Abstract
A specificity of Brassicaceous plants is the production of sulphur secondary metabolites called glucosinolates that can be hydrolysed into glucose and biocidal products. Among them, isothiocyanates are toxic to a wide range of microorganisms and particularly soil-borne pathogens. The aim of this study was to investigate the role of glucosinolates and their breakdown products as a factor of selection on rhizosphere microbial community associated with living Brassicaceae. We used a DNA-stable isotope probing approach to focus on the active microbial populations involved in root exudates degradation in rhizosphere. A transgenic Arabidopsis thaliana line producing an exogenous glucosinolate and the associated wild-type plant associated were grown under an enriched 13CO2 atmosphere in natural soil. DNA from the rhizospheric soil was separated by density gradient centrifugation. Bacterial (Alphaproteobacteria, Betaproteobacteria, Gammaproteobacteria and Acidobacteria), Archaea and fungal community structures were analysed by DGGE fingerprints of amplified 16S and 18S rRNA gene sequences. Specific populations were characterized by sequencing DGGE fragments. Roots of the transgenic plant line presented an altered profile of glucosinolates and other minor additional modifications. These modifications significantly influenced microbial community on roots and active populations in the rhizosphere. Alphaproteobacteria, particularly Rhizobiaceae, and fungal communities were mainly impacted by these Brassicaceous metabolites, in both structure and composition. Our results showed that even a minor modification in plant root could have important repercussions for soil microbial communities.
Similar content being viewed by others
Introduction
Root exudates are critical in determining the nature of plant–microorganisms interaction in the rhizosphere and shape microbial community structure present in the vicinity of the roots (Butler et al., 2003; Bais et al., 2006; Costa et al., 2006; Paterson et al., 2007; Broeckling et al., 2008; Haichar et al., 2008). Some studies have focused on the implication of a specific plant defense-signaling pathway as a driving force in structuring native soil microbial communities. Salicylic and jasmonic acids signal molecules reduce natural endophytic and epiphytic bacterial diversity on leaves of Arabidopsis thaliana (Kniskern et al., 2007). Systemic acquired resistance can alter the rhizosphere bacterial communities (Hein et al., 2008). The glucosinolate–myrosinase chemical defense system is specific of Brassicaceae, Capparaceae, Caricaceae, Resedaceae and Moringaceae plants (Fahey et al., 2001). This system influences the active microbial community in the rhizosphere of canola (Rumberger and Marschner, 2003) but its real impact on the native microbial community in the rhizosphere of Brassicaceae remains unknown.
Glucosinolates are specific secondary metabolites stored in plant cell vacuoles. When plant tissues are damaged, glucosinolates come into contact with the enzyme myrosinase located in the cell wall, cytoplasm or in separate cells and are hydrolysed into glucose, sulphates and biocidal products such as isothiocyanates, nitriles and ionic thiocyanates (Grubb and Abel, 2006; Halkier and Gershenzon, 2006). This defense system is active against aerial herbivores, pests or pathogens (Halkier and Gershenzon, 2006). The incorporation and degradation of Brassicaceous tissues in soil releasing active hydrolysis products is used to tentatively control soil-borne pests (Angus et al., 1994; Brown and Morra, 1997). The effects on fungal and bacterial responses of the glucosinolate-myrosinase defense system have already been described. However, these studies implicated pure strains of pathogen organisms and in vitro approaches (Brabban and Edwards, 1995; Kirkegaard et al., 1996; Smith and Kirkegaard, 2002; Souza-Fagundes et al., 2004). The incorporation of crucifer tissues showed a variety of effects on non-target soil microbial populations in studies based on isolation and culture of microorganisms (Scott and Knudsen, 1999; Bending and Lincoln, 2000; Cohen et al., 2005). However, such approaches do not reflect the actual microbial community as 0.1 to 10% of total population in soil can be cultured according to current methods (Keller and Zengler, 2004).
In this study, we focused on the influence of glucosinolates and hydrolysis products on native total microbial populations, through the production of a glucosinolate compound released in situ from living Brassicaceae roots in natural soil.
We used an original culture-independent stable isotope probing (SIP) approach to describe plant microorganisms fine scale interactions in the rhizosphere. This culture-independent approach allows discriminating root exudate-assimilating microbial populations in plant rhizosphere from those dormant or degrading soil organic matter (Rangel-Castro et al., 2005; Haichar et al., 2008) and to link the identity of microorganisms to their function in soil (Radajewski et al., 2000; Lu et al., 2006).
We chose a transgenic A. thaliana plant in which CYP79A1 gene from sorghum (Sorghum bicolor) was introduced, which led to the accumulation of up to 3% dry matter of p-hydroxybenzylglucosinolate, an aliphatic glucosinolate derived from tyrosine (Bak et al., 1999; Kristensen et al., 2005). We report that the aliphatic hydroxybenzylglucosinolate is only produced in transgenic A. thaliana plant roots and that the modifications of glucosinolate profiles by introduction of CYP79A1 gene leads to specific changes in the active microbial community on the roots but also in the rhizosphere of A. thaliana growing in natural soil.
Materials and methods
Plant growth and 13C labelling
Glucosinolates biosynthesis pathways are linked to metabolic pathways of other compounds, like the plant hormone indole-3-acetic acid (Halkier and Gershenzon, 2006) and perturbation of glucosinolate metabolism in plant mutants could cause severe phenotypes (Yan and Chen, 2007). The A. thaliana transgenic line CYP79A1 (Bak et al., 1999) was selected for its glucosinolate unusual profile and for its similar phenotype than wild-type plant. It results from the introduction of the CYP79A1 gene from sorghum which led to the production of high levels of the tyrosine-derived glucosinolate p-hydroxybenzylglucosinolate not known to accumulate in wild-type A. thaliana ecotype Columbia (Bak et al., 1999). Transgenic CYP79A1 plant seeds were provided by Halkier BA (Royal Veterinary and Agricultural University, Copenhagen, Denmark) and together with wild-type Col, were grown in triplicate in soil under 13CO2 following the protocol described by Haichar et al. (2008), to compare their rhizospheric microbial communities. Briefly, after seed sterilization, plants were grown in pots for 18 days before being continuously labelled for 25 days by injection of pure (>99% atom 13C) 13CO2 (Purchased from Cortec Net, Paris, France) monitored to maintain an isotope excess at >80% atom 13C (Figure 1). Replicates of transgenic CYP79A1 and Columbia plants were grown in a twin growth chamber under the same conditions except unlabelled CO2 for the analysis of glucosinolate profiles in plants. Triplicate of soil pots without plants were also incubated in the same conditions.
