Abstract
Intricate relationships between microorganisms structure the exchange of molecules between taxa, driving their physiology and evolution. On a global scale, this molecular trade is an integral component of biogeochemical cycling. As important microorganisms in the world’s oceans, diatoms and bacteria have a large impact on marine biogeochemistry. Here, we describe antagonistic effects of the globally distributed flavobacterium Croceibacter atlanticus on a phylogenetically diverse group of diatoms. We used the model diatom Thalassiosira pseudonana to study the antagonistic impact in more detail. In co-culture, C. atlanticus attaches to T. pseudonana and inhibits cell division, inducing diatom cells to become larger and increase in chlorophyll a fluorescence. These changes could be explained by an absence of cytokinesis that causes individual T. pseudonana cells to elongate, accumulate more plastids and become polyploid. These morphological changes could benefit C. atlanticus by augmenting the colonizable surface area of the diatom, its photosynthetic capabilities and possibly its metabolic secretions.
Similar content being viewed by others
Introduction
The ocean is a dilute environment with a microbial density of ~106 bacteria per milliliter of seawater (Whitman et al., 1998) and 10–1000 times fewer phytoplankton. To survive low nutrient concentrations, marine microbes commonly form close associations with one another to facilitate metabolic and nutrient exchanges. Consequently, interacting networks of phytoplankton and bacteria recycle most fresh organic carbon in the sunlit layers of the upper ocean, while a relatively small fraction of photosynthetic products remains dissolved in the water column or sinks to the sea floor (Azam et al., 1994; Azam, 1998).
Marine microbes experience a complex and heterogeneous chemical milieu that is unlike the macro-environment described by bulk measurements (Stocker, 2012). A thin boundary layer of fluid termed ‘the phycosphere’ surrounds algal cells (Bell and Mitchell, 1972). In the phycosphere, molecular gradients are governed by diffusion (Lazier and Mann, 1989) and altered by the activities of bacteria that accumulate through chemotaxis (Stocker and Seymour, 2012; Smriga et al., 2016) and reproduction. As the thickness of diffusive boundary layers scales with surface area (Karp-Boss et al., 1996), larger phytoplankton may harbor more bacteria than smaller phytoplankton. Bacteria modify the phycosphere by excreting metabolites, consuming metabolites produced by the algae or other bacteria, consuming and regenerating nutrients or making nutrients more bioavailable. As a result, bacteria turn the phycosphere into a dynamic micro-niche by shaping the environmental signals perceived by algae. Over time, co-evolution between algae and bacteria is likely to have resulted in specific interactions between these two groups.
Diatoms are an ecologically important group of phytoplankton that have a major role in supporting fisheries and driving the marine carbon cycle (Armbrust, 2009; Agusti et al., 2015). Specific bacterial phyla, mainly the Proteobacteria and Bacteroidetes (Amin et al., 2012), are commonly associated with diatoms. Beneficial and detrimental bacteria can be isolated from phytoplankton cultures, but their roles in the ocean and their mechanisms of antagonistic activity remain under-studied (Mayali and Azam, 2004; Amin et al., 2012). Beneficial bacteria typically provide a useful service to the diatom in exchange for organic nutrients; for example, Ruegeria pomeroyi provides vitamin B12 to Thalassiosira pseudonana in exchange for the organic sulfur compound 2,3-dihydroxypropane-1-sulfonate (Durham et al., 2015). Many detrimental bacteria are from the phylum Bacteroidetes or the class Gammaproteobacteria (Mayali and Azam, 2004) and are thought to harm phytoplankton by secreting proteases or other algaecides (Lee et al., 2000; Paul and Pohnert, 2011; Seyedsayamdost et al., 2011).
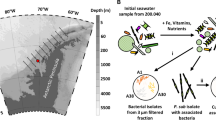
Previously, we isolated and identified bacteria belonging to the Sulfitobacter, Hyphomonas, Limnobacter, Marinobacter and Croceibacter genera from several cultures of the coastal diatom Pseudo-nitzschia multiseries originating from the Atlantic and Pacific Oceans. To probe interactions between the diatom and these bacteria, we made P. multiseries cultures axenic and performed a series of co-culture experiments with the different bacterial isolates. Some isolates increased the growth, whereas others inhibited the growth of P. multiseries (Amin et al., 2015). Here we report on the activities of Croceibacter atlanticus, an antagonistic flavobacterium from the P. multiseries consortium, which impacts diatom DNA content, cell division and cell morphology. This type of antagonistic effect has not, to our knowledge, been previously reported in the marine environment and could have important implications for understanding bloom dynamics and organic matter fluxes in the ocean.
Materials and methods
Culture acquisition and maintenance
Nitzschia sp. RCC 80 was acquired from the Roscoff Culture Collection (Roscoff, France); Thalassiosira weissfloggii CCMP 1052, T. oceanica CCMP 1003 and T. pseudonana CCMP 1335 from the National Center for Marine Algae and Microbiota (NCMA, East Boothbay, ME, USA); Pseudo-nitzschia multiseries CLNN-17 and P. multiseries CLN-47 from Stephen Bates and Claude Léger of Fisheries and Oceans Canada (Moncton, NB, Canada). P. multiseries CLN-47 is the product of a cross performed ca 2001 between environmental strains CL-143 and CL-147 collected from the Bay of Fundy, Canada; P. multiseries CLNN-17 is the product of a cross between CLN-47’s siblings CLN-35 and CLN-48 performed at the end of 2006. P. multiseries PC9 and GGA2, and P. fraudulenta OC1 were isolated from Penn Cove WA, in 2010, from Golden Gardens WA in 2010, and from Newport Beach CA in 2011 by Michael Carlson, and P. multiseries IOES-1 from East Sound WA in 2010 by Irina Oleinikov. Pseudo-nitzschia species were identified using PCR primers to amplify and sequence a polymorphic region of the internal transcribed spacer 1 (Hubbard et al., 2008).
All strains were maintained as semicontinuous batch cultures in f/2 medium (Guillard, 1975) at 13 °C in a 16-h light/8- h dark diurnal cycle (~120 μE m−2 s−1) except RCC 80 and CCMP 1003, which were maintained at 20 °C in 24- h light conditions (~120 μE m−2 s−1). For experimental work, Pseudo-nitzschia strains and CCMP 1335 were acclimated to the synthetic seawater medium Aquil (Price et al., 1989). To monitor diatom division rates, in vivo chlorophyll a fluorescence was measured daily with a 10-AU fluorometer (Turner Designs, Sunnyvale, CA, USA) 1–2 h before the end of the dark period.
