Abstract
Microbes are an essential component of marine food webs and biogeochemical cycles, and therefore precise estimates of their biomass are of significant value. Here, we measured single-cell biomass distributions of isolates from several numerically abundant marine bacterial groups, including Pelagibacter (SAR11), Prochlorococcus and Vibrio using a microfluidic mass sensor known as a suspended microchannel resonator (SMR). We show that the SMR can provide biomass (dry mass) measurements for cells spanning more than two orders of magnitude and that these estimates are consistent with other independent measures. We find that Pelagibacterales strain HTCC1062 has a median biomass of 11.9±0.7 fg per cell, which is five- to twelve-fold smaller than the median Prochlorococcus cell’s biomass (depending upon strain) and nearly 100-fold lower than that of rapidly growing V. splendidus strain 13B01. Knowing the biomass contributions from various taxonomic groups will provide more precise estimates of total marine biomass, aiding models of nutrient flux in the ocean.
Similar content being viewed by others
Introduction
Per-cell microbial biomass estimates are extremely important in parameterizing ecological and biogeochemical models (Ducklow, 2000). Beyond the average, the full distribution of single-cell biomass may also be important in biophysical models. However, single-cell biomass is non-trivial to determine. Established techniques include CHN analyzers (Lee and Fuhrman, 1987) and high-temperature catalytic oxidation (Fukuda et al., 1998), which when combined with cell counts can be used to estimate average biomass and elemental mass per cell. Alternatively, transmission electron microscopy, X-ray microanalysis and particle volume sensors based on the Coulter principle (also known as resistive pulse sensing) provide single cell mass or volume distributions (for example, Kogure and Koike, 1987; Fagerbakke et al., 1996; Loferer-Krößbacher et al., 1998). However, particle volume sensors are generally not sensitive enough to resolve the smallest marine bacteria and TEM-based analyses are difficult to scale-up since they require significant labor, technical skill and image processing.
Here we demonstrate the use of a micromechanical mass sensor to measure the single-cell biomass (dry mass) distributions of isolates from several ubiquitous marine bacterial groups including Pelagibacter (SAR11), Prochlorococcus and Vibrio. The SAR11 clade is estimated to have a global abundance of 2.4 × 1028 cells, and is the most abundant marine bacterial group (Morris et al., 2002). Prochlorococcus is the most abundant primary producer on Earth with a global estimate of 2.9 × 1027 cells (Flombaum et al., 2013) and supports a significant fraction of the secondary production that occurs in warm oligotrophic surface waters. Unlike Pelagibacter and Prochlorococcus, which are abundant open-ocean organisms (Partensky et al., 1999; Morris et al., 2002; Flombaum et al., 2013), Vibrio is commonly found in more productive waters at concentrations ~103 cells per ml (Takemura et al., 2014); however, massive, short-lived blooms have recently been documented, during which Vibrios can represent dominant community members (up to 50% of total bacteria; Gilbert et al., 2012; Westrich et al., 2016).
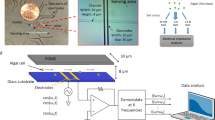
To measure single-cell biomass, we used suspended microchannel resonators (SMRs) - microcantilever-based microfluidic mass sensors that directly measure single-cell buoyant mass (Burg et al., 2007). The SMR consists of a hollow vibrating microcantilever with an internal microfluidic channel, which changes its resonant frequency proportionally to a cell’s buoyant mass whenever a cell flows through the interior of the cantilever. A cell’s buoyant mass is its total mass minus the mass of the fluid it displaces. To obtain dry mass (biomass), we combine information from paired buoyant mass measurements performed in H2O and D2O (Feijó Delgado et al., 2013). In pure H2O, a cell’s buoyant mass is only the buoyant mass of its dry material, as its intracellular water is neutrally buoyant. Similarly, in heavy water (D2O)—which permeates the cell and replaces internal H2O—a cell’s buoyant mass is also only the buoyant mass of its dry material. We exploit this property to obtain the density of a cell’s dry material (termed its dry density) with which we can convert from buoyant mass in H2O or D2O to biomass (Feijó Delgado et al., 2013), as shown in Figure 1a. We fixed cells so they would not lyse under hypoosmotic conditions, resuspended them in H2O or D2O and then measured their buoyant mass distributions. We then use these distributions to calculate the single-cell biomass distributions and uncertainty in their associated statistics (Supplementary Methods).
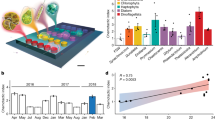
Measuring single-cell biomass (dry mass) of marine microbial isolates via Archimedes’ principle. (a) Paired measurements of a population of cells in H2O (ρfluid=1.0 g ml−1) and D2O (ρfluid=1.1 g ml−1) yields the dry density of the population, enabling conversion of buoyant mass distributions to dry mass distributions. (b) Biomass distributions for various cell types. (c) Log-log plot of genome size vs median single-cell biomass. Colors are as in (b).
