Abstract
In situ click chemistry is a target-guided synthesis technique for discovering potent protein ligands by assembling azides and alkynes into triazoles inside the affinity site of a target protein. We report the rapid discovery of a new and potent inhibitor of bacterial chitinases by the use of in situ click chemistry. We observed a target-templated formation of a potent triazole inhibitor of the chitinase-catalyzed chitin hydrolysis, through in situ click chemistry between a biologically active azide-containing scaffold and structurally unrelated alkyne fragments. Chitinase inhibitors have chemotherapeutic potential as fungicides, pesticides and antiasthmatics. Argifin, which has been isolated and characterized as a cyclopentapeptide natural product by our research group, shows strong inhibitory activity against chitinases. As a result of our efforts at developing a chitinase inhibitor from an azide-bearing argifin fragment and the application of the chitinase template and a library of alkynes, we rapidly obtained a very potent and new 1,5-disubstituted triazole inhibitor against Serratia marcescens chitinase (SmChi) B. The new inhibitor expressed 300-fold increase in the inhibitory activity against SmChiB compared with that of argifin. To the best of our knowledge, our finding of an enzyme-made 1,5-disubstituted triazole, using in situ click chemistry is the second example reported in the literature.
Similar content being viewed by others
Introduction
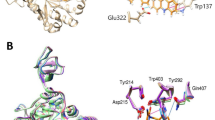
Chitin, the second most abundant polysaccharide in nature, is a constituent of fungal cell walls, the exoskeletons of crustaceans and insects, and the microfilarial sheaths of parasitic nematodes.1, 2, 3 Accumulation of chitin by organisms is modulated by chitin synthase-mediated biosynthesis and by chitinase-mediated hydrolytic degradation. Thus, chitinases are expected to be specific targets for antifungal, insecticidal and antiparasitic agents (for a review, see Andersen et al.4). They also offer significant potential for the treatment of asthma and other diseases in humans.5 Argifin (1) (Figure 1),6, 7, 8 a member of a family of natural cyclic peptides produced by microorganisms, was isolated in our laboratory from the cultured broth of the fungal strain (Gliocladium sp. FTD-0668), and was found to be a potent inhibitor of blowfly6 and Serratia marcescens chitinases (SmChi).9, 10, 11, 12 Recently, argifin complexes with fungal (Aspergillus fumigatus), human and bacterial chitinases have been resolved by X-ray crystallography.11, 12 These studies revealed that there are at least four conserved hydrogen-bond interactions between the Nω-methylcarbamoyl-L-arginine moiety and the polar groups arrayed in the hydrolytic pocket of the family 18 chitinases examined to date. The remarkable fidelity of the hydrogen-bonding network between the chitinases and the argifin ligand implicates its critical role in revealing the micro- to nanomolar range of inhibition. In fact, van Aalten and co-workers revealed through X-ray analysis that the ability of the Nω-methylcarbamoyl group to penetrate fully into the active-site pocket of chitinases strongly correlated with the inhibition of chitin hydrolysis.13 Hence, we concluded that the Nω-methylcarbamoyl-L-arginine core represents an ideal anchor to derivatize and elaborate better chitinase inhibitors.