The bacteria Marinobacter sp. SA14 and C. atlanticus SA60 were isolated from P. multiseries CLNN-17 in 2010; C. atlanticus HV2 was isolated from P. multiseries GGA2 in 2012. These strains were identified by their 16S ribosomal RNA gene sequence (Amin et al., 2015; GenBank accession no.: KM033246, KM033280, KM033233). We acquired the C. atlanticus type strain HTCC 2559 from the American Type Culture Collection (ATCC, Manassas, VA, USA). All bacterial isolates were stored in 15% glycerol stocks at −80 °C.
Pseudo-nitzschia strains IOES-1, PC9, GGA2 and OC1 were ‘cured’ of associated bacteria by adapting a protocol from Shishlyannikov et al. (2011), treating cultures with the detergent Triton X-100 (20 μg ml−1) and an antibiotic cocktail (per ml: 50 μg streptomycin, 67 μg gentamycin, 20 μg ciprofloxacin, 2.2 μg chloramphenicol and 100 μg ampicillin) as described in Amin et al. (2015). CCMP 1335 is maintained bacteria-free by the National Center for Marine Algae and Microbiota (NCMA). The absence of bacteria in these cultures was verified periodically by examining SYBR green-stained cultures with epifluorescence microscopy and by checking for bacterial growth in marine broth (MB; per l: 5 g peptone, 0.5 g yeast extract and 750 ml 0.2 μm filtered seawater; ZoBell, 1941).
Co-culture experiments
To assess the range of possible hosts for C. atlanticus, we performed a series of co-culturing experiments between different diatoms and the three C. atlanticus strains. Before each experiment, bacterial glycerol stocks were plated at room temperature on marine agar (MB with 1.5% w/v agar). Single colonies were inoculated into MB and grown overnight with shaking at 30 °C. Triplicate co-cultures were initiated by inoculating an exponentially growing diatom culture (initial relative chlorophyll a fluorescence units ~0.5) with ~4 × 105 bacteria ml−1 (initial bacterial concentration). This concentration of C. atlanticus was chosen because it is within the range of concentrations certain flavobacteria reach in the euphotic zone during a diatom bloom (Teeling et al., 2012). Control cultures were treated in each experiment with 0.01–0.20% (v/v) MB to match the initial concentrations of growth media added with bacteria.
More comprehensive experiments were conducted with T. pseudonana CCMP 1335 and C. atlanticus SA60 with the goal of honing in on the mechanism of antagonistic activity. T. pseudonana cells were grown exponentially in the organic carbon-free artificial medium Aquil (Price et al., 1989) to ~105 cells ml−1 and diluted to an initial concentration of 2500 cells ml−1 for the experiment. Overnight cultures of C. atlanticus were grown in MB to a density of ~109 cells ml−1 and either added directly to the diluted T. pseudonana cultures or added after washing 2 × with either Aquil, fresh MB, or the spent MB from an overnight grown culture after the bacteria were removed by centrifugation and the media filtered through a 0.2 μm filter. Washing consisted of centrifuging 1 ml cell cultures at 3220 × g and 10 °C for 5 min and replacing the supernatant with 1 ml media. These experiments were designed to determine whether C. atlanticus secretes a growth inhibitor into MB in the absence of diatoms, and whether C. atlanticus can grow on diatom-produced metabolites. As C. atlanticus is non-motile except for surface gliding (Cho and Giovannoni, 2003; McBride and Zhu, 2013), we increased the initial concentration of C. atlanticus added to each co-culture to ~2 × 106 cells ml−1 for an initial ratio of about 800 bacteria per diatom cell, enhancing the encounter rate with diatoms and increasing the likelihood of early attachment. For diatom control treatments, we added 0.2% v/v of fresh MB, fresh Aquil media, or 0.2% v/v of the sterile-filtered spent overnight MB media to cultures of T. pseudonana growing in Aquil.
Samples (1–2 ml) for flow cytometry were collected each day 0.5–1.5 h after the start of the light period and preserved in 1% buffered paraformaldehyde and 0.05% glutaraldehyde, incubated 20–30 min, flash frozen in liquid nitrogen and stored at −80 °C (Marie et al., 1999). Samples (1–2 ml) for epifluorescence microscopy were taken from a single replicate of each sample and were preserved in 8–9 ml Aquil with 2% glutaraldehyde and stored in the dark at 4 °C.
Specific growth rates (μ, day−1) were estimated from in vivo chlorophyll a fluorescence (relative fluorescence units) or cell counts (flow cytometry) measured during the exponential growth phase. Specific growth rate is defined as the natural log cell number or fluorescence at time t2 over the cell number or fluorescence at time t1 divided by the difference between t2 and t1. Percent growth inhibition for the co-cultures was defined as the difference between μcontrol and μco-culture divided by μcontrol multiplied by 100. Final cell concentrations (C, cells ml−1) were defined as the maximum value for each replicate growth curve measured by flow cytometry cell counts. Means were calculated from triplicate cultures unless otherwise indicated.
In a separate experiment, we tested the hypothesis that C. atlanticus requires amino acids as a supplementary nutrient in co-culture. T. pseudonana was grown exponentially in Aquil to ~3 × 105 cells ml−1 and diluted to an initial concentration of ~2000 cells ml−1. Overnight cultures of C. atlanticus were grown in MB to a density of ~109 cells ml−1, washed 2 × with Aquil and added to T. pseudonana at an initial concentration of ~2 × 106 cells for an initial ratio of 1000 bacteria per diatom cell. Each co-culture was amended with MEM Amino Acids solution (1:200 dilution, product no.: M5550, Sigma-Aldrich Inc., St Louis, MO, USA), MEM non-essential amino-acid solution (1:400 dilution, product no.: M7145, Sigma-Aldrich Inc.), a mix of both solutions, or nothing (duplicate samples each). After a 5-day incubation period, flow cytometry samples were collected as described previously.
Epifluorescence microscopy
Microscopy was used to provide phenotypic evidence of the interaction and to interpret flow cytometry results. Polyvinyl alcohol basal solution (Mowiol 4-88, Sigma-Aldrich Inc.) was prepared as described by Lunau et al. (2005) and stored as 200 μl aliquots in amber microcentrifuge tubes at −20 °C. Fresh staining solution was prepared by adding 2 μl of freshly prepared 1 m ascorbic acid and 4 μl SYBR Green I (Invitrogen, Waltham MA, USA) to 200 μl basal solution. Samples preserved in 2% glutaraldehyde were gently filtered onto black polycarbonate membrane filters (0.2 μm pore size, 25 mm diameter, Poretics, Osmonics Inc., Minnetonka, MN, USA). The filters were air dried on glass slides cleaned with 70% ethanol and then attached to the slide with 5 μl of staining solution. An additional 14 μl of staining solution was pipetted onto a cover slip that was flipped onto the filter. Air bubbles were squeezed out and the cover slip was sealed in place with clear nail polish. The samples were examined by epifluorescence microscopy with a 470/40 nm blue excitation 515 nm long-pass emission filter set (× 100 magnification, Eclipse 80i, Nikon, Tokyo, Japan; 11001v2, Chroma Corp., Rockingham, VT, USA) on the day of slide preparation and sections of the slide were imaged (MicroPublisher 3.3 RTV, QImaging, Surrey, British Columbia, Canada). Slides were stored in the dark at −20 °C. All microscopy images can be accessed from the figshare online digital repository (doi: 10.6084/m9.figshare.3471488).