Harvesting procedure and DNA extraction
Roots of each plant replicate were separated from rhizospheric soil, washed with sterile water and both rhizospheric soil and roots were frozen in liquid nitrogen and stored at −80 °C. Total nucleic acids were extracted from 3 × 1.5 g of soil per plant sample and from entire root systems as described by Ranjard et al. (2003). Integrity of total DNA was verified on agarose gel and DNA concentrations were assessed using PicoGreen kit staining (Molecular Probes, Paris, France) following the manufacturer's protocol. DNA solutions were then stored at −20 °C. The 13C enrichment of total DNA extracted from rhizosphere was checked by a measure of δ13C (‰) by isotope ratio mass spectrometry coupled with an elemental analyser (IRMS, Delta+ and Conflo, Thermofinnigan, Thermo-electron corp, Bremen, Germany). One to 2 μg of each DNA solution were placed into 5 × 9 mm tin capsule (Elemental Microanalysis Limited, UK) and submitted to mass spectrometry. δ13C (‰) was determined using the equation:
δ13C (‰)=[(Rsample−Rstandard)−1] × 1000
where R=13C/12C. The Rstandard was Pee Dee Belemite (Wang and Hsieh, 2002).
Density gradient separation and fractionation
CsCl density gradient centrifugation was performed to separate 13C-labelled and unlabelled nucleic acids extracted from rhizospheric and unplanted soil samples as described by Haichar et al. (2007). Approximately 5 μg of DNA were loaded in the density gradient and centrifuged in 4.9 ml polyallomer Optiseal tube in an NVT 90 rotor (Beckman) at 20 °C, 45 000 r.p.m., for 72 h. Centrifuged gradients were fractionated from the top to the bottom into approximately equal 200 μl by using 1 ml syringes fitted with 21-G needles. Density of each fraction was determined by weighing. DNA concentrations were fluorometrically quantified using PicoGreen staining kit. For each gradient, one fraction representative of 13C-labelled (‘heavy’) DNA and one fraction representative of unlabelled (‘light’) DNA were chosen according to a standardized method based on previous studies in the laboratory (Haichar et al., 2007, 2008) following buoyant density and DNA content of each fraction in comparison with control gradient containing unlabelled soil DNA and 13C labelled DNA from Escherichia coli. The heavy fraction was considered to contain DNA representative of populations involved directly or not, in root exudates assimilation and the light one, to contain DNA from populations degrading soil organic matter or inactive (Figure 1) (Rangel-Castro et al., 2005; Haichar et al., 2008).
Microbial community analysis
Nucleic acids were purified from CsCl salts using GeneClean kit (MP Biomedical). PCR amplifications were performed from root DNA and from light and heavy fractions obtained from the rhizospheric soil of each plant. Alphaproteobacteria, Betaproteobacteria, Gammaproteobacteria, Acidobacteria, Archaea and fungal communities were targeted by amplification of 16S or 18S rRNA gene fragments with adapted primers listed in Table 1, using a nested PCR approach (Muyzer et al., 1993) according to the authors recommendations.
PCR products were analyzed by denaturing gradient gel electrophoresis (DGGE) allowing identification of specific populations through band excisions. DGEE was performed with the Dcode Universal Mutation Detection System (BIO-Rad Laboratories, France) on polyacrylamide gels with a denaturant gradient between 28% and 58% (100% denaturant equals 7 M urea and 40% (v/v) formamide). Aliquots of PCR products were loaded on the gel and electrophoresis was carried out with 1 × Tris-acetate-EDTA buffer at 60 °C and at 75 V for 17 h. Gels were silver-stained and scanned. ImageQuant TL one-dimensional gel analysis software (V2003, Amersham Biosciences, France) was used to determine band presence and intensity as described by McCaig et al. (2001). Intensities were normalized to avoid background bias and the DGGE matrices obtained were analysed using principal component analysis (PCA) ADE-4 software (Thioulouse et al., 1997). Statistical ellipses representing 90% confidence on PCA plots were used to compare DGGE profiles. If two ellipses representing two different treatments do not overlap, the treatments have significant different DGGE profiles with an alpha risk of 10%.
After electrophoresis the denaturing gels were stained with a 1:10 000 dilution of SYBR Green I nucleic acid stain (Sigma-Aldrich, USA) and prominent bands were excised from DGGE gels under UV. DNA was eluted in sterile water at 65 °C for 30 min and used as a template for PCR amplification under the conditions mentioned above. In most cases as reported by Mahmood et al. (2005), further rounds of band excision, PCR amplification and DGGE analysis were necessary before sequencing the PCR products (Genome express, Meylan, France). A particular attention was paid on 13C-labelled populations and those impacted by glucosinolates, specifically in case of minor DGGE bands. The NCBI BLASTN search tool (Altschul et al., 1990) was used to determine the most similar sequences in GenBank database.
Analysis of glucosinolates and derivatives
The roots of six replicates of both Col and CYP79A1 plants grown with unlabelled CO2 were freeze dried before glucosinolate extractions to avoid activation of ‘glucosinolate-myrosinase’ system and glucosinolate hydrolysis. Glucosinolates were extracted by 70% v/v methanol following Mellon et al. (2002) and Wathelet et al. (2004). Analysis of glucosinolates was then carried out using a HPLC/DAD/ESI-MS (HP 1100 series, Agilent Technologies) with a C-18 Nucleodur sphinx reversed-phase column (250 × 4.6 mm, 5 μm, Macherey Nagel). The system was managed by Chemstation agilent software (Agilent Technologies). Mobile phase was a linear gradient of water with 0.1% v/v trifluoroacetic acid (TFA) (A) and methanol with 0.1% v/v TFA (B) at a flow rate of 1 ml min−1. The linear gradient was 0 to 5 min 0% solvent B; 5 to 25 min 0% to 22% solvent B; 25 to 40 min 22% to 50% solvent B; 40 to 45 min 50% to 100% solvent B. Then, for 10 min the column was washed and equilibrated before the next injection. Spectra were recorded between 200 to 600 nm and the response at 228 nm was used for quantification. Mass spectrometry operating conditions were: gas temperature 340 °C at a flow rate of 11 l min−1, nebulizer pressure 30 p.s.i, quadripole temperature 30 °C, capillary voltage 4000 V and fragmentor 150. Full scan spectra from m/z 60 to 900 in both positive and negative ion mode were obtained. Identification was based on the molecular weight and previously published information.