Herein we report our early results of the design and simplification of azide-bearing Nω-methylcarbamoyl-L-arginine substrate as a smaller analog of macrocyclic peptide natural product 1 and the use of target-guided synthesis (TGS) (for reports of TGS, see Rideout,14 Rideout,15 Ingelese and Benkovic,16 Boger et al.,17 Maly et al.,18 Nicolaou et al,19 Greasley et al.,20 Nguyen and Huc,21 Nicolaou et al.,22 Kehoe et al.,23 Poulin-Kerstein and Dervan,24 and Hu et al.25) for the screening of new and more potent chitinase inhibitors, using the 1,3-dipolar cycloaddition26 between an azide ligand and a library of acetylenes. The in situ click chemistry for drug discovery is dependent on irreversibly reacting reagents that are inert under physiological conditions,27 as shown earlier by the discovery of highly potent inhibitors of acetylcholine esterase,28, 29, 30, 31 carbonic anhydrase II32 and HIV-1 protease.33
Results and discussion
Preparation of the azide-bearing inhibitors
Initially, we investigated the appropriate inhibitors to develop in situ click chemistry from the Nω-methylcarbamoyl-L-arginine scaffold. Although a simple arginine-derived inhibitor 2 discovered by van Aalten and co-workers had been reported independently,13 it showed low inhibitory activity against SmChi in contrast to argifin 1, which was examined by our group (Figure 2). We synthesized the azide-bearing inhibitor 3 as a reactive scaffold for capturing complementary acetylenic reagents to form triazole-linked inhibitors by TGS. Competition assay with 4-methylumberiferyl diacetylchitobiose (4-MU-(GluNAc)2)34, 35 showed that this azide-bearing inhibitor 3 expressed a low inhibitory activity similar to that of the azide-lacking inhibitor 2, which are in striking contrast to the potency of the natural product 1. Hence, amide derivatives of azide 3 with amines other than methylamine were made and tested to see whether the binding could be restored to a level that would make azide 3 a sufficiently good anchor at the active site, to be used for the capture of alkyne-bearing candidates through in situ triazole formation. Fortunately, the dibenzylamide analog 5 of azide 3 emerged as a potent inhibitor (0.045 and 0.58 μM IC50 values against SmChiA and B, respectively). Interestingly, the IC50 value against SmChiB of 5 was 10-fold stronger than that of parent 1. The monobenzylamide 4 was also active, but less than 5. As seen in Figure 2, compounds 2–5 can be ranked by inhibition constants as 5>4>3≈2. We therefore used potent azide analog 5 as a target ‘anchor’ molecule for in situ click chemistry.
Screening for enzyme-templated reaction
The in situ click chemistry experiments were carried out in parallel in 96-well microtiter plates to explore the chitinase-accelerated reaction using a mixture of SmChi A, B and C1 (Sigma Co., Tokyo, Japan). Utilization of the mixed SmChi has the advantage of accelerating the identification of new inhibitors against each isozyme of chitinase through a one-off screening. Although a single isozyme or multiple isozymes of chitinase may participate in the formation of triazoles under this particular screening condition, the identification of the actual templating isozyme or isozymes can be determined in a follow-up assay using separate isozymes. Consequently, azide 5 (100 μM) and 71 structurally diverse alkynes (300 μM) were incubated in the presence of SmChi A, B and C1 (9.6 mUnit ml−1) in 10% methanol containing phosphate buffer solution at pH 7.0 (Scheme 1). The formation of the triazole products was monitored by HPLC and mass spectrometry by selected ion recording detection (LCMS-SIR, also known as HPLC and mass spectrometry in selected ion monitoring (LCMS-SIM)) after 20 h at 37 °C. After analysis of each reaction mixture, only alkyne 6 (IC50>30 μM) was sufficiently accelerated in its cycloaddition with azide 5 in the presence of the enzymes to yield a detectable amount of triazole 7 (at this point not distinguished as to whether a syn- or anti-substituted triazole) in the background with great reproducibility by LCMS-SIR measurement (Figure 3). In effect, the chitinases had performed as a reaction vessel of molecular scale to create its own better inhibitor.
Results of in situ click chemistry between 5 and 6, monitored by LCMS-SIR. (a) Authentic sample of 7 from thermal reaction (100 °C, 12 h), apparently single peak (4.9 min) of 7 (anti: syn=3: 2) was observed; (b) reaction (37 °C, 20 h) between 5 (100 μM) and 6 (300 μM) in the presence of SmChi (9.6 mUnit ml−1); (c) without SmChi (background reaction).