Results and discussion
Previous work on natural bacterial assemblages has found nearly three orders of magnitude variation in single-cell biomass, from three femtograms to over a picogram (Loferer-Krößbacher et al., 1998). In accordance with this natural variation, we find that median biomass varies nearly 100-fold between the cultivated isolates from abundant marine bacterial clades. Pelagibacter median single-cell biomass was between 12 and 16 fg, Prochlorococcus between 60 and 158 fg and V. splendidus, depending on the growth stage, between 150 and 1000 fg (Figure 1b,Table 1). These values are consistent both with our measurements of buoyant mass in seawater-based media (Supplementary Figure S1) and with literature values, which is summarized below. Upon initial cultivation, Pelagibacterales strain HTCC1062 was reported to be extremely small, with an estimated cell volume of ca. 0.01 μm3 determined by TEM (Rappe et al., 2002). The carbon content of HTCC1062 was later estimated at 5.8 fg C per cell (Tripp et al., 2008), which corresponds to 11.6 fg of total biomass if carbon accounts for half the cell’s biomass. Our direct estimates of single-cell biomass for HTCC1062 and HTCC7211 are consistent with these previous reports and support the notion that Pelagibacterales are among the smallest known free-living cells. Previous estimates of Prochlorococcus biomass range from 15 to 94 fg C per cell (or 30–188 fg total biomass, assuming 50% carbon content) and were derived from strains belonging to the HLI clade (Bertilsson et al., 2003; Buitenhuis et al., 2012), the same as strain MED4 used here. Here we find median dry mass for Prochlorococcus to be between 60 and 158 fg, with higher values corresponding to the first direct biomass measurements of low-light-adapted Prochlorococcus (NATL2A and MIT9313), which we find can be 2-fold higher than their high-light-adapted relatives. We also note that across our Prochlorococcus and Pelagibacter strains, biomass increases monotonically with genome size (Figure 1c).
To our knowledge, the dry mass of Vibrio splendidus has not been previously measured; however, X-ray microanalysis of Vibrio natriegens yielded a geometric mean dry mass of 850 fg for exponential-phase cells and 145 fg for stationary-phase cells (Fagerbakke et al., 1996). Such drastic differences between exponential and stationary phase cells—exceeding 5-fold mass changes—have also been observed in E. coli (Feijó Delgado et al., 2013; Loferer-Krößbacher et al., 1998) and are correlated with a substantial reduction in RNA:protein ratio.
Our measurement also provides information on within-strain size variation. Strikingly, we found that the coefficient of variation (estimated using a robust metric—see Supplementary Methods) was highly consistent across strains, ranging from 26 to 30%. For unsynchronized cells, deterministically growing either linearly or exponentially from mass m0 to 2m0 and then dividing symmetrically, one would expect a robust CV of ~25%. While we expect our Pelagibacter and Vibrio populations to be unsychronized, the Prochlorococcus strains were grown under diel light conditions and thus were fixed toward the end of the day, just before division begins, so likely at their maximal size. This suggests that unsynchronized Prochlorococcus would likely have a broader size distribution than Pelagibacter or Vibrio. Estimates of cell-to-cell mass variation may be useful in constraining biophysical models of marine microbial behavior and could ultimately inform how uniquely a mass identifies a microbe or its growth state.
Our results show that SMR can provide single cell biomass estimates spanning nearly two orders of magnitude among marine bacteria, a variation that needs to be taken into account when considering the importance of different taxonomic groups in the global carbon cycle. Moreover, Pelagibacter and Prochlorococcus strains also demonstrate considerable biomass variation within taxonomic groups that may reflect the ecological constraints that different ecotypes or populations live under. We propose that SMR micromechanical mass sensors are an efficient means to determine biomass under different ecological conditions to further refine estimates of global microbial biomass.
References
Bertilsson S, Berglund O, Karl DM, Chisholm SW . (2003). Elemental composition of marine Prochlorococcus and Synechococcus: Implications for the ecological stoichiometry of the sea. Limnol Oceanogr 48: 1721–1731.
Buitenhuis ET, Li WKW, Lomas MW, Karl DM, Landry MR, Jacquet S . (2012). Picoheterotroph (Bacteria and Archaea) biomass distribution in the global ocean. Earth Syst Sci Data 4: 101–106.
Burg TP, Godin M, Knudsen SM, Shen W, Carlson G, Foster JS et al. (2007). Weighing of biomolecules, single cells and single nanoparticles in fluid. Nature 446: 1066–1069.
