Abstract
A wide range of animals have been reported to show kin-biased behaviours, such as reduced aggressiveness and increased food sharing among relatives. However, less is known about whether wild animals also associate with relatives under natural conditions, which is a prerequisite to facilitate kin-biased behaviours and hence kin selection. We tested, by means of microsatellite polymorphism, correlations between pair-wise relatedness and pair-wise metric distance in wild brown trout (Salmo trutta L.) under natural conditions in two streams. Our data show that young-of-the-year as well as older trout found close together also had a higher genetic relatedness in one of the two streams, whereas no relationship was found in the other stream. Very few half and full siblings were found in the second stream and under these conditions it is unlikely that kin-biased behaviours will receive positive selection. We discuss the underlying mechanisms for the observed structure and we specifically address the issue of whether the grouping of related individuals could reflect dispersal from the same spawning redds, or if it reflects active association with relatives, possibly conferring kin-selected advantages.
Similar content being viewed by others
Introduction
The distribution of individuals on microgeographical scales will affect the possibilities for individuals to interact. If interacting individuals are genetically closely related, kin-biased behaviours may be favoured by kin selection. Kin selection theory predicts that animals behave differently towards related versus unrelated conspecifics. This has also been demonstrated empirically in a range of animals (reviews by Holmes and Sherman, 1983; Waldman, 1987, 1988). For instance, kin recognition and kin selection is regarded as highly important for the evolution of helper at the nest systems in many bird species (review by Komdeur and Hatchwell, 1999). Another example is the Belding's Ground Squirrel, Spermophilus beldingi (Merriam), females that give alarm calls more readily when in the vicinity of related individuals (Holmes and Sherman, 1983).
Kin-biased behaviours cannot, however, evolve unless the majority of these behaviours are directed towards related individuals. Hamilton (1964) suggested that proximity could serve as a cue for kin recognition without the individual actually recognising conspecifics as kin or non-kin. Limited or no dispersal can lead to predictable clustering of close relatives and hence, kin-biased behaviour may receive positive selection as the majority of interactions are among relatives. Kin-biased behaviours would, however, be subject to stronger selection if individuals could minimise misdirected kin-biased behaviours by being able to identify and assess the degree of genetic relatedness to other individuals. A variety of animals ranging from tunicates to humans are documented to discriminate between related and nonrelated conspecifics (reviewed by Holmes and Sherman, 1983; Waldman, 1987, 1988).
Many salmonid species, including brown trout Salmo trutta (L.), are territorial and defend their territories vigorously, at least during some stage of their development. These properties have made salmonids suitable for studies of kin recognition and kin-biased behaviour. Evidence of kin-biased aggressive behaviour and preference for water scented by siblings has been demonstrated in artificially bred sibling groups of juvenile Atlantic salmon S. salar (L.), Arctic charr Salvelinus alpinus (L.), brown trout and rainbow trout Oncorhynchus mykiss (Waldbaum), both under laboratory and seminatural conditions (reviews by Brown and Brown, 1996; Olsén, 1999). There is, however, less knowledge concerning the degree to which these salmonids also associate with related conspecifics in wild populations under natural conditions.
Some attempts have been made to study if salmonids show kin-biased distribution under natural conditions in relation to the degree of relatedness, but the results are equivocal. Mjølnerød et al (1999) found weak, but significant spatial association between related Atlantic salmon. Fontaine and Dodson (1999) found no evidence that juvenile Atlantic salmon siblings occupy adjacent territories. Carlsson and Carlsson (2002) suggested that sibling competition among juvenile brown trout might lead to nonrandom distribution of related fish, with related fish found further apart.
The distribution of individual salmonids with respect to relatedness is likely to be influenced by the age of the individuals. Young-of-the-year (YOY) salmonids hatch in close proximity to related individuals (eg Elliott, 1987). Limited dispersal from spawning redds could thus lead to high relatedness among neighbouring individuals (Hansen et al, 1997). The association of individuals from the same redds is, however, expected to decay with time due to stochastic movements induced by, for instance, predators, floods and droughts or directional movements caused by ontogenetic shifts and seasonal movements (reviewed by Gowan et al, 1994).