Preparations and inhibitory activities of triazole 7
Subsequently, azide 5 and alkyne 6 were subjected to copper(I)-catalyzed azide–alkyne cycloaddition conditions (CuAAC)36, 37, 38 along with the ruthenium-catalyzed azide-alkyne cycloaddition reaction conditions (RuAAC),39, 40 to prepare pure regioisomers of 7 allowing the identification of the regiochemistry of the triazole formed by the enzymes. As expected, pure 1,4- and 1,5-disubstituted triazole products (anti -7 and syn -7) were obtained (Scheme 2). Having both pure triazoles in hand, we turned our attention to the identification of the generated triazole analog by TGS and the participating isozymes of the enzyme for this in situ click chemistry. For the determination of IC50 values against each SmChi isozyme, anti- and syn -7 were assessed in a competition assay with 4-MU-(GluNAc)2. As shown in Figure 4, the inhibitory activities of both regioisomers of 7 against SmChi A and C1 were almost the same as those of 5. On the other hand, syn -7 displayed high inhibitory activity against SmChi B (IC50 value of 0.022 μM), which is approximately 30-fold stronger than that of 5 (approximately 300-fold improved potency as compared with the natural product 1). These results strongly indicate that syn -7 is most likely formed in situ by the SmChi B isozyme in the enzyme mixture.
Acceleration effect of syn-triazole formation through SmChiB
Analysis of syn-anti selection for the in situ screening by LCMS-SIR revealed that a combination of azide 5 and alkyne 6 had led to the accelerated formation of syn -7 in the presence of pure (His)6-SmChiB in an enzyme-dose-dependent manner (Figure 5). Moreover, no syn-triazole formation was observed in the control incubation containing SmChi B and the same azide and alkyne in the presence of a natural product argadin acting as a potent inhibitor (see Supplementary Information for structure)41 (IC50 values against SmChiB, 33±2.8 nM), thereby validating syn -7 as an in situ hit and confirming that its formation required the enzyme active site to be accessible. Interestingly, the regioisomer anti -7, which is predominantly formed under pure thermal Huisgen conditions, is less active against SmChiB than the ‘anchor’ molecule 5, which probably presents the –CH2–N3 group in a unique position when 5 and the protein form their complex. At this point, the chitinase–5 complex is most likely a single entity presenting the azide properly to the ‘well-suited’ alkyne ligand and correctly binding to the complex so that a syn-triazole selectively clicks into its existence. Now the two small molecules have become one, and this enzyme-templated molecule invariably binds more strongly than either component alone. The high kinetic barrier would effectively lower 5 and 6 until they are attracted to each other to form the thermodynamically unfavorable syn-isomer.
Identification of syn-anti selection for in situ click chemistry between 5 (100 μM) and 6 (300 μM), monitored by LCMS-SIR under different conditions from Figure 3. (a) Authentic sample of 7 from thermal reaction (100 °C, 12 h): syn -7 (9.0 min) and anti -7 (9.5 min) (2:3 ratio); (b) reaction with pure (His)6-SmChiB (192 μgml−1; 37 °C, 20 h); (c) (His)6-SmChiB (96 μgml−1; 37 °C, 20 h); (d) (His)6-SmChiB (48 μgml−1; 37 °C, 20 h); (e) (His)6-SmChiB (192 μgml−1), argadin (100 μM; 37 °C, 20 h); and (f) without enzyme (background reaction).
Conclusion and remarks
We discovered a highly active inhibitor of chitinase through in situ click chemistry. The strategy used an azide substituent appended to an active domain excised, as it were, from the more complex natural macrocyclic peptide 1. The SmChi, which in this case was specifically SmChiB, acts as both mold and template for triazole formation between a unique pair of azide and alkyne fragments. Indeed, a number of analogs, based on bioactive molecules, would have to be synthesized to reveal the SAR for the affinity of a specific target molecule (e.g. enzymes) and to identify superior materials for traditional lead discovery. In the process of in situ click chemistry, the highly exergonic nature of triazole formation makes the process completely irreversible and thereby locks in unique information, a kind of embedded message of the encounter. More practically, it allowed us to discover a lead template for the discovery of a selective chitinase inhibitor directed toward the functions of SmChi, without the need for long and costly analog syntheses. Ongoing studies are focused on optimizing the structure and further evaluating bioactivities against various types of chitinases, as well as promoting efforts to seek entirely new inhibitors, not only for chitinases but also for thorough application and refinement of the basic in situ methodology.
