluster groups I and II was not significant (P=0.25).
In multivariate analysis, using the variables of patient age (≤55 years vs >55 years old), cluster group designation (cluster groups I and II vs cluster group III), and tumor stage (stages I and II vs stages III and IV) in regression modeling, the cluster group designation was prognostically significantly independent of patient stage and age (P=0.014; data not shown). When the analysis was expanded to include cluster group designation, tumor stage, patient age, FIGO grade, and tumor cell type as variables, only FIGO grade, patient age, and FIGO stage were of independent significance, while tumor cell type and cluster group designation approached but did not reach independent prognostic significance (0.05<P<0.10; Table 4).
After re-staining and re-analysis of the cases by an independent team of pathologists, the clustering was repeated and again identified three cluster groups, with significantly different patient outcomes for patients in the three different cluster groups (data not shown). Interlaboratory reproducibility in assignment of individual cases to cluster groups was compared and there was very good reproducibility (κ=0.79, concordance in case assignment in 174/194 (89.6%) of cases).
Discussion
Endometrial carcinoma is the most common female genital tract cancer and has the fourth highest overall incidence in women, following breast, lung, and colorectal carcinoma.1 In endometrial carcinoma, the pathologist's role is pivotal in prognostication and in guiding the use of postoperative adjuvant therapy. In early-stage endometrial carcinoma (stages I and II), depth of myometrial invasion (substage), histological grade, and tumor cell type are important predictors of patient outcome and used to guide therapy.2, 4 A dualistic model of endometrial carcinogenesis is widely accepted.5, 28, 29, 30, 31, 32, 33, 34 Most endometrial carcinomas are of type I, arising from precursor hyperplasia driven by estrogen excess. These tumors express ER and PR, may have microsatellite instability (20%), and may have PTEN mutation (40%). Typically these are low-grade carcinomas and are endometrioid in type, with a low proliferative index. Type II tumors arise independent of estrogen excess and the most commonly described genetic abnormality is mutation of p53. p53 mutation is almost always associated with aneuploidy and does not appear to coexist with PTEN mutations. Type II tumors are high-grade carcinomas, typically serous in type, and have a high proliferative index. Not all endometrial carcinoma fit into this dualistic model, and the type I and type II designations describe ‘two loose clinicopathological clusters’ rather than firm diagnostic categories.35 They do not, for example, account for mixed serous/endometrioid tumors of the endometrium. While the type I and type II designations are helpful in understanding the pathogenesis of endometrial carcinoma, a more clinically applicable subclassification system is desirable.
We stained a series of cases of endometrial carcinoma seen consecutively at our center with a panel of antibodies. This panel was chosen primarily based on their known role in endometrial carcinogenesis or previously demonstrated significance as prognostic markers in endometrial carcinoma (p53,26, 36 PTEN,37, 38 ER and PR,39, 40 bcl-2,41, 42, 43 HER241, 44, 45) or their role in cell-cycle regulation (p21, p27, E2F-1) (reviewed in Israels and Israels46). We did not use MIB-1 (Ki67) immunostaining as we have previously shown that it shows only a weak correlation with mitotic activity in uterine serous carcinomas.28 Additional markers included proteins implicated in regulation of hormone response (Gfi-147), markers of squamous differentiation (and thus possible markers of endometrioid cell type; CK5/6 and p6348, 49), and a protein associated with serous differentiation (B72.350). Not surprisingly, given the criteria for selection of the immunopanel, most markers studied were prognostically significant in univariate analysis.