Isothiocyanates were tentatively extracted from rhizospheric soil on four replicates of each plant type Col and CYP79A1. Plants were grown in a growth chamber as previously described with some modifications to limit volatilization of isothiocyanates. After 4 weeks, rhizospheric soil of each replicate was carefully collected. Isothiocyanates were immediately extracted with ethyl acetate following a slightly modified procedure of Gimsing and Kirkegaard (2006). They were separated by GC/MS (GC 6890; mass detector HP 5973; Agilent Technologies) using a column HP—INNOWAX (30 m × 0.25 mm with a 0.25 μm film; Agilent Technologies). Helium was used as the carried gaz at 1 ml min−1. One μl of sample was injected in splitless mode at 230 °C. The temperature program was set at 50 °C for 1 min, 50–105 °C at 3 °C min−1, 105–200 °C at 8 °C min−1, 200–240 °C at 30 °C min−1 and 240 °C for 2 min. Mass detection of the samples was performed at 270 °C.
Results
Glucosinolates and their hydrolysis product analysis
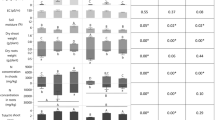
The glucosinolate profiles of CYP79A1 transgenic A. thaliana plants were compared with the profiles of wild-type plants. We focused on root compartment, as studies on glucosinolate content of CYP79A1 and wild-type plants concerned leaves (Bak et al., 1999; Kristensen et al., 2005). Glucosinolates content of six replicates of seedling root tissues at post-germination stage reveal a different profile for CYP79A1 plants. Besides the exclusive production of p-hydroxybenzylglucosinolate by CYP79A1 plants (Figure 2, peak 2), another minor unidentified compound is detected only in CYP79A1 plants (Figure 2, peak 5), and two compounds show significant reduced amplitude in these plants (Figure 2, peaks 1 and 6). Those variations are in agreement with previous results obtained with CYP79A1 leaves (Bak et al., 1999; Kristensen et al., 2005).
Glucosinolates extracted from A. thaliana roots. (a) Representative HPLC chromatogram obtained for CYP79A1 (A1) plant. Stars indicate peaks not detected in root sample of Col plant. (b) Representative HPLC chromatogram for Col plant. (c) Amplitude of each major glucosinolate peak detected by HPLC analyses per gram of Col and A1 plant roots. Means values of six replicates with standard deviations. Stars indicate significant differences in peak amplitude between the two plant types (P=0.05).
In the rhizosphere soil, isothiocyanates could not be successfully detected because either very low amounts of isothiocyanates were released by plant roots or isothiocyanates turnover was so rapid in the rhizosphere that they were undetectable.
Influence of glucosinolates and derivatives on the structure of root microbial communities
We separated roots and rhizospheric soil because plant root tissues are the source of glucosinolates and derivatives, and the highest effect on soil microflora was expected in this fraction (Figure 1). Actually, PCA analyses show significant differences between Col and CYP79A1 plants for Alphaproteobacteria, Acidobacteria and fungal root populations (Figure 3). DGGE bands excised were sequenced and affiliated to the closest relatives to identify impacted populations.
Denaturing gradient gel electrophoresis fingerprint of ribosomal gene fragments from root samples. (A) Fungi, (B) Alphaproteobacteria, (C) Acidobacteria and their corresponding principal component analyses for (a) fungi, (b) Alphaproteobacteria, (c) Acidobacteria. Three replicates for each plant type, wild-type Col and transgenic CYP79A1 (A1). Statistical ellipses drawn over the plot replicates represent 90% confidence. Arrowheads indicate examples of DGGE bands specific of one plant type. Legends:  Col plants;
Col plants;  A1 plants. Band numbers are reported in Table 2.
A1 plants. Band numbers are reported in Table 2.
One Acidobacteria-specific population, closely related to an uncultured clone from a soil sample of radish rich area, is only present in wild type Col plant (Table 2). Alphaproteobacteria populations associated with plant roots are all close to members of Rhizobiales, underlining the selective effect of the roots on this group (Table 2, Figure 3a). In the Agrobacterium/Rhizobium genera, two populations are only associated with CYP79A1 roots and two other populations are located on Col roots (Table 2).
Fungal populations associated with plant roots are close to Basidiomycota, Ascomycota and Chytridiomycota (Table 2). One specific fungal population with a very intense band is only present on CYP79A1 plant roots (Figure 3). This band is assigned to Syncephalis depressa (Zoopagomycotina), a fungal species known to be a parasite of other fungi.
No significant differences between Col and CYP79A1 plants are detected by PCA on roots for Betaproteobacteria, Gammaproteobacteria and Archaea communities (Supplementary Figure S1). All identified populations of these taxa are detected to be associated with both plant type roots (Table 2).
Influence of glucosinolates and derivatives on the structure of rhizosphere microbial communities
The application of DNA-SIP technique in the rhizosphere allowed distinction between microbial populations actively assimilating root exudates (fresh carbon source) from dormant and/or soil organic matter degrading microorganisms (assimilating ancient carbon source) (Figure 1). In situ assimilation of root-derived carbon by soil microorganisms and sufficient incubation time are confirmed by the ∂13C mean value of total DNA recovered from rhizosphere of 13C-labelled plants that reach 261 (±64) ‰ compared to ∂13Cvalue of 17 (±13) ‰ obtained for DNA recovered from unplanted soil.
The DGGE patterns of rhizospheric soil-inhabiting microorganisms are different from those of root-colonizing microorganisms as illustrated for Alphaproteobacteria (Figure 4a). Principal component analyses reveal significant differences between light and heavy DNA fractions (Supplementary Figure S2 and S3). The most important difference based on both PC1 and PC2 axes is observed for Alphaproteobacteria and Archaea from transgenic CYP79A1 plant. The ability of some populations to assimilate fresh (13C) or ancient (12C) sources of soil carbon explains most of the differences in the community structure of these groups 87 and 92%, respectively (Supplementary Figure S3). For all other cases, significant effect becomes slighter, only on the PC2 axis explaining less than 18% of the variability. Visual comparison of corresponding DGGE profiles reveals, for Alphaproteobacteria and Betaproteobacteria, replicable apparition or clear enrichment of specific bands from heavy DNA fractions corresponding to exudate degrading populations compared with light DNA fractions (Supplementary Figure S4). DNA-SIP allows pointing out a ‘rhizosphere effect’ of both wild-type Col and transgenic CYP79A1 plant exudates on the structure of the majority of tested taxa. Some specific populations active in the rhizosphere through assimilation of exudates were identified and are presented in Table 3. These populations are not detected in the heavy fractions of unplanted soil showing that 13C enrichment does not come from an autotrophic activity (Table 3).