ImageJ (Rasband, 2016), image analysis software, was used to measure diatom length and diameter (or just diameter depending on the cell orientation), plastid surface area, and to count nuclei from control and co-culture treatments at t=2 days and t=6 days. Each image was split into three channels Red Green Blue (RGB) and measurements were made on cells that were in focus. The wand tool was used in the red channel to measure the surface area covered by plastids for each diatom cell. The number of nuclei per cell was counted in the green channel. Length and diameter measurements (or minimum and maximum lengths) were also taken with the straight-line selection tool in the green channel, using nonspecific SYBR Green staining or attached bacteria as a guideline for the cell boundaries.
Flow cytometry
Flow cytometry was used both to count T. pseudonana and unattached C. atlanticus cells as well as to compare the light scatter and emission properties of the T. pseudonana population in co-culture and mono-culture. Flow cytometry measurements were made on an influx flow cytometer (BD/Cytopeia Inc., Seattle, WA, USA) equipped with two 10% neutral density filters and a 488 nm laser; data collection was triggered by forward light scatter. Samples were stained with SYBR Green I (1:10 000 dilution) and incubated in the dark on ice for at least 20 min. To minimize coincidence and improve population resolution, flow rates and sample concentrations were adjusted to achieve event rates below 1000 s−1. Two-micrometer beads were added to each sample as an internal standard (PolySciences Inc., Warrington, PA, USA). Samples were run for 5 min before data collection to allow the stain to equilibrate. Samples were run until at least 200 T. pseudonana cells were counted. Forward light scatter, side scatter and emission at 692 nm (40 nm band pass) (chlorophyll a fluorescence) were measured on a log scale. A flow meter was used to measure the volume sampled.
Flow cytometry standard files were processed with the flowCore and flowStats libraries in R (Ellis et al., 2015b; Hahne et al., 2015). Each population of cells was semiautomatically gated with the lymphGate function from the flowStats library after a rectangular pre-selection. T. pseudonana was gated in the forward scatter and 692/40 nm emission dimensions, whereas C. atlanticus and the 2 μm beads were gated in the forward scatter and 530/40 nm emission dimensions. These gating parameters were chosen to effectively separate the population of chlorophyll a-containing particles from the unattached bacteria. The accuracy of gated populations was visualized with scatter plots from the flowViz library (Ellis et al., 2015a). T. pseudonana distributions were normalized to the 2 μm beads before analysis. Beads were inadvertently not added to a T. pseudonana (t=3 days) replicate and a T. pseudonana+fresh MB (t=5 days) replicate before data collection; these samples were therefore removed from further analysis. Flow cytometry data can be accessed from the figshare online digital repository (doi: 10.6084/m9.figshare.3471455).
Statistical analyses
R was used to plot in vivo chlorophyll a fluorescence and flow cytometry data, and the R stats package (https://stat.ethz.ch/R-manual/R-devel/library/stats/DESCRIPTION) was used to perform all statistical analyses (v. 3.22.6). Two sample t-tests with a significance cut-off value of 0.05 were used to compare growth rates and final cell concentrations (μ, C) of different treatments as well as microscopy measurements made with ImageJ. A chi-squared test with a significance cutoff value of 0.05 was used to compare the distributions of nuclei counted in different treatments.
When discussing the co-culture treatments, we sometimes refer to T. pseudonana cells with attached bacteria as ‘colonized cells.’ The flow cytometry data were not normally distributed according to Shapiro–Wilk test results. Q-Q plots showed that control populations tended to have a slightly positive skew while colonized populations skewed increasingly negative over the course of the experiment, making data transformations intractable. We therefore used one-tail Wilcoxon rank sum tests with a significance cutoff of 0.05 to test for significant differences between the forward scatter, side scatter, and red fluorescence per cell of the T. pseudonana populations over time. Given that nonparametric tests with large sample sizes can be sensitive to small statistical differences with little biological meaning, we designed the test to detect a significant shift for a given time point if the colonized T. pseudonana population mean was greater than the control population mean by as much as the upper quartile of all samples from the control population, a relatively consistent metric over time. Replicate samples were merged to compare control and colonized treatments over time.
C. atlanticus genomes
We used the sequenced genomes of C. atlanticus to look for evidence of metabolic deficiencies in the bacterium that could be supplemented by metabolites from diatoms. C. atlanticus strain Pm1 was sequenced by the Joint Genome Institute as part of the P. multiseries genome project. The C. atlanticus strain Pm1 genome corresponds to the first scaffold in the prokaryotic assembly file (http://genome.jgi-psf.org/Psemu1/download/Psemu1_prokaryotic_scaffolds.fasta.gz). The C. atlanticus type strain genome sequence (HTCC 2559) is publically available from GenBank (accession number CP002046). Strain SA60 was sequenced at the Center for Environmental Genomics (Seattle, WA, USA). DNA from SA60 was extracted using a DNeasy Blood & Tissue Kit (Qiagen, Valencia, CA, USA), sheared into 1–2-kb fragments (Hydroshear, Applied Biosystems, Waltham, MA, USA), and a 50 bp × 50 bp mate-paired library was generated (Genomic Solutions Ltd, Cambridgeshire, UK). Emulsion PCR was used to attach the library to beads (Life Technologies, Research and Development Unit, Carlsbad, CA, USA); approximately 56 million beads were deposited and sequenced with a SOLiD 4 Next Generation Sequencer (Life Technologies). Sequencing generated 98 959 690 reads (or 49 479 845 mate pairs), which resulted in 74 025 907 reads (12 275 915 singlets and 30 874 996 mate pairs) after quality control. The short-read de Bruijn graph assembler Velvet was used for de novo contig assembly (Zerbino and Birney, 2008) and five scaffolds were generated with SEAStAR graph_ops (Iverson et al., 2012). Scaffold order was determined by alignment to the HTCC 2559 genome.
