Abstract
We estimate gene flow and patterns of genetic diversity in Albizia lebbeck, an invasive leguminous tree in the dry forest of southwestern Puerto Rico. Genetic diversity estimates calculated for 10 populations of 24 trees each indicated that these populations may have been formed from multiple introductions. The presence of unique genotypes in the northernmost populations suggests that novel genotypes are still immigrating into the area. This combination of individuals from disparate locations led to high estimates of genetic diversity (He=0.266, P=0.67). Indirect estimates of gene flow indicate that only 0.69 migrants per generation move between populations, suggesting that genetic diversity within populations should decrease due to genetic drift. Since migration–drift equilibrium was not found, however, this estimate needs to be viewed with caution. The regular production of pods in this outcrossing species (tm=0.979) indicates that sufficient outcross pollen is received to insure successful reproduction. Direct estimates of gene flow indicate that between 44 and 100% of pollen received by trees in four small stands of trees (n<11) was foreign. The role of gene flow in facilitating the spread of this invasive plant species is discussed.
Similar content being viewed by others
Introduction
The spread of an introduced plant species across a landscape depends largely upon the species' mode of reproduction and its dispersal capabilities. Reproductively independent selfing species can establish a new population with a single individual, allowing rapid colonization of new sites (Baker, 1965). For predominantly outcrossing species, however, the seed production needed for population growth and the colonization of new sites depends upon the availability of compatible pollen. In newly colonized sites, high levels of gene flow from established populations can provide the genetic material necessary to exploit new habitats (Sakai et al, 2001). Additionally, gene flow between populations derived from different source areas can increase overall genetic diversity, perhaps leading to the formation of invasive genotypes (Ellstrand and Schierenbeck, 2000). Characterizing levels of gene flow in exotic species, therefore, should increase our understanding of why certain species become successful invaders.
The development of genetic markers has greatly increased our ability to accurately estimate population-genetic measures such as gene flow. The easiest, and most widely used, approach to estimating gene flow uses population-genetic models, especially that of Wright (1951), to describe the level of gene flow necessary to produce the observed distribution of genetic diversity among a set of populations. This approach relies on the assumption that the homogenizing action of migration balances genetic divergence among populations by genetic drift (Wright, 1951; Slatkin, 1993). Consequently, the genetic structure seen among populations will be primarily affected by this equilibrium between migration and genetic drift.
Among introduced species, genetic structure is often heavily influenced by human activities (Godt and Hamrick, 1991; Sakai et al, 2001). Since many generations may pass before these historical footprints disappear, and for migration–drift equilibrium to be achieved (Slatkin, 1993), indirect measures of gene flow for recently introduced species may not be appropriate.
Direct measures provide an alternative method for estimating contemporary levels of gene flow. In one approach, paternity analysis, a direct estimate of gene flow via pollen, is obtained by determining the pollen donor (ie, the ‘father’) of a large number of seeds from a population. Although requiring considerable field and lab work, this approach has fewer assumptions than indirect methods, and is the only reliable means of estimating contemporary gene flow for species not in migration–drift equilibrium. Gene flow, studied directly, is extensive in many tree species, with pollen often traveling more than a kilometer (Hamrick and Nason, 2000).
In this study, we estimate gene flow into isolated populations of the introduced tree, Albizia lebbeck, using both indirect and direct measures. We determine whether populations are in migration–drift equilibrium, and therefore amenable to indirect estimates of gene flow. Using direct measures, we determine the proportion of pollinations that involve immigrant pollen, and whether population size and distance to nearest-neighbor influence rates of pollen flow. We also examine patterns of genetic diversity for clues to the success of A. lebbeck as an invasive species.
Materials and methods
Study populations were located near the tip of southwestern Puerto Rico near the Sierra Bermeja mountain range (Figure 2). The mostly flat topography is crossed by several low mountain ranges (200–225 m in elevation). The region is classified as subtropical dry forest (sensu Holdridge, 1967), receiving less than 1000 mm of rain per year. Forest vegetation is scattered across a landscape that has been largely cleared for agriculture and cattle pastures. Exotic grasses cover much of the site, as is typical for tropical areas cleared of forest (Daehler, 1998; Cabin et al, 2001). Especially common is African Guinea Grass (Panicum maximum Jacq.; J Schwagerl, personal communication), which can burn in the dry season, although fires are not widespread.
A. lebbeck (L.) Benth. (Fabaceae, Mimosoideae), a tree native to tropical Asia (Little and Wadsworth, 1964), was introduced into the British Caribbean islands in 1782 (Howard, 1954), and has been in Puerto Rico at least since the beginning of the 20th century (Urban, 1905). This shade tree, primarily used ornamentally (Little and Wadsworth, 1964), has escaped cultivation and is now found in a variety of natural and disturbed Puerto Rican habitats.