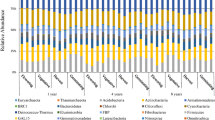
Denaturing gradient gel electrophoresis fingerprints of ribosomal gene fragments from light DNA fraction (L), heavy DNA fraction (H) for wild-type Col plant, transgenic CYP79A1 (A1) plant, three replicates for each treatment, (a) Alphaproteobacteria, 16S rRNA gene fragments from rhizosphere soil and root samples. White left-directing arrowhead indicates a 13C-labelled Rhizobium population present in both plant type rhizosphere but only on A1 plant roots. White right-directing arrowhead indicates a 13C-labelled Rhizobium population present on both plant type rhizosphere and roots. (b) Fungi, 18S rRNA gene fragments from rhizospheric soil. Black arrowhead indicates a population close to Syncephalis depressa, specific of A1 plant rhizosphere. Band numbers are reported in Tables 2 and 3.
Concerning the effect of the exogenous glucosinolate produced by CYP79A1 transgenic plants, we detected significant differences in heavy DNA profiles in rhizospheric soil for Alphaproteobacteria, Gammaproteobacteria and fungal communities (Supplementary Figure S5). Plant genotype-specific heavy DNA bands are detected on DGGE profiles except for Gammaproteobacteria where only variations of band intensities are observed. DGGE profiles obtained with light DNA are similar and no difference was detected.
Alphaproteobacteria are highly influenced in rhizospheric soil by the modification of glucosinolate content. They are more diverse than on roots and affiliated to members of Rhizobiales, Rhodobacterales and Caulobacterales (Table 3). Three populations close to Mesorhizobium, Bosea and Rhizobium are only present in the heavy DNA fraction of CYP79A1 plant rhizosphere. Some other active Rhizobium populations are present in rhizospheric soil for both plant types, clearly implicated in the assimilation of root exudates but independently of glucosinolate profiles (Table 3). Mesorhizobium, frequently isolated from plant nodules (Wang et al., 2002; Weir et al., 2004), had a plant growth promoting effect on Indian mustard Brassica campestris (Chandra et al., 2007) when Rhizobium species known for their symbiotic association with members of the Fabaceae (Leguminosae) or their ability to form tumours with many plants (Hirsch, 2004) were already isolated from rhizosphere in non-symbiotic context (Yanni et al., 1997; Berge et al., 2009). Bosea populations seemed to have active roles in the plant rhizosphere (Martin-Laurent et al., 2006; Dandie et al., 2007) and could grow on specific compounds produced by the root of transgenic Lotus Oger et al. (2004). DGGE profiles obtained for roots and rhizospheric populations show that one 13C-labelled Rhizobium impacted by glucosinolate modifications in roots is also present in rhizosphere but clearly not susceptible to glucosinolate content (Figure 4a).
Fungal community is clearly influenced by glucosinolate profile in rhizospheric soil, with a specific and intense band mainly present in heavy DNA (Figure 4b). As for root compartment, this band is assigned to the fungal parasite S. depressa (Table 3). This population is particularly susceptible to plant glucosinolate content and able to use, directly or indirectly through its host, 13C-labelled root exudates. Other populations close to Basidiomycota, Ascomycota and Chytridiomycota are identified in light and heavy DNA fractions of rhizospheric soil for both plant types (Table 3). These populations are not susceptible to glucosinolate profile modifications in CYP79A1 plant.
No significant difference between Col and CYP79A1 plants are detected for Acidobacteria, Betaproteobacteria and Archaea communities by PCA analyses (Supplementary Figure S5). All identified populations for these taxa are detected in both plant types rhizosphere (Table 3).
However, we identify one 13C-labelled archaeal population implicated in root exudates assimilation, only detected in CYP79A1 plant rhizosphere. This population closely related to an uncultured crenarchaeote from soil (Table 3), does not significantly change global DGGE profile because of high variability for this group. Crenarchaeota populations were considered as the most abundant ammonia-oxidizing organisms in soil ecosystems (Leininger et al., 2006).
Discussion
We measure effective production of the exogenous glucosinolate in root tissues of A. thaliana CYP79A1 transgenic line compared with wild-type plants. Root growth is inevitably accompanied by superficial cell destruction leading to hydrolysis of glucosinolates and release of their degradation products. Living Brassicaceae roots could also release these metabolites through root exudation as shown for Brassica napus (Choesin and Boerner 1991) and mustard roots (Schreiner and Koide 1993) and directly impact microbe living on roots. Populations closely associated with plant roots directly exposed to these compounds but also rhizospheric populations are impacted by this glucosinolate profile change. We hypothesize that glucosinolates and their hydrolysis products diffuse from roots into the rhizosphere where they are diluted and could be degraded by extracellular myrosinase in soil (Borek et al., 1996) as shown in canola rhizosphere (Rumberger and Marschner, 2003). This gradient of concentrations could explain that some Acidobacteria and Rhizobium populations are influenced by the modification of glucosinolate content in the root compartment but not in the rhizosphere. A. thaliana roots are very small and amounts of glucosinolates and derivatives proportional to roots size could be very low which could preclude their detection by GC. In the rhizosphere, these compounds could act as a signal working at very low concentration and having a short lifespan. In this fraction the modification of glucosinolate content in CYP79A1 plant has an impact mainly on exudate consumer 13C-labelled populations. It could indicate that this signal is a part of the plant driving force-selecting populations for its root exudate degradation.
DNA-SIP and continuous labelling of plant allow showing the global impact of glucosinolates and derivatives on active and growing microbial communities in rhizosphere. It could also be interesting to follow the temporal variations of microbial community structure during the plant growth (Petersen et al., 2002; Brown et al., 2003). An RNA-SIP approach combined with pulse labelling could be adequate for detection of punctual changes in active populations at a determined stage during plant culture. RNA is considered as a more sensitive marker for time course experiment because it does not require cell division for isotope incorporation (Whiteley et al., 2006).