Methods
Details of the experimental procedures and characterization of the new compounds, list of the acetylenes library, the expression of each Serratia marcescens chitinases and IC50 determination can be found in the Supplementary Information.
In situ click chemistry experiment using a mixture of SmChiA, B and C1
Azide 5 (5 μl; 2.0 mM in MeOH) was diluted in 0.1 M phosphate buffer (pH 7.0) (60 μl). Subsequently, the required alkynes (5 μl; 6.0 mM in MeOH) were added to the azide solution, followed immediately by the addition of a solution of SmChi A, B and C1, which was purchased from Sigma Co. (C-7809-1UN) (for the ratio of the SmChi isozymes, see Supplementary Figure 10 in the Supplementary Information), in 0.1 M phosphate buffer (30 μl; 32 mUnit ml−1) to give final concentrations of 9.6 mUnit ml−1 SmChi, 100 μM azide 5 and 300 μM of each alkyne (Supplementary Figure 7) in 10% MeOH/phosphate buffer (100 μl). After mixing thoroughly, the reaction mixtures were incubated at 37 °C for 20 h, then diluted with 100 μl MeOH to restrain the enzyme activity and injected directly into the LC/UV-MS instrument to perform LCMS-SIR analysis (Column: Senshu Pak Pegasil (Senshu Scientific Co., Tokyo, Japan) ODS 2ø × 50 mm; conditions for HPLC: gradient 10% MeCN (0.05% TFA)/H2O (0.1% TFA) to 100% MeCN (0.05% TFA) over 8 min, flow 0.3 ml min−1, detect 200–400 nm, temp 20 °C; MS-SIR: cone voltage 60 V, source temp 110 °C, desolvation temp 350 °C, selected mass 704 [M+H]+). The triazole products were identified by their retention times and molecular weights. Control experiments without chitinases (azide 5 (5 μl; 2.0 mM in MeOH) and alkynes (5 μl; 6.0 mM in MeOH) in 0.1 M phosphate buffer (pH 7.0) (90 μl) at 37 °C for 20 h) were run consecutively.
One unit of chitinases can liberate 1.0 mg of N-acetyl-D-glucosamine from chitin per hour at pH 6.0 at in a 2-h assay.
Expression and purification of (His)6-SmChiB
SmChiB gene was amplified by PCR and cloned in frame to pTrcHis B vector (Invitrogen, Tokyo, Japan) to give pTrcHisSmChiB. E. coli strain DH5α harboring plasmid pTrcHisSmChiB was harvested and disrupted by the same procedure used for expression of crude SmChiB. After centrifugation, the supernatant was applied to HisTrap Chelating HP column (Amersham Biosciences, Tokyo, Japan), and the bound proteins were eluted by a stepwise gradient of imidazole in 100 mM phosphate buffer. (His)6-SmChiB was eluted at a concentration of over 150 mM imidazole and showed a molecular mass of 53 kDa on SDS-PAGE (Supplementary Figure 10). The combined fraction was concentrated and imidazole was removed using HiTrap Desalting column (Amersham BioSciences). The protein concentration (320 μg ml−1) was determined by Bradford method using protein quantification kit-rapid (Dojindo, Kumamoto, Japan) with BSA as a standard protein (0.80 absorbance at 595 nm for diluted protein solution ( × 10)) (Supplementary Figure 10). The chitinolytic activity of purified (His)6-SmChiB was assayed by the procedure in the Supplementary Information (Supplementary Figure 10).
In situ formation of syn-7 by SmChiB template reaction
A chromatographic and analytical method different from in situ click screening was used for determining the syn-anti selection of the in situ click chemistry product 7 by (His)6-SmChiB. The assignment was accomplished by comparing the retention times of the in situ products with authentic samples prepared by the thermal reaction of triazole formation.