Traits that A. lebbeck possesses, which are typical of invasive tree species, include perfect flowers, generalist pollinators, retention of fruit on the tree (Reichard and Hamilton, 1997), nitrogen fixation (Daehler, 1998), use of pioneer habitat, high seed production, high growth rates (Kolar and Lodge, 2001), and multiseeded fruits (Sakai et al, 2001). These traits presumably have helped A. lebbeck invade this extensively grazed island landscape, an environment particularly susceptible to invasion by non-native species (Lonsdale, 1999).
A variety of generalist pollinators, predominantly bees and butterflies, visit the perfect flowers of A. lebbeck (B Dunphy, personal observation). The long (10–20 cm) light pods, which contain between six and nine seeds, can be blown several hundred meters across open habitats with a strong wind, although a large number typically fall beneath the maternal tree (B Dunphy, personal observation).
Indirect estimates of gene flow
Indirect gene flow estimates were based on 10 populations of 24 individuals each (Figure 2). All populations were taken from low-elevation open fields with minimal forest cover. The minimum distance separating two populations was 0.5 km; the maximum was 22.3 km. A minimum of 20 cm2 of leaf tissue was collected from each tree, and shipped on ice to the laboratory at the University of Georgia within 48 h of collection. Leaves were crushed in a potassium phosphate extraction buffer (Mitton et al, 1979) and resulting extracts were stored on filter paper wicks at −70°C.
Horizontal starch gel electrophoresis was used to assay allozyme diversity. Five gel buffer system gel–electrode combinations and 13 enzyme stains resolved 21 loci on 11.0% starch gels. Gel and electrode buffer recipes followed Soltis et al (1983). Enzymes stained (and loci resolved) on System 4 (a Tris/citrate gel and buffer) were isocitrate dehydrogenase (Idh), diaphorase (Dia), and phosphoglucoisomerase (Pgi-1 and Pgi-2). System 7 (a Tris-citric acid buffer gel and lithium hydroxide and boric acid tray buffer) was stained for amino-acid transferase (Aat). A modified System 8 (a discontinuous LiOH system) resolved leucine aminopeptidase (Lap) and glutamate dehydrogenase (Gdh). System 11 (a histidine gel and citric acid buffer) was stained for malate dehydrogenase (Mdh-1, Mdh-2, and Mdh-3), 6-phosphogluconate dehydrogenase (6Pgd-1 and 6Pgd-2), UTP-glucose-1-phosphate (Ugpp-1 and Ugpp-2), shikimate dehydrogenase (Skdh), and fructose-1,6-di-phosphatase (F16-1 and F16-2). System 6 (a Tris-citric acid buffer gel and sodium hydroxide and boric acid tray buffer) resolved fluorescent esterase (Fe-1 and Fe-2) and triose-phosphate isomerase (Tpi-1 and Tpi-2). Stain recipes were modified from Soltis et al (1983) except for diaphorase (Cheliak and Pitel, 1984) and UTP-glucose-1-phosphate (Manchenko, 1994). For enzymes with more than one locus, isozymes were numbered sequentially, with the lowest number assigned to the most anodal banding zone.
Standard measures of genetic variation were calculated for each population and for the species by pooling data for all the individuals analyzed. Genetic diversity parameters estimate percentage of polymorphic loci (P), the mean number of alleles per locus (A) and per polymorphic locus (AP), the effective number of alleles per locus (Ae), observed heterozygosity (Ho), and expected heterozygosity (He). Genetic structure was analyzed using Weir and Cockerham's (1984) estimators of F-statistics, calculated with FSTAT (Goudet, 1995). Fixation indices (FIS) for individual populations were calculated with Lynsprog, developed by MD Loveless and A Schnabel, which provides χ2 values for the indices.
Indirect gene flow estimates, in terms of the effective number of immigrants per generation (Nem), were made as follows (Wright, 1951):

where Ne is the effective number of individuals in the population, m is the immigration rate, and FST is the proportion of total genetic diversity due to differences among populations (Wright, 1951). A key assumption is that populations are in migration–drift equilibrium, which is indicated by a statistically significant positive relationship between pairwise estimates of FST/(1−FST) and the geographic distance separating the pair (Rousset, 1997).
Direct measures of gene flow
Four stands of trees were chosen from the same geographic location as the populations used for indirect estimates of gene flow (Figure 2). Distances from each stand to the nearest conspecific tree ranged from 60 to 450 m. From all stands, a total of 651 pods were analyzed.