Target members of Alphaproteobacteria and fungal community appear to be mostly impacted by the change in glucosinolate profile in root and rhizosphere compartments. Fungi modifications are expected according to literature. They were reported to be more sensitive than bacteria to these compounds (Smith and Kirkegaard, 2002). The formation of antimicrobial products from glucosinolates has been also suggested to explain the inability of Brassicaceae plants to form arbuscular mycorrhizal (Vierheilig et al., 2000; Roberts and Anderson, 2001). As an exception, Thlaspi praecox and Biscutella laevigata form arbuscular mycorrhizal symbiosis but only during the reproductive period that coincided with low level of glucosinolates in roots (Pongrac et al., 2008). However, the development of arbuscular mycorrhizal fungi in the host roots involved a complex cellular relationship both controlled by the plant branching factors strigolactones (Akiyama et al. 2005) and the Common Symbiosis Pathway (CSP) plant-signalling pathway. This pathway is common to arbuscular mycorrhizal, rhizobial and actinorhizal root symbioses representing mutualistic relationships in plant root (Markmann et al., 2008). Strigolactone is found in root exudates of many species including A. thaliana (Goldwasser et al., 2008) but most CSP gene orthologs were not predicted in the A. thaliana genome, whereas a non nodulating mycorrhized plant, O. sativa possesses all CSP ortholog candidates (Zhu et al., 2006). Nothing is known about possible a relationship in A. thaliana between the presence of the glucosinolate pathway and the absence of CSP, two specificities in Brassicaceae plants. Both could have suppressive independent effects on mycorrhyzas (Pieterse and Dicke, 2007) and perhaps on some other fungi.
Changes linked to glucosinolates and derivatives in Alphaproteobacteria community, particularly Rhizobiaceae, were less explored in the literature. Ground root tissue of Brassica was toxic to Bradyrhizobium pure strains (Trinick and Hadobas, 1995) and in vitro colonization of Brassica napus roots by the Alphaproteobacteria Azorhizobium caulinodans was affected by plant glucosinolate content (O'Callaghan et al., 2000). In our study, we show different susceptibilities of spontaneous Rhizobiaceae populations exposed to Brassicaceae roots. This unexpected high reactivity could have a role in the selection of populations that are important to explore because rhizobia are of particular interest in soil, according to possible positive or negative effect on plant health and growth.
Differential effects on microbial populations that we observed could result from variable direct susceptibility of populations towards antimicrobial properties of hydrolysis products of glucosinolates (Smith and Kirkegaard, 2002; Kliebenstein et al., 2005; Brader et al., 2006). Beside their antimicrobial effect, glucosinolates and their hydrolysis products could also stimulate microbial growth. In soil, glucosinolates are hydrolysed by microorganisms (Fahey et al., 2001) and microbial degradation is the main mechanism for disappearance of isothiocyanates (Rumberger and Marschner, 2003). Microorganisms could perceive these metabolites as specific signal compounds stimulating growth but also use them as a nutritional substrate (Zeng et al., 2003; Souza-Fagundes et al., 2004).
These compounds could also act indirectly on populations through nutritional or toxic impact on their hosts, competitors, antagonists or predators. Indeed, balance between populations was strictly governed by interaction among community members, each of them might be affected positively or negatively by each other (Trosvik et al., 2008). It should be the case for the most impacted fungi S. depressa, only present in CYP79A1 transgenic plants, that is a parasite of another fungi. Host sensitivity to glucosinolate composition could impact S. depressa population development.
We could not exclude that CYP79A1 plants are not only modified in glucosinolates profile because of a known link with other pathways, particularly those implicated in plant defense. Kristensen et al. (2005) showed that insertion of CYP79A1 gene and accumulation of the novel glucosinolate had only marginal inadvertent effects on the plant transcriptome and metabolome compared with wild type. However, Brader et al. (2006) showed that the accumulation of the novel glucosinolate could reduce the induction of jasmonate-mediated pathway but led to increased levels of salicylic acid. These plant defense pathways could modify the soil bacterial community structure (Kniskern et al., 2007; Hein et al., 2008).
In conclusion, we show that the production of only one exogenous glucosinolate by CYP79A1 plant alters microbial community structure on roots and rhizosphere. The original defense system ‘glucosinolate-myrosinase’ can be considered as an effective factor of microbial selection in root environment of Brassicaceae where inter-strain competition is strong for substrate assimilation. Even small advantages or disadvantages caused by glucosinolates and their hydrolysis products would select the fittest strains. This factor of selection is probably also effective on other organisms such as those of soil microfauna. Further studies are now necessary to draw conclusions about changes in microbial functions because of possible functional redundancy in soil (Wertz et al., 2006).
An interesting area for further study would be to understand the determinants and molecular mechanisms sustaining the differential response of non-targeted microbial populations towards this plant defense system, particularly in a context of living Brassicaceae used for soil pest control and disease suppression. Brassicaceae species are extensively used as biofumigants towards various soil-borne pests and pathogens (Angus et al., 1994; Brown and Morra, 1997). Our results show that this agricultural practice could also impact non-target microorganisms in soil with possible consequences on following cultures. Recent invasion of eastern North American forests by the non-native garlic mustard well illustrates these ecological issues. Although no single mechanism appears to explain its success, one of the most important is the root release of benzyl isothiocyanate of this Brassicaceae (Rodgers et al., 2008) inhibiting the growth of ectomycorrhizal fungi and reducing competitive abilities of native trees (Wolfe et al., 2008). The domination of this new species in forests finally changed the ecology and the function of these systems (Rodgers et al., 2008).
There is also a strong interest to alter levels of specific glucosinolates or generate novel glucosinolates in crop plants that could be an alternative to the use of pesticides. Concerns have been raised about the environmental impact associated with genetically modified organisms particularly GM crop. Variable and transient impacts on non-target microorganisms were already shown (De Vries et al., 1999; Lottmann et al., 2000; Dunfield and Germida, 2003; Castaldini et al., 2005; Rasche et al., 2006). We show here that manipulation of glucosinolate content in Brassicaceae could have some repercussions on the microbial community in their close soil environment. Possible consequences on specific microbial functions implicated in ecosystem functioning should be then evaluated.
References
Akiyama K, Matsuzaki K, Hayashi H . (2005). Plant sesquiterpenes induce hyphal branching in arbuscular mycorrhizal fungi. Nature 435: 824–827.
Altschul SF, Gish W, Miler W, Lipman DJ . (1990). Basic local alignment search tool. J Mol Biol 215: 403–410.