Azide 5 (5 μl; 2.0 mM in MeOH) was diluted in 0.1 M phosphate buffer (pH 7.0) (60 μl). Subsequently, the alkyne 6 (5 μl; 6.0 mM in MeOH) was added to the azide solution, followed immediately by the additon of a solution of (His)6-SmChiB (320 μg ml−1) in 0.1 M phosphate buffer (30 μl) to give a final concentration of 96 μg ml−1(His)6-SmChiB, 100 μM azide 5 and 300 μM alkyne 6 in 10% MeOH/phosphate buffer (100 μl). After mixing thoroughly, the reaction mixtures were kept at 37 °C for 20 h, then cooled to 20 °C and diluted with 100 μl MeCN to restrain the enzyme activity and injected directly into the LC/UV-MS instrument to perform LCMS-SIR analysis (Column: Develosil (Nomura Chemical Co., Aichi, Japan) C30-UG-5, 2ø × 150 mm; conditions for HPLC: isocratic 29% MeCN(0.05% TFA)/H2O(0.1%TFA), flow 0.3 ml min−1, detect 200–400 nm, temp 20 °C; MS-SIR: cone voltage 95 V, source temp 110 °C, desolvation temp 350 °C, selected mass 704 [M+H]+). The triazole products were identified by their retention times and molecular weights. In concurrence, experiments to investigate the dose dependency of the enzyme for production of syn -7 were carried out under the same condition by using different concentrations of (His)6-SmChiB (final concentrations 192 and 48 μg ml−1 (His)6-SmChiB, respectively, 100 μM azide 5 and 300 μM of alkyne 6 in 10% MeOH/phosphate buffer). From these comparisons, dose-dependent production of syn -7 was clearly observed (Figure 5). LCMS-SIR peaks of syn- and anti -7 were confirmed by co-injection of authentic sample 7 (from thermal reaction) and a crude mixture of enzyme template reaction ((His)6-SmChiB 192 μg ml−1, 100 μM azide and 300 μM alkyne at 37 °C, 20 h) (Supplementary Figure 11). Control experiments in the presence of the known SmChiB inhibitor Argadin41 (IC50 33±2.8 nM) (final concentration: 192 μg ml−1 SmChiB, 100 μM argadin, 100 μM azide 5 and 300 μM alkyne 6) and in the absence of enzyme (100 μM azide 5 and 300 μM alkyne 6) were run consecutively.
References
Boot, R. G. et al. Identification of a novel acidic mammalian chitinase distinct from chitotriosidase. J. Biol. Chem. 276, 6770–6778 (2001).
Shahabuddin, M., Toyoshima, T., Aikawa, M. & Kaslow, D. C. Transmission-blocking activity of a chitinase inhibitor and activation of malarial parasite chitinase by mosquito protease. Proc. Natl Acad. Sci. USA 90, 4266–4270 (1993).
Shibata, Y., Foster, L. A., Bradfield, J. F. & Myrvik, Q. N. Oral administration of chitin down-regulates serum IgE levels and lung eosinophilia in the allergic mouse. J. Immunol. 164, 1314–1321 (2000).
Andersen, O. A., Dixon, M. J., Eggleston, I. M. & van Aalten, D. M. F. Natural product family 18 chitinase inhibitors. Nat. Prod. Rep. 22, 563–579 (2005).
Zhu, Z. et al. Acidic mammalian chitinase in asthmatic Th2 inflammation and IL-13 pathway activation. Science 304, 1678–1682 (2004).
Ōmura, S. et al. Argifin, a new chitinase inhibitor, produced by Gliodadium sp. FTD-0668. I. Taxonomy, fermentation, and biological activities. J. Antibiot. 53, 603–608 (2000).
Arai, N., Shiomi, K., Iwai, Y. & Ōmura, S. Argifin, a new chitinase inhibitor, produced by Gliodadium sp. FTD-0668. II. Isolation, physico-chemical properties, and structure elucidation. J. Antibiot. 53, 609–614 (2000).
Shiomi, K. et al. Structure of argifin, a new chitinase inhibitor produced by Gliocladium sp. Tetrahedron Lett. 41, 2141–2143 (2000).