From each pod, only a single seed was chosen for analysis. This minimized pseudoreplication due to correlated mating, a situation common to mimosoid legumes in which a single pollen donor sires all seeds within a fruit. Seeds chosen for analysis were soaked for 15 min in concentrated sulfuric acid, rinsed with water, and planted at the Plant Biology Greenhouses at the University of Georgia in Athens, GA. Germination rates exceeded 90% and most seedlings emerged within 2–3 weeks. If a seed did not germinate, or if a seedling died, a replacement was taken from the same pod. Once true leaves appeared, roughly 20 cm2 of leaf tissue was collected and processed for allozyme analyses as described above. A large number of seeds had exit holes created by the bruchid beetle, Merobruchus paquetae Kingsolver. These ‘prescarified’ seeds were directly planted into the soil, and germinated at about the same rate as the acid-scarified seeds without exit holes.
Multilocus estimates of outcrossing were computed with MLTR (Ritland, 2002). Paternity analysis was used to estimate directly gene flow levels. Genotypes of all trees from a seedling's stand that could potentially have sired the seed were compared against the seedling's genotype. An initial estimate of gene flow was made by determining the percentage of seedlings that could not have been sired by potential fathers within the stand (ie, apparent gene flow). Allozyme trials yielded an exclusion probability of 0.91 for A. lebbeck, indicating that for a given seed, 91% of potential sires will be correctly excluded (Chakroborty et al, 1988). Since a pollen gamete produced outside of the population could be indistinguishable from one created by a sire within the population, the initial estimate is an underestimate of the true gene flow rate. The estimation of the ‘cryptic’ gene flow rate is essential to obtain accurate and unbiased estimates of the total rate of gene flow.
To estimate cryptic gene flow, we used the program GFLOW (available from BKD), which employs a slight modification of the technique developed by Devlin and Ellstrand (1990). This technique uses maximum likelihood to determine the total (apparent+cryptic) gene flow rate that would most likely yield the observed apparent gene flow rate (Devlin and Ellstrand, 1990). Average allele frequencies from the 10 populations used to estimate genetic diversity provided the ‘pollen pool’ frequencies necessary for this analysis.
Once total gene flow rates were available for individual trees, an average gene flow rate for the stand was calculated. This was calculated as the average, weighted by seed number, for all the trees from the stand for which gene flow was estimated.
Results
Diversity statistics and population structure
Of the 21 loci surveyed, 14 (66.7%) were polymorphic in at least one of the populations sampled (Table 1). At the species level, there were 3.29 alleles per polymorphic locus (APs), the effective number of alleles (Aes) was 1.58, and expected heterozygosity (Hes) was 0.266. At the population level, on average, 48.9% of the loci were polymorphic, with R100S and R306 having the highest values (66.7 and 61.9%, respectively). Refuge, Vivarium, and Tinaja shared the lowest value, 36.8%. There were 1.36 effective alleles per locus (range 1.24–1.62), and 2.37 alleles at polymorphic loci (range 2.14–2.64). Genetic diversity (He) ranged from 0.151 (Parguera) to 0.293 (R100S), with a mean of 0.189 (Table 1). Highest diversity values were for the three populations north of Rte. 101 (Figure 1).
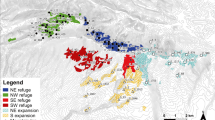
Study populations of A. lebbeck in southwestern Puerto Rico. Populations of adult trees used in indirect estimates of gene flow are numbered 1–10. The four stands of trees used for direct measures of gene flow are lettered (A–D). Stippled regions are low mountains (maximum height 200–225 m). Diagonal lines show the location of the Cabo Rojo National Wildlife Refuge.
Estimates of Wright's F-statistics and their 95% confidence limits were FIS=0.205 (0.078–0.339), FIT=0.414 (0.291–0.537), and FST=0.263 (0.184–0.359). A χ2 analysis for allele frequency heterogeneity among populations yielded highly significant (P<0.001) differences among populations for all polymorphic loci except Idh (P<0.025). A deficit of heterozygotes, indicated by positive FIS values (Table 2), was found for 10 loci, while four loci had an excess of heterozygotes. Across loci, there was a significant deficit of heterozygotes (χ2=145.54; P<0.001). Very high FIS values were found for Fe-1, Lap, Skdh, and 6pgd-2, loci with novel genotypes in the two northern populations R100S and R306. The very low FIS observed for Tpi-1, a locus monomorphic for most populations, was due to an excess of heterozygotes in R100S (FIS=−0.345) and especially R306, where the excess was statistically significant (FIS=−0.469, χ2=5.27, P<0.05). Of 96 fixation indices calculated, 29 (30%) were significantly different from zero (P<0.05). A total of 27 of these 29 indices were positive, indicating a deficit of heterozygotes. All populations except Refuge had at least one fixation index significantly different from zero. R306 had the most, with eight loci, followed by R100S with five loci.