Angus JF, Gardner PA, Kirkegaard JA, Desmarchelier JM . (1994). Biofumigation: isothiocyanates released from Brassica roots inhibit growth of the take-all fungus. Plant Soil 162: 107–112.
Bais HP, Weir TL, Perry LG, Gilroy S, Vivanco JM . (2006). The role of root exudates in rhizosphere interactions with plants and other organisms. Ann Rev Plant Biol 57: 233–266.
Bak S, Olsen CE, Petersen BL, Møller BL, Halkier BA . (1999). Metabolic engineering of p-hydroxybenzylglucosinolate in Arabidopsis by expression of the cyanogenic CYP79A1 from Sorghum Metaboli. Plant J 20: 663–671.
Bending GD, Lincoln SD . (2000). Inhibition of soil nitrifying bacteria communities and their activities by glucosinolate hydrolysis products. Soil Biol Biochem 32: 1261–1269.
Berge O, Lodhi A, Brandelet G, Santaella C, Roncato M, Christen R et al. (2009). Rhizobium alamii sp. nov., an exopolysaccharide producing species isolated from legume and non-legume rhizospheres. Int J Syst Evol Microbiol 59: 367–372.
Borek V, Morra MJ, McCaffrey JP . (1996). Myrosinase activity in soil extracts. Soil Sci Soc Am J 60: 1792–1797.
Brabban AD, Edwards C . (1995). The effects of glucosinolates and their hydrolysis products on microbial growth. J Appl Bacteriol 79: 171–177.
Brader G, Mikkelsen MD, Halkier BA, Palva ET . (2006). Altering glucosinolate profiles modulates diseases resistance in plants. Plant J 2006: 758–767.
Broeckling CD, Broz AK, Bergelson J, Manter DK, Vivanco JM . (2008). Root exudates regulate soil fungal community composition and diversity. App Environ Microbiol 74: 738–744.
Brown PD, Tokuhisa JG, Reichelt M, Gershenzon J . (2003). Variation of glucosinolate accumulation among different organs and developmental stages of Arabidopsis thaliana. Phytochem 62: 471–481.
Brown PD, Morra MJ . (1997). Control of soil-borne plant pests using glucosinolate-containing plants. Adv Agron 61: 167–231.
Butler JL, Williams MA, Bottomley PJ, Myrold DD . (2003). Microbial community dynamics associated with rhizosphere carbon flow. Appl Environ Microbiol 69: 6793–6800.
Castaldini M, Turrini A, Sbrana C, Benedetti A, Marchionni M, Mocali S et al. (2005). Impact of Bt corn on rhizospheric and soil eubacterial communities and on beneficial mycorrhizal symbiosis in experimental microcosms. Appl Environ Microbiol 71: 6719–6729.
Chandra S, Choure K, Dubey RC, Maheshwari DK . (2007). Rhizosphere competent Mesorhizobium MP6 induces root hair curling, inhibits Sclerotinia sclerotiorum and enhances growth of Indian mustard (Brassica campestris). Braz. J Microbiol 38: 124–130.
Choesin DN, Boerner REJ . (1991). Allyl isothiocyanate release and the allelopathic potential of Brassica napus (Brassicaceae). Plant Physiol Biochem 78: 1083–1090.
Cohen MF, Yamasaki H, Mazzola M . (2005). Brassica napus seed meal soil amendment modifies microbial community structure, nitric oxide production and incidence of Rhizoctonia root rot. Soil Biol Biochem 37: 1215–1227.
Costa R, Gotz M, Mrotzek N, Lottmann J, Berg G, Smalla K . (2006). Effects of site and plant species on rhizosphere community structure as revealed by molecular analysis of microbial guilds. FEMS Microbiol Ecol 56: 236–249.
Dandie CE, Miller MN, Burton DL, Zebarth BJ, Trevors JT, Goyer C . (2007). Nitric oxide reductase-targeted real-time PCR quantification of denitrifier populations in soil. App Environ Microbiol 73: 4250–4258.
De Vries J, Harms K, Broer I, Kriete G, Mahn A, Düring K et al. (1999). The bacteriolytic activity in transgenic potatoes expressing a chimeric T4 lysozyme gene and the effect of T4 lysozyme on soil- and phytopathogenic bacteria. System Appl Microbiol 22: 280–286.
Dunfield KE, Germida JJ . (2003). Seasonal changes in the rhizosphere microbial communities associated with field-grown genetically modified canola (Brassica napus). Appl Environ Microbiol 69: 7310–7318.
Fahey JW, Zalemann AT, Talaley AT . (2001). The chemical diversity and distribution of glucosinolates and isothiocyanates among plants. Phytochem 56: 5–51.
Fierer N, Jackson JA, Vilgalys R, Jackson RB . (2005). Assessment of soil microbial community structure by use of taxon-specific quantitative PCR assays. Appl Environ Microbiol 71: 4117–4120.
Gimsing AL, Kirkegaard JA . (2006). Glucosinolate and isothiocyanate concentration in soil following incorporation of Brassica biofumigants. Soil Biol Biochem 38: 2255–2264.
Goldwasser Y, Yoneyama K, Xie X, Yoneyama K . (2008). Production of strigolactones by Arabidopsis thaliana responsible for Orobanche aegyptiaca seed germination. Plant Growth Regul 55: 21–28.
Grubb CD, Abel S . (2006). Glucosinolate metabolism and its control. TRENDS Plant Sci 11: 89–100.
Haichar FZ, Marol C, Berge O, Rangel-Castro JI, Prosser JI, Balesdent J et al. (2008). Plant host habitat and root exudates shape soil bacterial community structure. ISME J 2: 1221–1230.
Haichar FZ, Achouak W, Christen R, Heulin T, Marol C, Marais MF et al. (2007). Identification of cellulolytic bacteria in soil by stable isotope probing. Environ Microbiol 9: 625–634.
Halkier BA, Gershenzon J . (2006). Biology and biochemistry of glucosinolates. Ann Rev Plant Biol 57: 303–333.
Hein JW, Wolfe GV, Blee KA . (2008). Comparison of rhizosphere bacterial communities in Arabidopsis thaliana mutants for systemic acquired resistance. Microb Ecol 55: 333–343.
Hirsch AM . (2004). Plant-microbe symbioses: A continuum from commensalism to parasitism. Symbiosis 37: 345–363.
Keller M, Zengler K . (2004). Tapping into microbial diversity. Nat Rev Microbiol 2: 141–150.