Mature male parr are found in many salmonid species and represent an alternative mating tactic (Gross, 1984). These males reach sexual maturity at young age stages and spawn with large migratory conspecifics. Intensified competition for matings and increased aggression are typically observed among mature male parr during spawning, and mature male parr are often attacked by larger conspecifics resulting in severe injuries or fatalities (Broberg et al, 2000). Therefore, ample opportunities for kin-biased behaviours among mature male parr should be present during spawning. By associating with, and showing high tolerance towards, related mature male parr, they could increase the chances that at least one or a few relatives spawn. By avoiding relatives, they could refrain from competing for mates with relatives. Hence, a nonrandom distribution of related mature male parr could be expected.
This study aims at detecting possible kin-biased distribution on microgeographical scales in wild brown trout under natural conditions. If limited dispersal rather than active association of kin is responsible for kin-biased distributions, we predict that it is most pronounced in YOY individuals and weaker in the older cohorts.
Materials and methods
Biological material
Trout were sampled by electrofishing in two small streams prior to spawning when mature fish are expected to assemble on or near the spawning grounds. A small tributary of the Findhu Glen Burn, southern Highlands, Scotland, which supports sea trout and resident trout, was sampled during late autumn 1999. YOY trout (N=93) were obtained by three successive removals of trout from a 50 m section, whereas the sample encompassing older fish (N=78) originated from a 215 m section (one removal). This tributary was later sampled by three successive removals to allow density estimates of older trout. Trout in this tributary most likely represent a fraction of a larger population as spawning also takes place in the main stream near the outlet of the tributary. A 125 m section of the Dollerup Møllebæk, Central Jutland, Denmark, that supports lake migratory trout and resident trout was sampled (N=103) in late autumn 2001. The stream was resampled a few weeks later to allow density estimates (two successive removals).
Both streams were mapped to the closest 10 cm and at the time of capture each individual caught in the mapped area was placed in a numbered 5-l plastic bag filled with water, and the corresponding number was marked on the map. In total, 89 YOY and 31 older trout were caught in the mapped area of the Findhu Glen Burn, and 103 trout were caught in the Dollerup Møllebæk mapped area. Individuals caught outside the mapped area, or for whom the position in the mapped area was uncertain, were included in the analyses of microsatellite variability and as background allele frequencies for relatedness analyses to increase the power of these analyses.
All caught fish were later anaesthetized in 2-phenoxy ethanol or benzokain, measured (total length) and sexed (external inspection). All mature males with total lengths less than 300 mm were considered to be male parr or small resident males. A small fin clip (tail fin) was sampled from all caught individuals and stored in 96% ethanol until analysed. Scales were sampled from all fish with lengths exceeding 80 mm in the Findhu Glen Burn sample. All fish were released alive after handling. The sample from the Findhu Glen Burn was divided into two groups, YOY trout (in the following abbreviated as FIN0+) and a second group encompassing older fish (in the following abbreviated as FINOLD); 41% of these fish were mature male parr or small mature resident males. The sample from Dollerup Møllebæk consisted of two groups: one consisted of trout aged 1+ or older (in the following abbreviated as DOLOLD, including 25% mature male parr and small mature resident males), caught from a 70 m section. The other group consisted of only mature male parr and small mature resident males from an additional 55 m (mature parr and small resident males from the additional 55 m in the Dollerup Møllebæk were pooled with the corresponding group from the first 70 m section, in the following abbreviated as DOLMAT).
Density estimates from electrofishing were calculated according to Bohlin et al (1989). In the case of Dollerup Møllebæk, all fish with total lengths exceeding 80 mm were included in the density estimate. These estimates ranged from 25.3 individuals/100 m2 (95% CI=21.3–40.8) in the FINOLD sample, to 71.7 (95% CI=67.5–75.8) in the Dollerup Møllebæk and 263.3 (95% CI=250.2–281.3) in the FIN0+ sample.
Microsatellites
Total genomic DNA was extracted from small fin-clips by proteinase K/lysis-buffer extraction (Laird et al, 1991) or by proteinase K/chelex extraction (Estoup et al, 1996). A total of 14 microsatellite loci were analysed: Ssa85, Ssa171, Ssa197 (O'Reilly et al, 1996), Ssa407UOS, Ssa408UOS, Ssa410UOS (Cairney et al, 2000), SsHaeIII14.20 (JL Goodier, unpublished data; Genebank accession number U10050), SSOSL311, SSOSL417 (Slettan et al, 1995), SSOSL438 (Slettan et al, 1996), T3-13 (Estoup et al, 1998), μ15, μ60 and μ73 (Estoup et al, 1993). More detailed information about PCR conditions will be provided upon request. Microsatellite length polymorphism was analysed on a Pharmacia ALF express automated sequencer, according to the manufacturer's recommendations.