Direct measures of gene flow
The multilocus estimate of outcrossing (tm) was 0.979 (SE=0.052). Gene flow rates were high, with values of 44% for JH, 78% for Manola, 100% for R100, and 81% for Tex. In Manola and Tex, stands where seeds were available from multiple maternal trees, variation within stands was greater for the former, which had trees with gene flow estimates as low as 65% and as high as 100%. The two values for Tex were 80 and 83% (Table 3).
An estimate of effective population size (Ne) was made for Manola and Tex, the two populations for which fruit production was recorded. Ne was calculated as Ne=1/∑((ci+pi)/2)2, where ci and pi are the relative female and male reproductive contribution of the ith individual, respectively (Crow and Denniston, 1988). Values of Ne were 7.26 for Manola and 4.33 for Tex. Combining Ne with the gene flow rates estimated above yields estimates of Nem of 2.83 (Manola) and 1.75 (Tex), although low effective population sizes limited maximum Nem to 3.63 and 2.17, respectively.
Indirect measures of gene flow
The percent of the variation distributed among populations (FST) was 26.3% (Table 2). Considered pairwise, only two FST values were between 0.1 and 0.2; the remaining values were either higher or lower. At the low end, 28.9% of FST values were below 5%. Two population pairs (Refuge and Texaco, Texaco and R100N) had values below 1%, despite a minimum of 3.4 km separating each pair. At the other extreme, 33.3% of pairs had FST values above 30% (13.3% were above 40%). All pairwise FST values above 0.300 involved either R100S or R306, two of the three populations north of R101. With these two populations removed, overall FST drops to 0.120.
No apparent relationship existed between genetic (Nei, 1972) and geographic distances. The genetic distance between the two most geographically distant populations (Texaco and Parguera) was 0.0187, a value less than 77.8% of other pairwise genetic distance values. Plotting FST/(1−FST) against interpopulation distances (Figure 2) yielded a nonsignificant, slightly negative slope, rather than the positive slope expected under migration–drift equilibrium (R2=0.039, P>0.10). Removing populations R100S and R306 increased the total variation in genetic distance explained to 6.6%, yet the relationship was still not statistically significant.
Unique alleles in northern populations
Two populations north of Rte. 101 (Figure 1), R100S and R306, differed substantially from the other populations, with an average genetic distance of 0.377 (SD 0.057); among the remaining populations, the average was 0.094 (SD 0.084). For four of the 14 polymorphic loci (Aat, Lap, Skdh, and 6pgd-2), the most common alleles in R100S and R306 were either absent in the other populations, or at frequencies below 15%. Some loci (Dia, 6pgd-2, Lap, and Fe-2) had alleles in R100S that were entirely missing from the other populations; the most common allele at Fe-2 in R100S was found only in that population. Additionally, three loci (Tpi-1, Skdh, and 6pgd-2) that were nearly monomorphic in the other populations were polymorphic in R100S and R306.
Individuals within R100S and R306 tended to be either free of these rare ‘northern population’ alleles, or to have them at multiple loci. Specifically, individuals with at least one ‘northern’ allele averaged 6.6 (95% CI 6.10–7.05) loci with ‘northern’ alleles. A lack of mixing of ‘northern’ with common alleles led to a substantial lack of heterozygotes, compared to random mating expectations in both R100S (FIS=0.416) and R306 (FIS=0.438). Three trees homozygous at several loci for ‘northern alleles’ also appear in R100N, the third population north of Rte. 101, although this was not enough to increase its genetic distance values with the seven southern populations.
Within individual loci, ‘northern’ alleles did not often mix with common alleles. A rare allele at Pgi-2 and one at Aat, in particular, were only found in homozygotes, despite the presence of common alleles within the same population. Similarly, two ‘northern’ alleles in 6pgd-2 formed heterozygotes only with the ‘3’ allele, an allele mostly restricted to R100S and R306. There were similar findings with Fe-2, Dia, Lap, and Skdh, although for each locus, there were a few individuals that were heterozygous for both ‘northern’ and common alleles. ‘Northern’ alleles show up in very limited amounts in populations to the south: one tree in Tex was homozygous for ‘northern’ alleles at Pgi-2, Aat, Skdh, and 6pgd-2; a second tree in Tex was heterozygous for a ‘northern’ allele at 6pgd-2; and two trees from JFinca were homozygous for a ‘northern’ allele at Lap.