All three genomes were submitted to the Integrated Microbial Genomes Expert Review (IMG ER) gene prediction and annotation pipeline (Markowitz et al., 2009). The genome sequences and their annotations are publically available through IMG (http://img.jgi.doe.gov submissions: 15010, 11665, 11666). LASTZ was used to align the Pm1 and SA60 genomes to the type strain (Harris, 2007); aligned segments were plotted (Guy et al., 2010) in R (v. 3.22.6). Overall genome percent identity was calculated from a bl2seq alignment as the number of matches × 100 divided by the alignment length between each genome and HTCC 2559.
The BioCyc organism-specific Pathway/Genome Databases (PGDBs; Caspi et al., 2014) were used to compare the amino-acid biosynthesis, vitamin biosynthesis and inorganic nutrient utilization pathways of T. pseudonana CCMP 1335 and C. atlanticus HTCC 2559. The HTCC 2559 PGDB is not curated and the T. pseudonana PGDB is missing some genes, likely due to lack of homology. For this reason, the results were verified in the SA60 genome and partial pathways were sometimes considered to be evidence that the whole pathway is present in T. pseudonana.
Results and discussion
C. atlanticus is commonly associated with P. multiseries in culture
We identified three strains of C. atlanticus from three different P. multiseries cultures based on 16S ribosomal DNA sequence identity with the C. atlanticus type strain HTCC 2559 isolated from the Sargasso Sea (Cho and Giovannoni, 2003). The P. multiseries isolates were collected from different locales and maintained in culture for varying lengths of time. C. atlanticus HV2 was isolated after 2 years in culture with P. multiseries GGA2. C. atlanticus strain SA60 was isolated from P. multiseries strain CLNN-17, although the bacterial consortium was likely derived from the original parental strains (CL-143, CL-147), both isolated from the Bay of Fundy ~10 years prior. The genome sequence of C. atlanticus strain Pm1 was identified in sequences derived from the P. multiseries strain CLN-47 genome project. CLN-47 is an F1 generation cross produced ca 2001 from parental strains CL-143 and CL-147 and sequenced after 5 years in culture (Supplementary Table S1); CLN-47 has since been lost and no available isolate is associated with the C. atlanticus Pm1 sequence. The detection of C. atlanticus in several diatom cultures with different origins and culturing time in the laboratory indicates that it is a common, long-term member of the bacterial consortia associated with P. multiseries in culture.
Antagonistic impact of C. atlanticus on diatoms
When co-cultured with P. multiseries IOES-1, C. atlanticus strains SA60, HV2 and HTCC 2559 all displayed a similarly negative impact on the growth of the diatom. Within hours of inoculating P. multiseries with the different strains of C. atlanticus, in vivo chlorophyll a fluorescence of the co-culture began to decline, leveling off within about 6 days (Figure 1). Similar results were obtained with the closely related diatom P. fraudulenta (Table 1). Epifluorescence microscopy revealed that C. atlanticus strain HV2 proliferated on the cell surface of both P. multiseries and P. fraudulenta (Supplementary Figure S1). These results differ significantly from co-culturing with other bacteria from the consortium. For example, in previous work Sulfitobacter sp. SA11 enhanced the growth of P. multiseries (Amin, et al., 2015), whereas Marinobacter sp. SA14 did not influence diatom growth (Figure 1). The antagonistic effect of C. atlanticus was unexpected given the observed long-term co-habitation of this bacterium with various P. multiseries strains in culture.
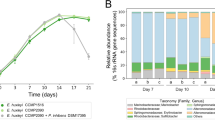
Effect of three different C. atlanticus isolates on the growth rates of P. multiseries IOES-1. The growth rates of P. multiseries are indistinguishable from each other when 0.05% v/v of either fresh MB media (n=2) or an overnight grown culture of Marinobacter sp. (strain SA14) at ~4 × 105 bacteria ml−1 (n=3) is added to diatom cultures at t=0 h. Growth rates and cell yield are significantly inhibited when overnight grown C. atlanticus isolates (n=3 each; Table 1, Bonferonni-adjusted P⩽0.05) are added to the diatom culture at a final concentration of ~4 × 105 bacteria ml−1. Error bars indicate±1 s.d. or the range (for n=3 or 2, respectively) and are shown when larger than symbol size.
All three C. atlanticus strains negatively impacted a variety of diatoms to varying degrees of severity (Table 1). Growth inhibition by C. atlanticus was not restricted to diatoms nor did it correlate with diatom phylogeny. Strong inhibition was observed with the closely related P. fraudulenta but not with Nitzschia sp. RCC 80; moderate inhibition was observed with two distantly related diatoms T. pseudonana and T. oceanica and no inhibition was observed with T. weissflogii (Table 1). Frazier et al. (2007) found that the pelagophyte Aureococcus anophagefferens was also inhibited by an isolate of C. atlanticus from Great South Bay, New York, although the extent of the inhibitory effect was not reported. Two of the seven tested diatoms were not impacted by the addition of C. atlanticus; neither of these diatom strains was axenic, raising the possibility that members of the pre-existing bacterial consortia from those diatoms mitigated the impacts of C. atlanticus. This possibility may explain how C. atlanticus remained a consistent member of the P. multiseries consortium without completely arresting diatom growth. Complex host-independent patterns of interaction have been observed in many host-dependent microbiomes (for example, Trosvik et al., 2010), and likely exist in the diatom microenvironment as well.
Mechanism of antagonistic effect of C. atlanticus on T. pseudonana
We chose to focus on the diatom-associated strain SA60 given the similarity in antagonistic phenotypes of the three C. atlanticus strains and their high genome sequence identities—the SA60 genome is 98% identical and the Pm1 genome is 99% identical to HTCC 2559 at the nucleotide level and both are missing a genome segment involved in alginate metabolism (Supplementary Figure S2). We examined in more detail, the interaction between strain SA60, and the model diatom T. pseudonana CCMP 1335 because the annotated whole genome of this diatom is available (Armbrust et al., 2004), with complementary metabolomic data (Longnecker et al., 2015), and because the cells are small enough to analyze with flow cytometry.
Epifluorescence microscopy revealed that in co-culture, C. atlanticus cells colonized the cell surface of T. pseudonana while a number of C. atlanticus cells remained unattached to the diatoms. As C. atlanticus is non-motile except for surface gliding (Cho and Giovannoni, 2003; McBride and Zhu, 2013), association with the diatom likely proceeds by random encounter followed by attachment and replication on the cell surface of the diatom. In acclimated non-axenic cultures and in nature, it may be that vertical transmission from parent to daughter cells is a more important mechanism of maintaining associations over multiple generations. The C. atlanticus population counted by flow cytometry consists of the unattached portion of the bacterial community—those cells that did not encounter T. pseudonana and any cells that detached from the diatom either naturally or during sample processing.