We next applied unsupervised (ie independent of all clinical and pathological parameters, apart from the immunostaining) hierarchical clustering analysis to the immunostaining data for the markers that approached or reached approached significance in univariate analysis. We were the first to apply hierarchical clustering analysis to immunostaining data13 and have previously used this approach (ie univariate survival analysis of individual markers followed by unsupervised hierarchical clustering analysis) to demonstrate that, for breast cancer, the use of multiple markers is superior to single markers for prognostication, and that different clustering algorithms tested yielded equivalent results in assignment of individual cases to specific cluster groups.15 Bair and Tibshirani51 independently described using this same analytical approach (selecting markers based on prognostic significance, followed by unsupervised clustering analysis) for analysis of gene expression data. They described this approach as ‘semi-supervised’ and were able to show the superiority of semi-supervised approaches over purely supervised or purely unsupervised analyses, using multiple sets of gene expression data from breast carcinoma and large-cell lymphoma. In endometrial carcinoma we identified three prognostically significant cluster groups by clustering analysis. Although hierarchical clustering was carried out based solely on immunostaining, independent of any clinical or pathological parameters, the cluster groups show a very strong correlation with these parameters, in particular tumor cell type and grade. Cluster group I consists predominantly of low-grade endometrioid carcinomas and corresponds to type I endometrial carcinoma. These tumors are ER positive and negative for proliferation markers or p53, and had the most favorable prognosis. Cluster group III consists predominantly of high-grade serous, clear cell, and mixed (serous and endometrioid) carcinomas, with a smaller number of high-grade endometrioid carcinomas, and has molecular features of type II endometrial carcinoma. These tumors were all ER negative, and expressed p53 and/or PTEN, consistent with known molecular events during carcinogenesis of type II tumors. This group had the least favorable outcome. Cluster group II includes tumors that were ER positive and showed p53, p21, and p27 expression. This group consists mainly of grades 2 and 3 endometrioid carcinomas, with a lesser number of mixed tumors and occasional serous and grade 1 endometrioid carcinomas. This group had an intermediate prognosis, and based on their immunoprofile these may represent examples of progression of type I tumors to more aggressive tumors. In at least some cases this occurs through mutation of p53 but, unlike type II tumors, these tumors are ER positive. The tumors in cluster group II are also more likely to express proliferation markers. It has previously been proposed that high-grade endometrial carcinoma may arise from two pathways, either de novo through p53 mutations (classic type II endometrial carcinoma) or through a pre-existing low-grade (type I) endometrioid carcinoma, arising in an environment of estrogen excess with accumulation of additional mutations and progression to a higher grade tumor.52 Our observations support this view, with the tumors in cluster groups II and III, respectively, probably representing examples of these two pathways.
This is the first study to assess the reproducibility of tumor classification based on an extended panel of immunomarkers. Immunostains for individual proteins can show a very high degree of variability, depending on the specific antibody chosen, antigen retrieval, detection method, etc. This is well demonstrated in a recent study by Pallares et al53 in which they found no significant correlation in immunostaining results obtained using four different commercially available anti-PTEN antibodies, and no correlation with PTEN mutational status. This is an all too common experience with experimental antibodies. In contrast, there can be a very high degree of interlaboratory agreement for antibodies routinely used in diagnostic laboratories. For example, in a comparison of ER staining between six laboratories, where the same cases were stained and interpreted according to each laboratories existing protocols, we found there to be excellent interlaboratory agreement (κ=0.84).54 In the current study we attempted to recapitulate clinical practice by having the slides stained in an independent laboratory, and interpreted by a different team of pathologists. These pathologists used the same guidelines for interpretation described in Material and methods, without a training session (ie their scoring was based on the application of written cutoffs, independent of the first set of pathologists, and without a teaching session to reduce interobserver variation in interpretation). Furthermore, the immunostaining protocols were those in current use in their laboratory, without an attempt to replicate the staining protocols used in the first round of immunostaining. We performed this only with those antibodies in routine clinical use (ER, bcl-2, p53, HER2); for antibodies not in routine use diagnostically this strategy could not be employed and staining was repeated in the originating laboratory and then interpreted by the second set of pathologists. The overall reproducibility of assignment of individual cases to specific cluster groups was very good (κ=0.79). This compares very favorably with interobserver reproducibility in assessment of either cell type or grade. The reported κ-value for interobserver variation in assessment of cell type (serous or clear cell vs endometrioid) is 0.70, and for interobserver variation of tumor grade by the FIGO system, the range of reported κ-values is 0.41–0.65.5, 6, 7, 8, 9, 10, 11, 12
There are strong correlations between tumor cell type, grade, and cluster group designation, which are the most important prognostic variables based on the characteristics of the tumor cells (as opposed to assessment of the extent of tumor spread, or stage). These three variables are, in theory, assessed independently but in practice cell type and grade are dependent, as serous and clear cell tumors are all considered to be high grade. In multivariate analysis, only tumor grade was of prognostic significance independent of tumor stage and patient age. Both tumor cell type and cluster designation approached significance as independent markers. Assignment of cases to cluster groups offers significant advantages over assessment of cell type or grade. It is performed without consideration of clinical information or routinely assessed histopathological variables, and shows better reproducibility than conventional assessment of cell type and grade. As well, the prognostic significance will improve with the use of better markers. Development of new markers to reflect the different pathways during carcinogenesis is particularly desirable. In contrast, it is likely that neither assessment of tumor cell type nor grade, as currently performed, has potential for significant improvement in either prognostication or reproducibility.