Not surprisingly, R100S and R306 grouped together in a UPGMA phenogram, well separated from the other eight populations (Figure 3). Six populations were grouped in a close cluster. There did not appear to be any correlation between relationships in the phenogram and geographic proximity. For instance, Refuge and Vivarium are close geographically (ca 0.50 km), but did not group together in the phenogram.
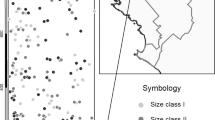
UPGMA phenogram of Nei's genetic distances (Nei, 1972) among 10 introduced A. lebbeck populations in southwestern Puerto Rico.
Populations R100S and R306 exerted a strong influence over the population statistics. With the exception of P, which was unchanged, all values decreased with removal of these two populations from the analysis: AP decreased from 3.29 to 2.79, A decreased from 2.52 to 2.19, Ae decreased from 1.58 to 1.39, and He changed from 0.266 to 0.206.
Discussion
Patterns of genetic diversity
Genetic diversity statistics for A. lebbeck were comparable or exceeded those reported for other outcrossing tree species by Hamrick and Godt (1996). The percent of polymorphic loci was similar (66.7 vs 65.5%), while gene diversity (He) was substantially higher (0.266 vs 0.180). Over a quarter of this diversity is due to variation among populations (FST=0.263 vs 0.094), a value normally seen for species with some degree of selfing. Since A. lebbeck is almost entirely outcrossing, the high FST values are likely due to historical population founding events, rather than a direct consequence of its mating system. Over time, the high rates of gene flow should homogenize populations, decreasing FST. Based upon direct estimates of Nem, FST should reach an equilibrium value of 0.097 (Manola) or 0.125 (Tex), values more consistent for species with similar life history traits (Hamrick and Godt, 1996).
Gene flow rates
Direct estimates of gene flow were high for all four stands. No relationship was seen between either nearest-neighbor distance or stand size and gene flow rates, suggesting that pollen readily moves across the distances represented in this study, and that reproduction and spread of A. lebbeck should not be pollen limited. The high gene flow rates reported for A. lebbeck are consistent with results for other tree species. Lone trees of the congeneric species, Albizia julibrissin, isolated by at least 1 km, produced large crops of outcrossed seeds (Godt and Hamrick, 1997). Mean pollen dispersal distance for Dinizia excelsa, an insect-pollinated Amazonian tree, was estimated to be 1509 m in open pasture (maximum 3.2 km) and 212 m in undisturbed forest (Dick, 2001; Dick et al, 2003). Pollen flowed an average of 191 m, with a maximum observed distance of 663 m in Neobalanocarpus heimii, a tropical emergent tree from Malaysia (Konuma et al, 2000). Three species of Ficus from Barro Colorado Island in Panama had gene flow levels greater than 90% and small populations of Spondias mombin on islands in Lake Gatun had pollen flow rates of 60–100% over distances as great as 1 km (Nason and Hamrick, 1997). In fragmented Honduran populations of Swietenia humilis, between 24 and 100% of the pollen traveled more than 900 m (White et al, 2002); an isolated tree received 71% of its pollen from over 4.5 km. Thus, long-distance pollen movement for insect-pollinated species such as A. lebbeck appears to be the rule rather than the exception.
The ready availability of suitable pollen is reflected in the consistent production of large numbers of fruits even in years when fruit production in other species was low (B Dunphy, personal observation). A high gene flow rate acts therefore not only to exchange genes among populations, but also to ensure a large fruit crop, especially in self-incompatible species such as A. lebbeck, providing seeds for population growth and the colonization of new sites.
A. lebbeck pods often remain on the branch for months, a trait promoting success as an invasive species (Reichard and Hamilton, 1997). Wind speeds near the study site average 11.2 mph, but wind gusts between 28.4 and 34.2 mph occur almost every month of the year (NOAA, 1995). Thus, pods are available when these high winds occur, allowing distant dispersal of seeds.
Comparison of direct and indirect measures of gene flow
Indirect and direct estimates of gene flow lead to slightly different predictions. According to population genetic theory (Wright, 1931), the indirect estimate, Nm=0.69, predicts that populations should grow more dissimilar over time. With the potentially problematic genotypes from populations R100S and R306 removed, Nm increased to 1.28, slightly within the key range of 1–4, where gene flow will slowly homogenize populations over time. Direct estimates, Nem=1.75 and 3.04, fall within this range as well. The apparent absence of migration–drift equilibrium for this invasive species indicates that the direct estimates are more reliable.