In the previous co-culture experiments (Figure 1,Table 1), an aliquot of overnight grown cultures of C. atlanticus cells in spent MB was added directly to the different diatom cultures. Although in each instance the added MB media was diluted 500–10 000-fold (Table 1), variable amounts of undefined organic matter, as well as any growth inhibitor potentially released by C. atlanticus into the MB, were also added to the co-cultures. We therefore carried out a series of experiments to determine potential sources of the antagonistic effects on diatoms.
First, we tested whether C. atlanticus constitutively released a diatom growth inhibitor into the MB during overnight growth in the absence of diatoms. An aliquot of filter-sterilized MB media from an overnight grown C. atlanticus culture was added to a T. pseudonana culture and did not significantly decrease the growth rate (μ1335+fresh MB=1.01±0.02 day−1, μ1335+spent MB=1.08±0.05 day−1, P=0.13) or final cell concentration (C1335+fresh MB=7.76±0.57 (× 105) cells ml−1, C1335+spent MB=7.71±0.80 (× 105) cells ml−1, P=0.94) of the diatom (Figure 2a). These results suggest that C. atlanticus does not constitutively release a growth inhibitor into MB under these conditions, at least at an effective concentration. By contrast, another study showed that the flavobacterium Kordia algicida constitutively secretes a protease that negatively impacts several diatoms (Paul and Pohnert, 2011). One diatom, Chaetoceros didymus, appears to deactivate the algicidal activity of K. algicida through production of its own counter-acting protease-degrading enzymes (Paul and Pohnert, 2013). It is not clear whether T. pseudonana can similarly degrade low concentrations of the C. atlanticus growth inhibitor.
Effects of C. atlanticus SA60 and its growth media on the growth of T. pseudonana CCMP 1335. (a) Filter-sterilized spent MB from an overnight culture of C. atlanticus has an indistinguishable effect relative to fresh MB on the growth of T. pseudonana, whereas a direct inoculate of overnight grown C. atlanticus drastically inhibits the growth of T. pseudonana; unattached C. atlanticus cells in co-culture display growth (inset). (b) C. atlanticus cells that were washed with Aquil do not grow in co-culture with T. pseudonana (inset) and do not impact the growth of T. pseudonana in co-culture. Symbols mark the mean of diatom and unattached bacterial cell concentration (circles, triangles) when grown alone, in the presence of C. atlanticus waste products, or in co-culture (white, gray, black; n=3 biological replicates±1 s.d. and are shown when larger than symbol size).
Second, we asked whether C. atlanticus required a component of the organic matter carried over in the MB inoculum to elicit the negative effect on the diatom. When C. atlanticus cells were washed with the organic carbon-free diatom medium Aquil before addition to the T. pseudonana culture, the bacteria did not divide in co-culture with T. pseudonana, did not attach to the T. pseudonana cells nor did they impact the growth rate or final cell concentration of the diatom (μ1335=0.93±0.05 day−1, μ1335+SA60=0.97±0.04 day−1, P=0.40; C1335=6.26±0.65 (× 105) cells ml−1, C1335+SA60=7.70±0.03 (× 105) cells ml−1, P=0.06; Table 2, Figure 2b). In contrast, a C. atlanticus overnight culture washed with spent MB before addition to the co-culture remained antagonistic indicating that the washing step itself did not impact the C. atlanticus phenotype (μ1335+SA60=0.39±0.07 day−1, μ1335+SA60 centrifuged=0.44±0.03 day−1, P=0.29; C1335+SA60=4.15±0.1 (× 104) cells ml–1, C1335+SA60 centrifuged=4.07±1.42 (× 104) cells ml–1, P=0.9; Supplementary Figure S3). This result suggests that specific nutrients present in MB and required in co-culture were not supplied by the diatom, at least at the start of the experiment.
To rule out that washing the cells in Aquil was toxic to the bacterium and inhibited its antagonistic activity, amino acids were added to Aquil-washed C. atlanticus in co-culture with T. pseudonana and bacterial growth was rescued (Supplementary Figure S4). The amino-acid formulations used in this experiment include two mixtures of amino acids (named essential and non-essential based on cultured mammalian cell nutrient requirements). C. atlanticus requires both mixtures for growth, implying that there are amino acids in both solutions that are required by C. atlanticus.
The extent of the negative impact when supplemented with MB appeared to be related to C. atlanticus growth rate in co-culture. When C. atlanticus cells were washed with spent MB rather than fresh MB, the unattached C. atlanticus cells displayed a lower growth rate in co-culture (μspent=1.81±0.08 day−1, μfresh=2.26±0.11 day−1, P=0.01) and reached a lower final cell concentration (Cspent=2.57±0.06 (× 107) cells ml–1, Cfresh=3.01±0.01 (× 107) cells ml–1, P<0.01). This lower C. atlanticus growth rate translated into a significantly reduced impact on the T. pseudonana growth rate (59% vs 71% inhibition, Table 2). These results reinforce the idea that the observed detrimental effect is driven by proliferation of C. atlanticus whose growth is supported by nutrients present in MB.
These co-culturing experiments show that the Aquil-washing step eliminates nutrients required for the growth of C. atlanticus in co-culture, and that early exponential phase cultures of T. pseudonana do not provide sufficient organic matter to support C. atlanticus cell division. C. atlanticus growth can, however, be rescued by the addition of amino acids or MB to co-culture. In the absence of nutrients, C. atlanticus does not attach to the diatom cells and the antagonistic phenotype is not induced. In contrast, when C. atlanticus cells are added to the diatom cultures along with sufficient organic nutrients, the unattached cells continue to divide, presumably attaining a cell density that provides a sufficiently high encounter rate with the diatom cells. C. atlanticus likely has a mechanism to recognize and attach to diatom cells; perhaps algal exudates serve as infochemicals that induce colonization after an encounter. Once C. atlanticus cells attach to a diatom, they appear to proliferate on the cell surface, likely due to the enhanced flux of diatom-released nutrients at the cell surface not seen in the bulk media.
Effects of C. atlanticus on the cell size, morphology and DNA content of T. pseudonana
Microscopy and flow cytometry measurements of diatoms in mono-culture and co-culture indicated that over the course of a few days, T. pseudonana cells with attached bacteria elongate (Figures 3a and b, Supplementary Figure S5), are filled with more plastids (Figure 3c), and typically possess more nuclei (Figure 3d) than cells without attached bacteria (Figure 4). In treatments where both C. atlanticus and MB were added, the vast majority of T. pseudonana cells hosted attached bacteria by the second day of co-culture (Figure 4a).