Important characteristics of a clinically relevant subclassification system for cancer are that it is reproducible, reflects genetic changes during oncogenesis (ie is biologically relevant), and is predictive of response to treatment and outcome. There has been substantial progress in achieving these goals in the case of breast carcinoma, through the use of panels of immunomarkers.15, 22, 23 We provide evidence that, in the future, the use of a panel of biologically relevant and technically well-validated immunomarkers could similarly allow reproducible subclassification of endometrial carcinoma and be used to guide therapeutic decision making.
References
Greenlee RT, Murray T, Bolden S, et al. Cancer statistics, 2000. CA Cancer J Clin 2000;50:7–33.
Tavassoli FA, Devilee P . WHO Classification of Tumors: Pathology and Genetics Tumors of the Breast and Female Genital Organs. IARC Press: Lyon, France, 2003.
Goff B . Uterine papillary serous carcinoma: what have we learned over the past quarter century? Gynecol Oncol 2005;98:341–343.
Prat J . Prognostic parameters of endometrial carcinoma. Hum Pathol 2004;35:649–662.
Alkushi A, Abdul-Rahman ZH, Lim P, et al. Description of a novel system for grading of endometrial carcinoma and comparison with existing grading systems. Am J Surg Pathol 2005;29:295–304.
Nedergaard L, Jacobsen M, Andersen JE . Interobserver agreement for tumour type, grade of differentiation and stage in endometrial carcinomas. APMIS 1995;103:511–518.
Nielsen AL, Thomsen HK, Nyholm HC . Evaluation of the reproducibility of the revised 1988 International Federation of Gynecology and Obstetrics grading system of endometrial cancers with special emphasis on nuclear grading. Cancer 1991;68:2303–2309.
Nordstrom B, Strang P, Lindgren A, et al. Carcinoma of the endometrium: do nuclear grade and DNA ploidy provide more prognostic information than do the FIGO and WHO classifications? Int J Gynecol Pathol 1996;15:191–201.
Lax SF, Kurman RJ, Pizer ES, et al. A binary architectural grading system for uterine endometrial endometrioid carcinoma has superior reproducibility compared with FIGO grading and identifies subsets of advance-stage tumors with favorable and unfavorable prognosis. Am J Surg Pathol 2000;24:1201–1208.
Scholten AN, Smit VTHBM, Beerman H, et al. Prognostic significance and interobserver variability of histologic grading systems for endometrial carcinoma. Cancer 2004;100:764–772.
Taylor RR, Zeller J, Lieberman RW, et al. An analysis of two versus three grades for endometrial carcinoma. Gynecol Oncol 1999;74:3–6.
Zaino RJ, Kurman RJ, Diana KL, et al. The utility of the revised international federation of gynecology and obstetrics histologic grading of endometrial adenocarcinoma using a defined nuclear grading system. A Gynecologic Oncology Group study. Cancer 1995;75:81–86.
Al-Kushi A, Irving J, Hsu F, et al. Immunoprofile of cervical and endometrial adenocarcinomas using a tissue microarray. Virchows Archiv 2003;442:271–277.
Eisen MB, Spellman PT, Brown PO, et al. Cluster analysis and display of genome-wide expression patterns. Proc Natl Acad Sci USA 1998;95:14863–14868.
Makretsov NA, Huntsman DG, Nielsen TO, et al. Hierarchical clustering analysis of tissue microarray immunostaining data identifies prognostically significant groups of breast carcinoma. Clin Cancer Res 2004;15:6143–6151.
Alizadeh AA, Eisen MB, Davis RE, et al. Distinct types of diffuse large B-cell lymphoma identified by gene expression profiling. Nature 2000;403:503–511.
Garber ME, Troyanskaya OG, Schluens K, et al. Diversity of gene expression in adenocarcinoma of the lung. Proc Natl Acad Sci USA 2001;98:13784–13789.