Suzuki, K. et al. Chitinases A, B, and C1 of Serratia marcescens 2170 produced by recombinant Escherichia coli: enzymatic properties and synergism on chitin degradation. Biosci. Biotechnol. Biochem. 66, 1075–1083 (2002).
Horn, S. J. et al. Comparative studies of chitinases A, B and C from Serratia marcescens. Biocatal. Biotransform. 24, 39–53 (2006).
Rao, F. V. et al. Specificity and affinity of natural product cyclopentapeptide inhibitors against A. fumigatus, human, and bacterial chitinases. Chem. Biol. 12, 65–76 (2005).
Houston, D. R. et al. High-resolution structures of a chitinase complexed with natural product cyclopentapeptide inhibitors: mimicry of carbohydrate substrate. Proc. Natl Acad. Sci. USA 99, 9127–9132 (2002).
Andersen, O. A. et al. Structure-based dissection of the natural product cyclopentapeptide chitinase inhibitor argifin. Chem. Biol. 15, 295–301 (2008).
Rideout, D. Self-assembling cytotoxins. Science 233, 561–563 (1986).
Rideout, D., Calogeropoulu, T., Jaworski, J. & McCarthy, M. Synergism through direct covalent bonding between agents: a strategy for rational design of chemotherapeutic combinations. Biopolymers 29, 247–262 (1990).
Inglese, J. & Benkovic, S. J. Multisubstrate adduct inhibitors of glycinamide ribonucleotide transformylase: synthetic and enzyme-assembled. Tetrahedron 47, 2351–2364 (1991).
Boger, D. L. et al. 10-formyl-5,8,10-trideazafolic acid (10-formyl-TDAF): a potent inhibitor of glycinamide ribonucleotide transformylase. Bioorg. Med. Chem. 5, 1817–1830 (1997).
Maly, D. J., Choong, I. C. & Ellman, J. A. Combinatorial target-guided ligand assembly: identification of potent subtype-selective c-Src inhibitors. Proc. Natl Acad. Sci. USA 97, 2419–2424 (2000).
Nicolaou, K. C. et al. Target-accelerated combinatorial synthesis and discovery of highly potent antibiotics effective against vancomycin-resistant bacteria. Angew. Chem. Int. Ed. 39, 3823–3828 (2000).
Greasley, S. E. et al. Unexpected formation of an epoxide-derived multisubstrate adduct inhibitor on the active site of GAR transformylase. Biochemistry 40, 13538–13547 (2001).
Nguyen, R. & Huc, I. Using an enzyme's active site to template inhibitors. Angew. Chem. Int. Ed. 40, 1774–1776 (2001).
Nicolaou, K. C. et al. Synthesis and biological evaluation of vancomycin dimers with potent activity against vancomycin-resistant bacteria: target-accelerated combinatorial synthesis. Chem. Eur. J. 7, 3824–3843 (2001).
Kehoe, J. W. et al. Tyrosylprotein sulfotransferase inhibitors generated by combinatorial target-guided ligand assembly. Bioorg. Med. Chem. Lett. 12, 329–332 (2002).
Poulin-Kerstien, A. T. & Dervan, P. B. DNA-templated dimerization of hairpin polyamides. J. Am. Chem. Soc. 125, 15811–15821 (2003).
Hu, X., Sun, J., Wang, H.-G. & Manetsch, R. Bcl-XL-templated assembly of its own protein-protein interaction modulator from fragments decorated with thio acids and sulfonyl azides. J. Am. Chem. Soc. 130, 13820–13821 (2008).
Huisgen, R. in 1,3-Dipolar Cycloaddition Chemistry, Vol. 1 (ed. Padwa, A.) 1–176 (Wiley, New York, 1984).
Sharpless, K. B. & Manetsch, R. In situ click chemistry: a powerful means for lead discovery. Expert Opin. Drug Discov. 1, 525–538 (2006).
Lewis, W. G. et al. Click chemistry in situ: acetylcholinesterase as a reaction vessel for the selective assembly of a femtomolar inhibitor from an array of building blocks. Angew. Chem. Int. Ed. 41, 1053–1057 (2002).