Microscopy measurements comparing colonized and control diatom cells after 2 and 6 days in culture. Box-and-whiskers plots of the (a) maximum and (b) minimum lengths and (c) the surface area per cell covered by plastids show the distribution of data including the median, quartiles, and range. N represents the number of cell measurements made. (d) Histograms of nucleus count per cell for colonized and control diatoms after 2 and 6 days in culture. Asterisks represent treatments that are statistically different from the control for the same incubation period (P<0.05).
Attachment of C. atlanticus to T. pseudonana and impacts on diatom morphology. (a) Representative epifluorescence micrographs from the experiment portrayed in Figure 2 show C. atlanticus attachment to T. pseudonana and morphological changes over the course of the experiment. Diatom plastids autofluoresce red, SYBR Green-stained DNA from the diatom nucleus and the bacteria fluoresce green. The co-culture micrographs for day 4 and 6 focus on the diatoms rather than the attached bacteria to show examples of polyploidy. Scale bar=10 μm. (b) Increased forward and side scatter values (blue, turquoise) over time show how C. atlanticus impacts the size and shape of T. pseudonana. Shifts in the 692/40 nm emission distribution (red) over time indicate that C. atlanticus impacts the amount of chlorophyll per cell. Colored polygons are representative distributions of T. pseudonana population scatter and fluorescence characteristics in co-culture; white polygons show the same for a control population. Distributions represent the fractions of total cells in a population.
In examining both the means and distributions of flow cytometry parameters and microscopy measurements, a time course of the impact of C. atlanticus on T. pseudonana was apparent. Within 2 days of the addition of C. atlanticus cells, the growth rate of T. pseudonana in co-culture decreased significantly relative to the control (Figure 2a), whereas the chlorophyll a fluorescence (692/50 nm emission) per cell in colonized populations increased significantly (P<0.001) (Figure 4b, Supplementary Figure S6). Microscopy measurements of the cellular surface area covered by plastids suggest that elevated chlorophyll a fluorescence is correlated with an increase in the number of plastids per cell (Figures 3c and 4b). By day 3, the forward scatter of colonized T. pseudonana populations increased (P<0.001) (Figure 4b,Supplementary Figure S6). Increased forward scatter is often correlated to increased cell size (Olson et al., 1989), although this relationship is sometimes affected by other refractive properties of the cell (Shapiro, 2003). However, microscopy measurements of T. pseudonana diameter and length (Figures 3a and b) lead us to conclude that increased forward scatter of colonized cells can be attributed to their increased cell size. By day 4, side scatter values also increase significantly (P<0.001) (Figure 4b, Supplementary Figure S6). The reason for these shifts are not known, but changes in side scatter distributions may be related to cell shape, cytoplasmic granularity and differences in internal cell structures (Shapiro, 2003). We also observed that T. pseudonana sank to the bottom of the culture tubes in treatments where the cells were colonized, possibly as a result of morphological changes. The similar pattern between cell length measurements and cell area covered by plastids suggests that the number of plastids scales with cell size.
Epifluorescence microscopy indicated that by day 6 the larger, SYBR Green-stained colonized T. pseudonana cells commonly possessed two or more nuclei rather than a single nucleus as observed in control cultures (Figure 3d). A significant proportion of cells without a nucleus were also observed on day 6 using microscopy (Figure 3d). This is suggestive of nuclear disintegration and DNA fragmentation, which accompanies apoptosis (Huang et al., 2005). C. atlanticus thus appears to manipulate the cell cycle of the diatom, stimulating both polyploidy and nuclear disintegration.
Increased DNA content is often correlated with enhanced cell size (Von Dassow et al., 2008; Koester et al., 2010) as well as increased metabolic activity (Wildermuth, 2010; Edgar et al., 2014). We hypothesize that manipulation of the diatom cell cycle could therefore be an effective strategy for C. atlanticus to induce morphological and physiological changes in its host that enhance the release of required metabolites. For example, C. atlanticus is unable to synthesize the amino acids isoleucine, valine, leucine and arginine, all of which T. pseudonana produces within its plastids and mitochondria, which may be more abundant now that the diatom is larger (Supplementary Table S2, Armbrust et al., 2004). In addition, the C. atlanticus genome encodes a variety of peptidases and carbohydrate active enzymes (CAZYmes) and C. atlanticus can grow in defined media with amino acids as its sole source of organic carbon and nitrogen (data not shown). Secretion of carbohydrates and peptides by T. pseudonana (Longnecker et al., 2015) could also fulfill C. atlanticus nutritional requirements.
Polyploidy occurs naturally in a variety of plant, mammalian and insect cell types (Edgar et al., 2014). It has also been observed at nutrient exchange points between fungal or bacterial pathogens or symbionts and their host cells (Wildermuth, 2010). As ploidy correlates linearly with gene expression and metabolic flux, it has been hypothesized that increased DNA content may be a common biological mechanism to cope with increased metabolic demands (Wildermuth, 2010). In particular, primary metabolism (glycolysis, respiration and fermentation), transport, and plastid-localized production of metabolites are sensitive to increases in ploidy (Wildermuth, 2010). In mammals, pathogenic bacteria are known to target cell cycle progression of their host cells through production of effectors known as cyclomodulins (Nougayrède et al., 2005). These effectors regulate the expression of cyclins and cyclin-dependent kinases, required for cell cycle progression. Analogously, growth-promoting and pathogenic bacteria can interfere with the plant cell cycle hormonal balance to modulate host cyclin and cyclin-dependent kinase activity (Stes et al., 2011). Whether C. atlanticus manipulates T. pseudonana cell division through a similar disruption of cyclin-mediated cell cycle progression requires further study.
Conclusion
Flavobacteria such as C. atlanticus have a reputation as fast-growing r-strategists that specialize on initiating organic matter degradation near the end of algal blooms (Teeling, et al., 2012). Our results raise the possibility that some flavobacteria could contribute to diatom bloom decline if they, like C. atlanticus, are capable of halting cell division. The colonized diatom population appears to survive until the late stationary phase when there is some indication of nuclear disintegration. As C. atlanticus is only capable of gliding motility, it cannot actively seek out a new host and must continue to colonize an arrested diatom cell. By causing T. pseudonana to elongate, C. atlanticus increases the surface area of its niche. In addition, C. atlanticus benefits from colonizing a live diatom that continues to synthesize photosynthetic products. However, once nutrients are depleted in solution and the diatom stops its metabolic activities, it would be more beneficial for C. atlanticus to consume all available organics from the failing diatom and detach from the cell before it sinks.