Sorlie T, Tibshirani R, Parker J, et al. Repeated observation of breast tumor subtypes in independent gene expression data sets. Proc Natl Acad Sci USA 2003;100:8418–8423.
van’t Veer LJ, Dai H, van de Vijver MJ, et al. Gene expression profiling predicts clinical outcome of breast cancer. Nature 2002;415:530–536.
Paik S, Shak S, Tang G, et al. A multigene assay to predict recurrence of tamoxifen-treated, node-negative breast cancer. N Engl J Med 2004;351:2817–2826.
Callagy G, Cattaneo E, Daigo Y, et al. Molecular classification of breast carcinomas using TMAs. Diagn Mol Pathol 2003;12:27–34.
Dalia M, El-Rehim A, Ball G, et al. High-throughput protein expression analysis using tissue microarray technology of a well-characterized series identifies biologically distinct classes of breast cancer confirming recent cDNA expression analyses. Int J Cancer 2005;3:340–350.
Jacquemier J, Ginestier C, Rougemont J, et al. Protein expression profiling identifies subclasses of breast cancer and predicts prognosis. Cancer Res 2005;65:767–779.
Creasman WT . Announcements: FIGO stages—1988 revision. Gynecol Oncol 1989;35:125–127.
Sherman ME, Bitterman P, Rosenshein NB, et al. Uterine serous carcinoma. A morphologically diverse neoplasm with unifying clinicopathologic features. Am J Surg Pathol 1992;16:600–610.
Alkushi A, Lim P, Coldman A, et al. Interpretation of p53 immunohistochemical staining in endometrial carcinoma: establishing a clinically relevant cut-off. Int J Gynecol Pathol 2004;23:129–137.
Liu CL, Prapong W, Natkunam Y, et al. Software tools for high-throughput analysis and archiving of immunohistochemistry staining data obtained with TMAs. Am J Pathol 2002;161:1557–1565.
Al-Kushi M, Lim P, Aquino-Parsons C, et al. Markers of proliferative activity are predictors of patient outcome for low-grade endometrioid carcinoma but not papillary serous carcinoma of the endometrium. Mod Pathol 2002;15:365–371.
Bokhman JV . Two pathogenetic types of endometrial carcinoma. Gynecol Oncol 1983;15:10–17.
Koshiyama M, Ueta M . Two kinds of endometrial neoplasia arising from different origins in the uterine corpus: comparison of p53 expression and sex steroid receptor status. Eur J Obstet Gynecol Reprod Biol 2002;104:167–170.
Lax SF, Kurman RJ . A dualistic model for endometrial carcinogenesis based on immunohistochemical and molecular genetic analyses. Verh Dtsch Ges Pathol 1997;81:228–232.
Lax SF, Kendall B, Tashiro H, et al. The frequency of p53, K-ras mutations, and microsatellite instability differs in uterine endometrioid and serous carcinoma: evidence of distinct molecular genetic pathways. Cancer 2000;88:814–824.
Matias-Guiu X, Catasus L, Bussaglia E, et al. Molecular pathology of endometrial hyperplasia and carcinoma. Hum Pathol 2001;32:569–577.
Sherman ME . Theories of endometrial carcinogenesis: a multidisciplinary approach. Mod Pathol 2000;13:295–308.
Hendrickson MR, Longacre TA, Kempson RI . The uterine corpus. In: Mills SE (ed). Sternberg's Diagnostic Surgical Pathology. Lippincott, Williams and Wilkins: Philadelphia, PA, 2004, pp 2460–2484.
Silverman MB, Roche PC, Kho RM, et al. Molecular and cytogenetic pretreatment risk assessment in endometrial carcinoma. Gynecol Oncol 2000;77:1–7.
Risinger JI, Hayes K, Maxwell GL, et al. PTEN mutation in endometrial cancers is associated with favorable clinical and pathological characteristics. Clin Cancer Res 1998;4:3005–3010.
Terakawa N, Kanamori Y, Yoshida S, et al. Loss of PTEN expression followed by Akt phosphorylation is a poor prognostic factor for patients with endometrial cancer. Endocr Relat Cancer 2003;10:203–208.
Carcangiu ML, Chambers JT, Voynick I, et al. Immunohistochemical evaluation of estrogen and progesterone receptor content in 183 patients with endometrial carcinoma. Part II: correlation between biochemical and immunohistochemical methods and survival. Am J Clin Pathol 1990;93:255–260.