Ducklow H. (2000). Bacterial production and biomass in the oceans. In: Microbial Ecology of the Oceans. John Wiley and Sons: New York, NY, USA, pp 85–120.
Fagerbakke KM, Heldal M, Norland S . (1996). Content of carbon, nitrogen, oxygen, sulfur and phosphorus in native aquatic and cultured bacteria. Aquat Microb Ecol 10: 15–27.
Feijó Delgado F, Cermak N, Hecht VC, Son S, Li Y, Knudsen SM et al. (2013). Intracellular water exchange for measuring the dry mass, water mass and changes in chemical composition of living cells. PloS One 8: e67590.
Flombaum P, Gallegos JL, Gordillo RA, Rincón J, Zabala LL, Jiao N et al. (2013). Present and future global distributions of the marine Cyanobacteria Prochlorococcus and Synechococcus. Proc Natl Acad Sci USA 110: 9824–9829.
Fukuda R, Ogawa H, Nagata T, Koike I . (1998). Direct determination of carbon and nitrogen contents of natural bacterial assemblages in marine environments. Appl Environ Microbiol 64: 3352–3358.
Gilbert JA, Steele JA, Caporaso JG, Steinbrück L, Reeder J, Temperton B et al. (2012). Defining seasonal marine microbial community dynamics. ISME J 6: 298–308.
Kogure K, Koike I . (1987). Particle counter determination of bacterial biomass in seawater. Appl Environ Microbiol 53: 274–277.
Lee S, Fuhrman JA . (1987). Relationships between biovolume and biomass of naturally derived marine bacterioplankton. Appl Env Microbiol 53: 1298–1303.
Loferer-Krößbacher M, Klima J, Psenner R . (1998). Determination of bacterial cell dry mass by transmission electron microscopy and densitometric image analysis. Appl Environ Microbiol 64: 688–694.
Morris RM, Rappé MS, Connon SA, Vergin KL, Siebold WA, Carlson CA et al. (2002). SAR11 clade dominates ocean surface bacterioplankton communities. Nature 420: 806–810.
Partensky F, Hess WR, Vaulot D . (1999). Prochlorococcus, a marine photosynthetic prokaryote of global significance. Microbiol Mol Biol Rev 63: 106–127.
Rappe MS, Connon SA, Vergin KL, Giovannoni SJ . (2002). Cultivation of the ubiquitous SAR11 marine bacterioplankton clade. Nature 418: 630–633.
Takemura AF, Chien DM, Polz MF . (2014). Associations and dynamics of Vibrionaceae in the environment, from the genus to the population level. Front Microbiol 5: 38.
Tripp HJ, Kitner JB, Schwalbach MS, Dacey JWH, Wilhelm LJ, Giovannoni SJ . (2008). SAR11 marine bacteria require exogenous reduced sulphur for growth. Nature 452: 741–744.
Westrich JR, Ebling AM, Landing WM, Joyner JL, Kemp KM, Griffin DW et al. (2016). Saharan dust nutrients promote Vibrio bloom formation in marine surface waters. Proc Natl Acad Sci USA 113: 5964–5969.
Acknowledgements
We thank Prof Stephen Giovannoni (Oregon State University) for providing the Pelagibacter isolates in this study. NC acknowledges support from an MIT Poitras fellowship. This work was funded in part by US National Science Foundation grant OCE-1129359 to MFP and SRM; a grant from the Simons Foundation (Grant numbers 337262 to MIT and 329108 to U. Hawaii) to SWC and by the Institute for Collaborative Biotechnologies through grant W911NF-09-0001 (SRM) from the US Army Research Office.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no conflict of interest.
Additional information
Supplementary Information accompanies this paper on The ISME Journal website
Supplementary information
Rights and permissions
About this article
Cite this article
Cermak, N., Becker, J., Knudsen, S. et al. Direct single-cell biomass estimates for marine bacteria via Archimedes’ principle. ISME J 11, 825–828 (2017). https://doi.org/10.1038/ismej.2016.161
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ismej.2016.161
This article is cited by
-
High-throughput determination of dry mass of single bacterial cells by ultrathin membrane resonators
Communications Biology (2022)
-
Differentiating Live Versus Dead Gram-Positive and Gram-Negative Bacteria With and Without Oxidative Stress Using Buoyant Mass Measurements
Current Microbiology (2022)
-
Pili allow dominant marine cyanobacteria to avoid sinking and evade predation
Nature Communications (2021)
-
Redox-informed models of global biogeochemical cycles
Nature Communications (2020)
-
Rapid detection of microbiota cell type diversity using machine-learned classification of flow cytometry data
Communications Biology (2020)