The presence of demographic forces disrupting migration–drift equilibrium is indicated by the simultaneous occurrence of loci with a deficit of heterozygotes and loci with an excess of heterozygotes (positive and negative FIS values, respectively; Table 2). This would not normally be seen in established populations, where all loci are exposed to the same demographic forces.
The lack of migration–drift equilibrium may be due to the relatively recent introduction of A. lebbeck into southwestern Puerto Rico, as the presence of unusual alleles in R100S and R306 suggests. Over time, as these alleles move through the landscape, it is unclear whether migration–drift equilibrium will be achieved. In the relationship between FST and geographic distance (Figure 2), there was a cloud of points, representing a range of geographic distances, with low FST values. These points all involve population pairs with at least one of the pair from populations Tinaja, Parguera, R100N, or R116W. These low FST values could be due to recent establishment of these populations from a common source, or from high gene flow between populations. Since the four populations are along roadsides (Parguera, R100N, and R116W) or in a fallow field (Tinaja), it is unlikely that they were deliberately planted. High gene flow, both via pollen, as demonstrated in the current study, and via seed, seems more likely. If this is the case, then the low FST values indicate that these must be older, more established populations for which enough time has passed for gene flow to homogenize differences in allele frequencies due to the original founder effects.
With the cloud of points removed, the relationship between FST and geographic distance becomes positive. The relationship may be accidental, however, since JFinca, which is the most remote population in this group of four, also tends to have high FST values with the other nine populations, including its nearest neighbors. This is reflected in the UPGMA phenogram (Figure 3), where JFinca was separated from the other populations. In addition, a pattern of isolation by distance tends to appear first among nearest neighbors (Slatkin, 1993). On average, however, an A. lebbeck population is more closely related to 4.3 other populations than to its nearest neighbor, so there is no evidence that isolation by distance has developed on the spatial scale examined.
Migrant genotypes in R100S and R306
Two of the three populations (R100N and R306) north of Rte. 101 were characterized by genotypes rare or absent from the other populations. These genotypes appear to have recently arrived in the area, since individuals in these two populations tended to have multilocus genotypes consisting of either ‘northern alleles’ or alleles common to the other eight populations. This disequilibrium should begin to break down within a generation of random mating, indicating that individuals with ‘northern’ genotypes are either newly established or that random mating is not occurring, perhaps due to differences in flowering phenology. A third possibility, the presence of a reproductively isolated cryptic species, is precluded by the presence of heterozygotes, albeit uncommon, that combine ‘northern’ with common alleles, demonstrating that mating can occur. Additionally, trees in R100S and R306 with ‘northern’ alleles had several loci with alleles common in the other eight populations.
The contribution of multiple source populations would explain the high genetic diversity values (0.266). Removing the two populations dominated by ‘northern’ alleles lowered He to 0.206, a value closer to the average (0.180) reported for species with similar life history characteristics (Hamrick and Godt, 1996). Introduced populations, derived from several source populations, often are more diverse than any single source population (Barrett and Husband, 1990). Kudzu (Pueraria lobata), for instance, had high levels of genetic diversity (0.290), presumably due to historically documented cases of multiple introductions (Pappert et al, 2000).
A. lebbeck's success as an invasive exotic
A. lebbeck has a number of reproductive characteristics that make it a successful invader, including even and large fruit production, large amounts of seed, and fruits that remain attached to the tree for long periods (Sakai et al, 2001). For many species, there is a strong genetic component to invasiveness, indicated by the presence of both invasive and noninvasive conspecific populations (eg, Sakai et al, 2001). Extensive gene flow may produce novel combinations of alleles, allowing invasive genotypes to form (Ellstrand and Schierenbeck, 2000), and providing the genetic diversity needed to adapt to a wide range of habitats during range expansion (Antonovics, 1976; Crawley, 1986; Hengeveld, 1990). In introduced populations of reed canary grass (Phalaris arundinacea), for instance, one genotype grew faster under strong competition, while a different genotype had the fastest growth rate with minimal competition (Morrison and Molofsky, 1999). The maintenance of genetic diversity may therefore be an important factor allowing this grass to invade successfully new habitats with varying levels of competition.
With few colonists, especially if all are drawn from the same source, genetic bottlenecks can lead to inbreeding depression (Ellstrand and Elam, 1993; Newman and Pilson, 1997), and lower genetic diversity, adversely affecting the populations' ability to adapt to an invaded habitat (Sakai et al, 2001). This may be why invasiveness often develops only after multiple introductions, especially by hybridization between individuals from source populations well isolated in their native range (eg, Pappert et al, 2000). In self-incompatible species, such as A. lebbeck, multiple invasions may also introduce a large number of self-incompatibility alleles, thus overcoming a potential barrier to population expansion.