Kadar N, Malfetano JH, Homesley HD . Steroid receptor concentrations in endometrial carcinoma: effect on survival in surgically staged patients. Gynecol Oncol 1993;50:281–286.
Berchuk A, Rodriguez G, Keeney RB, et al. Overexpression of HER-2/neu in endometrial cancer is associated with advanced stage disease. Am J Obstet Gynecol 1991;164:15–21.
Erdem O, Erdem M, Dursun A, et al. Angiogenesis, p53, and bcl-2 expression as prognostic indicators in endometrial cancer: comparison with traditional clinicopathologic variables. Int J Gynecol Pathol 2003;22:254–260.
Santin AD, Ballone S, Van Stedman S, et al. Amplification of c-erbB2 oncogene: a major prognostic indicator in uterine serous papillary carcinoma. Cancer 2005;104:1391–1397.
Prat J, Oliva E, Lerma E, et al. Uterine papillary serous adenocarcinoma: a 10-case study of p53 and c-erbB-2 expression and DNA content. Cancer 1994;74:1778–1783.
Sakuragi N, Ohkouchi T, Hareyama H, et al. Bcl-2 expression and prognosis of patients with endometrial carcinoma. Int J Cancer 1998;79:153–158.
Israels ED, Israels LG . The cell cycle. Oncologist 2000;5:510–513.
Kazanjian A, Wallis D, Au N, et al. Growth factor independence-1 is expressed in primary human neuroendocrine lung carcinomas and mediates the differentiation of murine pulmonary neuroendocrine cells. Cancer Res 2004;64:6874–6882.
Au N, Gown A, Cheang M, et al. P63 expression in lung carcinoma: a tissue microarray study of 408 cases. Appl Immunohistochem Mol Morphol 2004;12:240–247.
Hsu F, Nielsen TO, Alkushi A, et al. Multiple tumor microarrays are an effective quality assurance tool for diagnostic immunohistochemistry laboratories. Mod Pathol 2002;15:1374–1380.
Ordonez N . Role of immunohistochemistry in distinguishing epithelial peritoneal mesotheliomas from peritoneal and ovarian serous carcinomas. Am J Surg Pathol 1998;22:1203–1214.
Bair E, Tibshirani R . Semi-supervised methods to predict patient survival from gene expression data. PLoS Biology 2004;2:e108.
Carcangiu M, Chambers JT . Uterine papillary serous carcinoma: a study of 108 cases with emphasis on prognostic significance of associated endometrioid carcinoma, absence of invasion, and concomitant ovarian carcinoma. Gynecol Oncol 1992;47:298–305.
Pallares J, Bussaglia E, Martinez-Guitarte JL, et al. Immunohistochemical analysis of PTEN in endometrial carcinoma: a tissue microarray study with a comparison of four commercial antibodies in correlation with molecular abnormalities. Mod Pathol 2005;18:719–727.
Parker R, Huntsman DG, Lesack D, et al. Assessment of interlaboratory variation in the immunohistochemical determination of estrogen receptor status using a breast cancer tissue microarray. Am J Clin Path 2002;117:723–728.
Acknowledgements
This work was supported by an unrestricted educational grant to CBG and DH from sanofiaventis.
Author information
Authors and Affiliations
Corresponding author
Additional information
Supplementary Information accompanies the paper on Modern Pathology website (http://www.nature.com/modpathol)
Supplementary information
Rights and permissions
About this article
Cite this article
Alkushi, A., Clarke, B., Akbari, M. et al. Identification of prognostically relevant and reproducible subsets of endometrial adenocarcinoma based on clustering analysis of immunostaining data. Mod Pathol 20, 1156–1165 (2007). https://doi.org/10.1038/modpathol.3800950
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/modpathol.3800950
Keywords
This article is cited by
-
In search for biomarkers and potential drug targets for uterine serous endometrial cancer
Journal of Cancer Research and Clinical Oncology (2021)
-
Systemic recurrence of endometrial cancer more than 10 years after hysterectomy: a report of two cases and a brief review of the literature
Journal of the Egyptian National Cancer Institute (2020)
-
Targeted sequencing with a customized panel to assess histological typing in endometrial carcinoma
Virchows Archiv (2019)
-
The evolution of endometrial carcinoma classification through application of immunohistochemistry and molecular diagnostics: past, present and future
Virchows Archiv (2018)
-
Practical issues in the diagnosis of serous carcinoma of the endometrium
Modern Pathology (2016)