The mechanisms of diatom–bacteria interactions can give us new insights into the dynamics of marine microbial ecology. Although we do not yet know whether bacteria affect the cell cycle of diatoms in nature, we have evidence that C. atlanticus can affect the DNA content of T. pseudonana and prior evidence that Sulfitobacter sp. can upregulate G1-S phase cyclins of P. multiseries in culture (Amin et al., 2015). It may be that the potential for cell cycle manipulations is widespread in the marine environment and is a common ecological strategy. C. atlanticus sequences are prevalent across the global ocean (Yooseph et al., 2010), and the interaction appears to lack specificity. If C. atlanticus is capable of antagonizing a variety of phytoplankton in nature, then this microbe could have an important role in the remineralization of organic matter in the ocean. Interactions between C. atlanticus and other algal-associated bacteria may also be involved in mediating these antagonistic interactions.
References
Agusti S, González-Gordillo JI, Vaqué D, Estrada M, Cerezo MI, Salazar G et al. (2015). Ubiquitous healthy diatoms in the deep sea confirm deep carbon injection by the biological pump. Nat Commun 6: 7608.
Amin SA, Hmelo LR, Van Tol HM, Durham BP, Carlson LT, Heal KR et al. (2015). Interaction and signalling between a cosmopolitan phytoplankton and associated bacteria. Nature 522: 98–101.
Amin SA, Parker MS, Armbrust EV . (2012). Interactions between diatoms and bacteria. Microbiol Mol Biol Rev 76: 667–684.
Armbrust EV, Berges JA, Bowler C, Green BR, Martinez D, Putnam NH et al. (2004). The genome of the diatom Thalassiosira pseudonana: ecology, evolution, and metabolism. Science 306: 79–86.
Armbrust EV . (2009). The life of diatoms in the world’s oceans. Nature 459: 185–192.
Azam F . (1998). Microbial control of oceanic carbon flux: the plot thickens. Science 280: 694–696.
Azam F, Smith DC, Steward GF, Hagström Å . (1994). Bacteria-organic matter coupling and its significance for oceanic carbon cycling. Microb Ecol 28: 167–179.
Bell W, Mitchell R . (1972). Chemotactic and growth responses of marine bacteria to algal extracellular products. Biol Bull 143: 265–277.
Caspi R, Altman T, Billington R, Dreher K, Foerster H, Fulcher CA et al. (2014). The MetaCyc database of metabolic pathways and enzymes and the BioCyc collection of Pathway/Genome Databases. Nucleic Acids Res 42: D459–D471.
Cho J, Giovannoni SJ . (2003). Croceibacter atlanticus gen. nov., sp. nov., a novel marine bacterium in the family Flavobacteriaceae. Syst Appl Microbiol 26: 76–83.
Durham BP, Sharma S, Luo H, Smith CB, Amin SA, Bender SJ et al. (2015). Cryptic carbon and sulfur cycling between surface ocean plankton. Proc Natl Acad Sci USA 112: 453–457.
Edgar BA, Zielke N, Gutierrez C . (2014). Endocycles: a recurrent evolutionary innovation for post-mitotic cell growth. Nat Rev Mol Cell Biol 15: 197–210.
Ellis B, Gentleman R, Hahne F, Meur NL, Sarkar D . (2015a) flowViz: visualization for flow cytometry. R package version 1.28.22.
Ellis B, Haaland P, Hahne F, Meur NL, Gopalakrishnan N, Spidlen J et al. (2015b) flowCore: flowCore: basic structures for flow cytometry data. R package version 1.30.7.
Frazier AD, Rowe JM, Rentz CA, Gobler CJ, Wilhelm SW . (2007). Bacterial lysis of Aureococcus anophagefferens CCMP 1784 (Pelagophyceae). J Phycol 43: 461–465.
Guillard RRL . (1975). Culture of phytoplankton for feeding marine invertebrates. In: Smith WL, Chanley MH (eds). Culture of Marine Invertebrate Animals. Springer US: New York, NY, USA, pp. 29–60.
Guy L, Roat Kultima J, Andersson SGE . (2010). genoPlotR: comparative gene and genome visualization in R. Bioinformatics 26: 2334–2335.
Hahne F, Gopalakrishnan N, Khodabakhshi AH, Wong C, Lee K . (2015) flowStats: statistical methods for the analysis of flow cytometry data. R package version 3.22.6.
Harris RS . (2007) Improved Pairwise Alignment of Genomic DNA. The Pennsylvania State University. Available from: http://www.bx.psu.edu/∼rsharris/lastz/.
Huang X, Halicka HD, Traganos F, Tanaka T, Kurose A, Darzynkiewicz Z . (2005). Cytometric assessment of DNA damage in relation to cell cycle phase and apoptosis. Cell Prolif 38: 223–243.
Hubbard KA, Rocap G, Armbrust EV . (2008). Inter- and intraspecific community structure within the diatom genus pseudo-nitzschia (Bacillariophyceae). J Phycol 44: 637–649.
Iverson V, Morris RM, Frazar CD, Berthiaume CT, Morales RL, Armbrust EV . (2012). Untangling genomes from metagenomes: revealing an uncultured class of marine Euryarchaeota. Science 335: 587–590.
Karp-Boss L, Boss E, Jumars P . (1996). Nutrient fluxes to planktonic osmotrophs in the presence of fluid motion. Oceanogr Mar Biol Annu Rev 34: 71–107.
Koester JA, Swalwell JE, von Dassow P, Armbrust EV . (2010). Genome size differentiates co-occurring populations of the planktonic diatom Ditylum brightwellii (Bacillariophyta). BMC Evol Biol 10: 1.
Lazier JRN, Mann KH . (1989). Turbulence and the diffusive layers around small organisms. Deep Sea Res Part A Oceanogr Res Pap 36: 1721–1733.
Lee SO, Kato J, Takiguchi N, Kuroda A, Ikeda T, Mitsutani A et al. (2000). Involvement of an extracellular protease in algicidal activity of the marine bacterium Pseudoalteromonas sp. strain A28. Appl Environ Microbiol 66: 4334–4339.
Longnecker K, Kido Soule MC, Kujawinski EB . (2015). Dissolved organic matter produced by Thalassiosira pseudonana. Mar Chem 168: 114–123.
Lunau M, Lemke A, Walther K, Martens-Habbena W, Simon M . (2005). An improved method for counting bacteria from sediments and turbid environments by epifluorescence microscopy. Environ Microbiol 7: 961–968.