Manetsch, R. et al. In situ click chemistry: enzyme inhibitors made to their own specifications. J. Am. Chem. Soc. 126, 12809–12818 (2004).
Bourne, Y. et al. Freeze-frame inhibitor captures acetylcholinesterase in a unique conformation. Proc. Natl Acad. Sci. USA 101, 1449–1454 (2004).
Krasiéski, A. et al. In situ selection of lead compounds by click chemistry: target-guided optimization of acetylcholinesterase inhibitors. J. Am. Chem. Soc. 127, 6686–6692 (2005).
Mocharla, V. P. et al. In situ click chemistry: enzyme-generated inhibitors of carbonic anhydrase II. Angew. Chem. Int. Ed. 44, 116–120 (2005).
Whiting, M. et al. Inhibitors of HIV-1 protease by using in situ click chemistry. Angew. Chem. Int. Ed. 45, 1435–1439 (2006).
Brurberg, M. B. et al. Chitinase B from Serratia marcescens BJL200 is exported to the periplasm without processing. Microbiology 141, 123–131 (1995).
Hodge, A., Gooday, G. W. & Alexander, I. J. Inhibition of chitinolytic activities from tree species and associated fungi. Phytochemistry 41, 77–84 (1996).
Rostovtsev, V. V., Green, L. G., Fokin, V. V. & Sharpless, K. B. A stepwise Huisgen cycloaddition process: copper(I)-catalyzed regioselective ligation of azides and terminal alkynes. Angew. Chem. Int. Ed. 41, 2596–2599 (2002).
Tornøe, C. W., Christensen, C. & Meldal, M. Peptidotriazoles on solid phase: [1,2,3]-triazoles by regiospecific copper(I)-catalyzed 1,3-dipolar cycloadditions of terminal alkynes to azides. J. Org. Chem. 67, 3057–3064 (2002).
Chan, T. R., Hilgraf, R., Sharpless, K. B. & Fokin, V. V. Polytriazoles as copper(I)-stabilizing ligands in catalysis. Org. Lett. 6, 2853–2885 (2004).
Zhang, L. et al. Ruthenium-catalyzed cycloaddition of alkynes and organic azides. J. Am. Chem. Soc. 127, 15998–15999 (2005).
Boren, B. C. et al. Ruthenium-catalyzed azide-alkyne cycloaddition: scope and mechanism. J. Am. Chem. Soc. 130, 8923–8930 (2008).
Arai, N. et al. Argadin, a new chitinase inhibitor, produced by Clonostachys sp. FO-7314. Chem. Pharm. Bull. 48, 1442–1446 (2000).
Acknowledgements
This work was supported by the Grant of the 21st Century COE Program, Ministry of Education Culture, Sports, Science and Technology. We also thank Ms A Nakagawa, Ms N Sato and Dr K Nagai for various instrumental analyses.
Author information
Authors and Affiliations
Corresponding authors
Additional information
Supplementary Information accompanies the paper on The Journal of Antibiotics website (http://www.nature.com/ja)
Supplementary information
Rights and permissions
About this article
Cite this article
Hirose, T., Sunazuka, T., Sugawara, A. et al. Chitinase inhibitors: extraction of the active framework from natural argifin and use of in situ click chemistry. J Antibiot 62, 277–282 (2009). https://doi.org/10.1038/ja.2009.28
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ja.2009.28
Keywords
This article is cited by
-
Marine chitinolytic enzymes, a biotechnological treasure hidden in the ocean?
Applied Microbiology and Biotechnology (2018)
-
In situ click chemistry generation of cyclooxygenase-2 inhibitors
Nature Communications (2017)
-
A new application of click chemistry in situ: development of fluorescent probe for specific G-quadruplex topology
Scientific Reports (2015)
-
Catalytic site inhibition of insulin-degrading enzyme by a small molecule induces glucose intolerance in mice
Nature Communications (2015)
-
Solid-phase total synthesis of the chitinase inhibitor Argadin using a supported acetal resin
The Journal of Antibiotics (2009)