Kirkegaard JA, Wong PTW, Desmarchelier JM . (1996). In vitro suppression of fungal root pathogens of cereals by Brassica tissues. Plant Path 45: 593–603.
Klein AN, Frigon D, Raskin L . (2007). Populations related to Alkanindiges, a novel genus containing obligate alkane degraders, are implicated in biological foaming in activated sludge systems. Environ Microbiol 9: 1898–1912.
Kliebenstein DJ, Rowe HC, Denby KJ . (2005). Secondary metabolites influence Arabidopsis/Botrytis interactions: variation in host production and pathogen sensitivity. Plant J 44: 25–36.
Kniskern JM, Traw MB, Bergelson J . (2007). Salicylic acid and jasmonic acid signaling defense pathways reduce natural bacterial diversity on Arabidopsis thaliana. MPMI 20: 1512–1522.
Kristensen C, Morant M, Olsen CE, Ekstrøm CT, Galbraith DW, Møller BL et al. (2005). Metabolic engineering of dhurrin in transgenic Arabidopsis plants with marginal inadvertent effects on the metabolome and transcriptome. Proc Natl Acad Sci USA 102: 1779–1784.
Leininger S, Urich T, Schloter M, Schwark L, Qi J, Nicol GW et al. (2006). Archaea predominate among ammonia-oxidizing prokaryotes in soils. Nature 442: 806–809.
Lottmann J, Heuer H, de Vries J, Mahn A, During K, Wackernagel W et al. (2000). Establishment of introduced antagonistic bacteria in the rhizosphere of transgenic potatoes and their effect on the bacterial community. FEMS Microbiol Ecol 33: 41–49.
Lu Y, Rosencrantz D, Liesack W, Conrad R . (2006). Structure and activity of bacterial community inhabiting rice roots and the rhizosphere. Environ Microbiol 8: 1351–1360.
Mahmood S, Paton GI, Prosser JI . (2005). Cultivation-independent in situ molecular analysis of bacteria involved in degradation of pentachlorophenol in soil. Environ Microbiol 7: 1349–1360.
Markmann K, Giczey G, Parniske M . (2008). Functional adaptation of a plant receptor-kinase paved the way for the evolution of intracellular root symbioses with bacteria. PLoS Biol 6: 497–506.
Martin-Laurent F, Barrès B, Wagschal I, Piutti S, Devers M, Soulas G et al. (2006). Impact of the maize rhizosphere on the genetic structure, the diversity and the atrazine-degrading gene composition of cultivable atrazine-degrading communities. Plant Soil 282: 99–115.
McCaig AE, Glover LA, Prosser JI . (2001). Numerical analysis of grassland bacterial community structure under different land management regimes by using 16S ribosomal DNA sequence data and denaturing gradient gel electrophoresis banding patterns. Appl Environ Microbiol 67: 4554–4559.
Mellon FA, Bennett RN, Holst B, Williamson G . (2002). Intact glucosinolate analysis in plant extracts by Programmed Cone Voltage Electrospray LC/MS: performance and comparison with LC/MS/MS methods. Anal Biochem 306: 83–91.
Muyzer G, Waal Ecd, Uitterlinden AG . (1993). Profiling of complex microbial populations by denaturing gradient gel electrophoresis analysis of polymerase chain reaction-amplified genes encoding for 16S rRNA. App Environ Microbiol 59: 695–700.
Nicol GW, Glover LA, Prosser JI . (2003). Spatial analysis of archaeal community structure in grassland soil. Appl Environ Microbiol 69: 7420–7429.
O'Callaghan KJ, Stone PJ, Hu X, Griffiths DW, Davey MR, Cocking EC . (2000). Effects of glucosinolates and flavonoids on colonization of the roots of Brassica napus by Azorhizobium caulinodans ORS571. Appl Environ Microbiol 66: 2185–2191.
Oger PM, Mansouri H, Nesme X, Dessaux Y . (2004). Engineering root exudation of Lotus toward the production of two novel carbon compounds leads to the selection of distinct microbial populations in the rhizosphere. Microb Ecol 47: 96–103.
Paterson E, Gebbing T, Abel C, Sim A, Telfer G . (2007). Rhizodeposition shapes rhizosphere microbial community structure in organic soil. New Phytol 173: 600–610.
Petersen BL, Chen S, Hansen CH, Olsen CE, Halkier BA . (2002). Composition and content of glucosinolates in developing Arabidopsis thaliana. Planta 214: 562–571.
Pieterse CMJ, Dicke M . (2007). Plant interactions with microbes and insects: from molecular mechanisms to ecology. TRENDS Plant Sci 12: 564–569.
Pongrac P, Vogel-Mikuš K, Regvar M, Tolrà R, Poschenrieder C, Barceló J . (2008). Glucosinolate profiles change during the life cycle and mycorrhizal colonization in Cd/Zn hyperaccumulator Thlaspi praecox (Brassicaceae). J Chem Ecol 34: 1038–1044.
Radajewski S, Ineson P, Parekh NR, Murrell JC . (2000). Stable-isotope probing as a tool in microbial ecology. Nature 403: 646–649.
Rangel-Castro JI, Killham K, Ostle N, Nicol GW, Anderson IC, Scrimgeour CM et al. (2005). Stable isotope probing analysis of the influence of liming on root exudate utilization by soil microorganisms. Environ Microbiol 7: 828–838.
Ranjard L, Lejon DPH, Mougel C, Scheher L, Merdinoglu D, Chaussod R . (2003). Sampling strategy in molecular microbial ecology: influence of soil sample size on DNA fingerprinting analysis of fungal and bacterial communities. Environ Microbiol 11: 1111–1120.
Rasche F, Hödl V, Poll C, Kandeler E, Gerzabek MH, van Elsas JD et al. (2006). Rhizosphere bacteria affected by transgenic potatoes with antibacterial activities compared with the effects of soil, wild-type potatoes, vegetation stage and pathogen exposure. FEMS Microbiol Ecol 56: 219–235.
Roberts KJ, Anderson RC . (2001). Effect of garlic mustard [Alliaria petiolata (Beib. Cavara & Grande)] extracts on plants and arbuscular mycorrhizal (AM) fungi. Am Midl Nat 146: 146–152.
Rodgers VL, Stinson KA, Finzi AC . (2008). Ready or not, garlic mustard is moving in: Alliaria petiolata as a member of Eastern North American forests. BioSci 58: 426–436.
Rumberger A, Marschner P . (2003). 2-phenylethylisothiocyanate concentration and microbial community composition in the rhizosphere of canola. Soil Biol Biochem 35: 445–452.