The frequent occurrence of bruchid beetles, which do not appear to harm the embryo, may actually increase the ecological success of A. lebbeck. Species with germination requirements met over a range of environmental conditions tend to be more successful (Baker, 1974), especially those that lack a pregermination requirement (Reichard and Hamilton, 1997). Two types of seeds covering a broad range of possible environmental conditions are effectively produced in A. lebbeck: seeds with exit holes, capable of immediate germination; and intact seeds, requiring scarification, that can wait in the seed bank longer.
In conclusion, pollen appears to move freely over the distances examined. The A. lebbeck populations in southwestern Puerto Rico are not in migration–drift equilibrium, so indirect measures of gene flow cannot be used reliably. Although considerable genetic structure exists, this is probably due to the recent arrival of foreign genotypes from north of the study site. Migration–drift equilibrium may only appear over larger geographic distances. Because trees are not pollen limited, the capacity for seed dispersal, and the ability to deal with local environmental regimes, should dictate the rate at which this species spreads in the area studied. High gene flow rates maintain high genetic diversity, which may aid continued spread of the species by providing the environment-specific genotypes needed for adaptation to a varied set of conditions.
References
Antonovics J (1976). The nature of limits to natural selection. Ann Mo Bot Gard 63: 224–247.
Baker HG (1965). Characteristics and modes of origin of weeds. In: Baker HG, Stebbins GL (eds) The Genetics of Colonizing Species. Academic Press: New York. pp 147–172.
Baker HG (1974). The evolution of weeds. Ann Rev Ecol Syst 5: 1–24.
Barrett SCH, Husband BC (1990). The genetics of plant migration and colonization. In: Brown AHD, Clegg MT, Kahler AL, Weir BS (eds) Plant Population Genetics, Breeding and Genetic Resources. Sinauer: Sunderland, MA. pp 254–277.
Cabin RJ, Weller SG, Lorence DH, Flynn TW, Sakai AK, Sandquist D et al (2001). Effects of long-term ungulate exclusion and recent alien species control on the preservation and restoration of a Hawaiian tropical dry forest. Conserv Biol 14: 439–453.
Chakroborty P, Meagher TR, Smouse PE (1988). Parentage analysis with genetic markers in natural populations. 1. Paternity exclusion and expected proportions of offspring with unambiguous paternity. Genetics 118: 527–536.
Cheliak WM, Pitel JA (1984). Techniques for Starch Gel Electrophoresis of Enzymes for Forest Tree Species. Petawawa National Forestry Institute, Information Report P1-X-42. Canadian Forestry Service, Agriculture: Chalk River, Ontario.
Crawley MJ (1986). The population biology of invaders. Philos Trans R Soc London B 314: 711–729.
Crow JF, Denniston C (1988). Inbreeding and variance effective population numbers. Evolution 42: 482–495.
Daehler CC (1998). The taxonomic distribution of invasive angiosperm plants: ecological insights and comparison to agricultural weeds. Biol Conserv 84: 167–180.
Devlin B, Ellstrand NC (1990). The development and application of a refined method for estimating gene flow from angiosperm paternity analysis. Evolution 44: 248–259.
Dick CW (2001). Genetic rescue of remnant tropical trees by an alien pollinator. Proc R Soc London B 268: 2391–2396.
Dick CW, Etchelecu G, Austerlitz F (2003). Pollen dispersal of tropical trees (Dinizia excelsa: Fabaceae) by native insects and African honeybees in pristine and fragmented Amazonian rainforest. Mol Ecol 12: 753–764.
Ellstrand NC, Elam DR (1993). Population genetic consequences of small population size: implications for plant conservation. Ann Rev Ecol Syst 24: 217–242.
Ellstrand NC, Schierenbeck KA (2000). Hybridization as a stimulus for the evolution of invasiveness in plants? Proc Natl Acad Sci USA 97: 7043–7050.
Godt MJW, Hamrick JL (1991). Genetic variation in Lathyrus latifolius (Leguminosae). Am J Bot 78: 1163–1171.
Godt MJW, Hamrick JL (1997). Estimation of mating system parameters of Albizia julibrissin (Fabaceae). For Genet 4: 217–221.
Goudet J (1995). FSTAT (ver. 1.2): a computer program to calculate F-statistics. J Hered 86: 485–486.
Hamrick JL, Godt MJW (1996). Effects of life history traits on genetic diversity in plant species. Philos Trans R Soc London B 351: 1291–1298.