Marie D, Partensky F, Vaulot D, Brussard C . (1999). Enumeration of phytoplankton, bacteria, and viruses in marine samples. In: Robinson JP (ed). Current Protocols in Flow Cytometry. John Wiley & Sons, Inc.: New York, NY, USA, pp. 11.11.1–11.11.15.
Markowitz VM, Mavromatis K, Ivanova NN, Chen I-MA, Chu K, Kyrpides NC . (2009). IMG ER: a system for microbial genome annotation expert review and curation. Bioinformatics 25: 2271–2278.
Mayali X, Azam F . (2004). Algicidal bacteria in the sea and their impact on algal blooms. J Eukaryot Microbiol 51: 139–144.
McBride MJ, Zhu Y . (2013). Gliding motility and por secretion system genes are widespread among members of the phylum bacteroidetes. J Bacteriol 195: 270–278.
Nougayrède J-P, Taieb F, De Rycke J, Oswald E . (2005). Cyclomodulins: bacterial effectors that modulate the eukaryotic cell cycle. Trends Microbiol 13: 103–110.
Olson RJ, Zettler ER, Anderson OK . (1989). Discrimination of eukaryotic phytoplankton cell types from light scatter and autofluorescence properties measured by flow cytometry. Cytometry 10: 636–643.
Paul C, Pohnert G . (2013). Induction of protease release of the resistant diatom Chaetoceros didymus in response to lytic enzymes from an Algicidal bacterium. PLoS One 8: e57577.
Paul C, Pohnert G . (2011). Interactions of the Algicidal bacterium Kordia algicida with diatoms: regulated protease excretion for specific algal lysis. PLoS One 6: e21032.
Price NM, Harrison GI, Hering JG, Hudson RJ, Nirel PMV, Palenik B et al. (1989). Preparation and chemistry of the artificial algal culture medium Aquil. Biol Oceanogr 6: 443–461.
Rasband WS . (2016) ImageJ, U.S. National Institutes of Health, Bethesda, MD, USA. Available at: http://imagej.nih.gov/ij/, 1997–2015.
Seyedsayamdost MR, Case RJ, Kolter R, Clardy J . (2011). The Jekyll-and-Hyde chemistry of Phaeobacter gallaeciensis. Nat Chem 3: 331–335.
Shapiro HM . (2003) Physical parameters and their uses. In: Shapiro HM (ed). Practical Flow Cytometry. John Wiley & Sons, Inc.: New York, NY, USA, pp. 273–285.
Shishlyannikov SM, Zakharova YR, Volokitina NA, Mikhailov IS, Petrova DP, Likhoshway YV . (2011). A procedure for establishing an axenic culture of the diatom Synedra acus subsp. radians (Kütz.) Skabibitsch from Lake Baikal. Limnol Oceanogr Methods 9: 478–484.
Smriga S, Fernandez VI, Mitchell JG, Stocker R . (2016). Chemotaxis toward phytoplankton drives organic matter partitioning among marine bacteria. Proc Natl Acad Sci USA 113: 1576–1581.
Stes E, Vandeputte OM, El Jaziri M, Holsters M, Vereecke D . (2011). A successful bacterial coup d’État: how Rhodococcus fascians redirects plant development. Annu Rev Phytopathol 49: 69–86.
Stocker R . (2012). Marine microbes see a sea of gradients. Science 338: 628–633.
Stocker R, Seymour JR . (2012). Ecology and physics of bacterial chemotaxis in the ocean. Microbiol Mol Biol Rev 76: 792–812.
Teeling H, Fuchs BM, Becher D, Klockow C, Gardebrecht A, Bennke CM et al. (2012). Substrate-controlled succession of marine bacterioplankton populations induced by a phytoplankton bloom. Science 336: 608–611.
Trosvik P, Rudi K, Straetkvern KO, Jakobsen KS, Naes T, Stenseth NC . (2010). Web of ecological interactions in an experimental gut microbiota. Environ Microbiol 12: 2677–2687.
Von Dassow P, Petersen TW, Chepurnov VA, Armbrust EV . (2008). Inter- and intraspecific relationships between nuclear DNA content and cell size in selected members of the centric diatom genus Thalassiosira (Bacillariophyceae). J Phycol 44: 335–349.
Whitman WB, Coleman DC, Wiebe WJ . (1998). Prokaryotes: the unseen majority. Proc Natl Acad Sci USA 95: 6578–6583.
Wildermuth MC . (2010). Modulation of host nuclear ploidy: a common plant biotroph mechanism. Curr Opin Plant Biol 13: 449–458.
Yooseph S, Nealson KH, Rusch DB, McCrow JP, Dupont CL, Kim M et al. (2010). Genomic and functional adaptation in surface ocean planktonic prokaryotes. Nature 468: 60–66.
Zerbino DR, Birney E . (2008). Velvet: algorithms for de novo short read assembly using de Bruijn graphs. Genome Res 18: 821–829.
ZoBell CE . (1941). Studies on marine bacteria. I. The cultural requirements of heterotrophic aerobes. J Mar Res 4: 42–75.
Acknowledgements
We thank Michael Carlson, Irina Oleinikov, Lyndsey Swanson and Megan Schatz for supplying and maintaining diatom cultures; Chris Berthiaume and Rhonda Morales for doing the genome sequencing and assembly work; Rhonda Morales, Jarred Swalwell and Francois Ribalet for advice on flow cytometry data collection and processing. This work was supported by Gordon and Betty Moore Foundation grant GBMF3776 awarded to E Virginia Armbrust. Helena van Tol was partially supported by a Natural Sciences and Engineering Research Council of Canada Postgraduate Scholarship Master’s program award (NSERC PGS M 489334).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no conflict of interest.
Additional information
Supplementary Information accompanies this paper on The ISME Journal website
Supplementary information
Rights and permissions
About this article
Cite this article
van Tol, H., Amin, S. & Armbrust, E. Ubiquitous marine bacterium inhibits diatom cell division. ISME J 11, 31–42 (2017). https://doi.org/10.1038/ismej.2016.112
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ismej.2016.112
This article is cited by
-
Progress and challenges in exploring aquatic microbial communities using non-targeted metabolomics
The ISME Journal (2023)
-
Hyperactive nanobacteria with host-dependent traits pervade Omnitrophota
Nature Microbiology (2023)
-
Flavobacterial exudates disrupt cell cycle progression and metabolism of the diatom Thalassiosira pseudonana
The ISME Journal (2022)
-
A comparative whole-genome approach identifies bacterial traits for marine microbial interactions
Communications Biology (2022)
-
The ecological responses of bacterioplankton during a Phaeocystis globosa bloom in Beibu Gulf, China highlighted by integrated metagenomics and metatranscriptomics
Marine Biology (2022)