Schreiner RP, Koide RT . (1993). Mustards, mustard oils and mycorrhizas. Plant Physiol 123: 107–113.
Scott JS, Knudsen GR . (1999). Soil amendment effects of rape (Brassica napus) residues on pea rhizosphere bacteria. Soil Biol Biochem 31: 1435–1441.
Smith BJ, Kirkegaard JA . (2002). In vitro inhibition of soil microorganisms by 2-phenylethylisothiocyanate. Plant Pathol 51: 585–593.
Souza-Fagundes EM, Rosa LH, Gomes NCM, Santos MH, Pimente PF . (2004). Thiocyanate degradation by pure and mixed cultures of microorganisms. Braz J Microbiol 35: 333–336.
Thioulouse J, Chessel D, Dolédec S, Olivier JM . (1997). ADE-4: a multivariate analysis and graphical display software. Stat Comput 7: 75–83.
Trinick MJ, Hadobas PA . (1995). Formation of nodular structures on the non-legumes Brassica napus, B. campestris, B juncea and Arabidopsis thaliana with Bradyrhizobium and Rhizobium isolated from Parasponia spp or legumes grown in tropical soils. Plant Soil 172: 207–219.
Trosvik P, Rudi K, Næs T, Kohler A, Chan K-S, Jakobsen KS et al. (2008). Characterizing mixed microbial population dynamics using time-series analysis. ISME J 2: 707–715.
Vainio EJ, Hantula J . (2000). Direct analysis of wood-inhabiting fungi using denaturing gradient gel electrophoresis of amplified ribosomal DNA. Mycol Res 104: 927–936.
Vierheilig H, Bennett R, Kiddle G, Kaldorf M, Ludwig-Müller J . (2000). Differences in glucosinolate patterns and arbuscular mycorrhizal status of glucosinolate-containing plant species. New Phytol 146: 343–352.
Wang Y, Hsieh YP . (2002). Uncertainties and novel prospects in the study of the soil carbon dynamics. Chemosphere 49: 791–804.
Wang ET, Rogel MA, Sui XH, Chen WX, Martínez-Romero E, van Berkum P . (2002). Mesorhizobium amorphae, a rhizobial species that nodulates Amorpha fruticosa, is native to American soils. Arch Microbiol 178: 301–305.
Wathelet J-P, Iori R, Leoni O, Rollin P, Quinsac A, Palmieri S . (2004). Guidelines for glucosinolate analysis in green tissues used for biofumigation. Agroindustria 3: 257–266.
Weir BS, Turner SJ, Silvester WB, Park D-C, Young JM . (2004). Unexpectedly diverse Mesorhizobium strains and Rhizobium leguminosarum nodulate native legume genera of New Zealand, while introduced legume weeds are nodulated by Bradyrhizobium species. Appl Environ Microbiol 70: 5980–5987.
Wertz S, Degrange V, Prosser JI, Poly F, Commeaux C, Freitag T et al. (2006). Maintenance of soil functioning following erosion of microbial diversity. Environ Microbiol 8: 2162–2169.
Whiteley AS, Manefield M, Lueders T . (2006). Unlocking the ‘microbial black box’ using RNA-based stable isotope probing technologies. Cur Op Biotechnol 17: 67–71.
Wolfe BE, Rodgers VL, Stinson KA, Pringle A . (2008). The invasive plant Alliaria petiolata (garlic mustard) inhibits ectomycorrhizal fungi in its introduced range. J Ecol 96: 777–783.
Yan X, Chen S . (2007). Regulation of plant glucosinolate metabolism. Planta 226: 1343–1352.
Yanni YG, Rizk RY, Corich V, Squatini A, Ninke K, Philip-Hollingsworth S et al. (1997). Natural endophytic association between Rhizobium leguminosarum bv. trifolii and rice roots and assessment of potential to promote rice growth. Plant Soil 194: 99–114.
Zeng RS, Mallik AU, Setliff E . (2003). Growth stimulation of ectomycorrhizal fungi by root exudates of Brassicaceae plants: role of degraded compounds of indole glucosinolates. J Chem Ecol 29: 1337–1355.
Zhu H, Riely BK, Burns NJ, Ané J-M . (2006). Tracing nonlegume orthologs of legume genes required for nodulation and arbuscular mycorrhizal symbioses. Genet 172: 2491–2499.
Acknowledgements
We thank the ‘Groupe de Recherche Appliquées en Phytotechnologies’ (IBEB/SBVME, CEA Cadarache) for plant growth and labelling. We also thank Christine Marol (GRAP, CEA Cadarache) and Jérôme Balesdent (INRA, UR1119, Unité Géochimie des Sols et des Eaux) for δ13C analyses. We are grateful to Dr Barbara Ann Halkier (Denmark) for providing transgenic CYP79A1 plant seeds and to Agnès Gastaud for her helpful assistance during plant harvesting. We thank Philippe Roumagnac and Catherine Santaella for critical reading and pertinent suggestions and Arjan de Groot for english reviewing. We thank Christophe Merlin for the A. thaliana schematic representation. This work was supported by a CEA PhD grant and by the ANR ‘ECCO’ MICROGER program.
Author information
Authors and Affiliations
Corresponding author
Additional information
Supplementary Information accompanies the paper on The ISME Journal website (http://www.nature.com/ismej)
Supplementary information
Rights and permissions
About this article
Cite this article
Bressan, M., Roncato, MA., Bellvert, F. et al. Exogenous glucosinolate produced by Arabidopsis thaliana has an impact on microbes in the rhizosphere and plant roots. ISME J 3, 1243–1257 (2009). https://doi.org/10.1038/ismej.2009.68
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ismej.2009.68
Keywords
This article is cited by
-
A symbiotic footprint in the plant root microbiome
Environmental Microbiome (2023)
-
Study on the soil microbial community structure of the Rhizosphere of Camellia sinensis L. in Anping Village, Kaiyang County, Guizhou Province
Annals of Microbiology (2023)
-
Exploration of phyllosphere microbiomes in wheat varieties with differing aphid resistance
Environmental Microbiome (2023)
-
The Arabidopsis holobiont: a (re)source of insights to understand the amazing world of plant–microbe interactions
Environmental Microbiome (2023)
-
Green manure substitution for potassium fertilizer promotes agro-ecosystem multifunctionality via triggering interactions among soil, plant and rhizosphere microbiome
Plant and Soil (2023)