Hamrick JL, Nason JD (2000). Gene flow in forest trees. In: Boyle TJB, Young A, Boshier D (eds) Forest Conservation Genetics: Principles and Practice. CIFOR and CSIRO: Australia. pp 81–90.
Hengeveld R (1990). Dynamic Biogeography. Cambridge Univ. Press: London.
Holdridge LR (1967). Life Zone Ecology. Tropical Science Center: San José, Costa Rica.
Howard RA (1954). A history of the Botanic Garden of St. Vincent, British West Indies. Geogr Rev 44: 381–393.
Kolar C, Lodge DM (2001). Progress in invasion biology: predicting invaders. Trends Ecol Evol 16: 199–204.
Konuma A, Tsumura Y, Lee CT, Lee SL, Okuda T (2000). Estimation of gene flow in the tropical-rainforest tree Neobalanocarpus heimii (Dipterocarpaceae), inferred from paternity analysis. Mol Ecol 9: 1843–1852.
Little EL, Wadsworth FH (1964). Common Trees of Puerto Rico and the Virgin Islands. USDA Forest Service, Agr. Handbook 249.
Lonsdale WM (1999). Global patterns of plant invasions and the concept of invasibility. Ecology 80: 1522–1536.
Manchenko GP (1994). Handbook of Detection of Enzymes on Electrophoretic Gels. CRC Press: Ann Arbor, MI.
Mitton JB, Linhart YB, Sturgeon KB, Hamrick JL (1979). Allozyme polymorphisms detected in mature needle tissue of ponderosa pine. J Hered 70: 86–89.
Morrison SL, Molofsky J (1999). Environmental and genetic effects on the early survival and growth of the invasive grass Phalaris arundinacea. Can J Bot 77: 1447–1453.
Nason JD, Hamrick JL (1997). Reproductive and genetic consequences of forest fragmentation: two case studies of Neotropical canopy trees. J Hered 88: 264–276.
National Oceanic and Atmospheric Administration (NOAA) (1995). Wind Frequency Statistics and Monthly Peak Gusts for Ponce, Puerto Rico (1972–1990). Unpublished data from the Southeast Regional Climate Center, Columbia, South Carolina.
Nei M (1972). Genetic distance between populations. Am Nat 106: 283–292.
Newman D, Pilson D (1997). Increased probability of extinction due to decreased genetic effective population size: experimental populations of Clarkia pulchella. Evolution 51: 354–362.
Pappert RA, Hamrick JL, Donovan LA (2000). Genetic variation in Pueraria lobata (Fabaceae), an introduced, clonal, invasive plant of the Southeastern United States. Am J Bot 87: 1240–1245.
Reichard SH, Hamilton CW (1997). Predicting invasions of woody plants introduced into North America. Conserv Biol 11: 193–203.
Ritland K (2002). Extensions of models for the estimation of mating systems using n independent loci. Heredity 88: 221–228.
Rousset F (1997). Genetic differentiation and estimation of gene flow from F-statistics under isolation by distance. Genetics 145: 1219–1228.
Sakai A, Allendorf FW, Holt JS, Lodge DM, Molofsky J, With KA et al (2001). The population biology of invasive species. Ann Rev Ecol Syst 32: 305–332.
Slatkin M (1993). Isolation by distance in equilibrium and nonequilibrium populations. Evolution 47: 264–279.
Soltis DE, Haufler CH, Darrow DC, Gastony GJ (1983). Starch gel electrophoresis of ferns: a compilation of grinding buffers, gel and electrode buffers, and staining schedules. Am Fern J 73: 9–27.
Urban I (1905). Symbolae Antillanae seu Fundamenta Florae Indiae Occidentalis. In: Flora Portoricensis. Fratres Borntraeger: Leipzig. Vol 4, pp 193–352.
Weir BS, Cockerham CC (1984). Estimating F-statistics for the analysis of population structure. Evolution 38: 1358–1370.
White GM, Boshier DH, Powell W (2002). Increased pollen flow counteracts fragmentation in a tropical dry forest: an example from Swietenia humilis Zuccarini. Proc Natl Acad Sci USA 99: 2038–2042.
Wright S (1931). Evolution in Mendelian populations. Genetics 16: 97–159.
Wright S (1951). The genetical structure of populations. Ann Eugenics 15: 323–354.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Dunphy, B., Hamrick, J. Gene flow among established Puerto Rican populations of the exotic tree species, Albizia lebbeck. Heredity 94, 418–425 (2005). https://doi.org/10.1038/sj.hdy.6800622
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/sj.hdy.6800622