Statistics
The two samples from Dollerup Møllebæk were pooled and analysed as one sample in all cases except when noted. Exact tests of deviations from Hardy–Weinberg equilibrium (Guo and Thompson, 1992) and calculations of observed (HO) and expected (HE) heterozygosity were performed using the GENEPOP 3.1b software package (Raymond and Rousset, 1995). A randomization test, based on permuting multilocus genotypes between samples (Goudet et al, 1996), was used to test for genetic differences between YOY trout and older trout in the Findhu Glen Burn sample and performed using the FSTAT 2.9.3.2 program (Goudet, 1995).
The KINSHIP 1.3.1 software (Queller and Goodnight, 1989) was used for estimations and simulations of relatedness between pair of individuals (R). This estimator is symmetric and range from −1 to 1. The estimator compares the degree of relatedness between individuals as compared to the average intrapopulation relatedness, which is forced to zero.
The pair-wise relatedness and pair-wise metric distances were sorted for each combination of individuals within each sample. The data could, however, not be statistically analysed in this form; as the number of individuals within each sample ranged from 31 to 89 individuals, the number of pair-wise comparisons would be far greater. For instance, the number of individuals in the FINOLD group was 31 and the number of pair-wise comparisons would be 465. This would present problems of pseudoreplication and the degrees of freedom in statistical analyses would be inflated. To overcome this problem, a separate regression of relatedness on metric distance was calculated for each individual, following methods described in Knight et al (1999). We calculated the regression of relatedness on metric distance for all the pair-wise relatedness and pair-wise metric distance estimates involving a particular individual. A negative regression slope would indicate that the individual was found nearby related conspecifics and the opposite for a positive slope.
The minimum number of full sibling families (ie where the intragroup relatedness should be close to 0.5) within each sample was estimated using the Harbinger–Smith algorithm available in the software SGP (Sibling Group Partition, Smith et al, 2001). This algorithm uses a Markov Chain Monte–Carlo approach to estimate the minimum number of full sibling groups in a sample without parental genotype information. The algorithm group individuals into families by maximizing the overall score of pair-wise likelihood ratios of either being full siblings or unrelated.
Although the algorithm in SGP reports the number of full-sibling groups in the sample, it was also of interest to know which proportion of individuals in each sample had close relatives (ie half and full siblings). However, estimation of the number of individuals with close relatives within each sample is problematic since the close relatives of one individual would not only be incorporated in the estimation for the specific individual but also for all its relatives. Hence, the estimates would be influenced by pseudoreplication. We therefore choose to estimate the number of individuals within each sample that had no close relatives. However, as this estimation would be affected by the difference in sample sizes, we randomly selected 31 individuals (ie the number of individuals in the smallest sample, FINOLD) from samples FIN0+, DOLOLD and DOLMAT, 100 times. The pair-wise relatedness between the 31 individuals was retrieved from the results of the relatedness analysis. We then calculated the proportion of individuals within each permutated sample that had no close relatives. The average proportion of individuals without close relatives from the 100 permutations were then compared between samples.
In all cases of multiple tests, significance levels were adjusted using the sequential Bonferroni technique (Rice, 1989).
Results
Microsatellite variability
The number of alleles per locus within samples ranged from three at locus μ60 observed in all samples, to 26 at locus Ssa410UOS in the Dollerup Møllebæk population (Table 1). Nine loci showed significant deviations from Hardy–Weinberg expectations in the FIN0+ sample (Table 1). Deviations from Hardy–Weinberg expectations were found at eight loci in the FINOLD sample, whereas only three loci showed significant deviations in the Dollerup Møllebæk sample (Table 1). The departures from Hardy–Weinberg expectations were not generally correlated with heterozygosity deficiency or heterozygosity excess (Table 1). Heterozygosity deficiencies were only observed in one sample, FINOLD, at two loci (SsHaeIII14.20 and μ15) but not in the FIN0+ sample (that originates from the same stream). Null alleles would generally be indicated by heterozygosity deficiency. Hence, if the heterozygosity deficiency were caused by null-alleles, we would expect both samples (FINOLD and FIN0+) to show heterozygosity deficiency at these loci. Heterozygosity excess was only observed at two loci (Ssa171 in the Dollerup Møllebæk sample and Ssa410OUS in the FIN0+ sample). Hence, we find it unlikely that technical artefacts such as null alleles or ‘ghost-bands’ were responsible for the deviations from Hardy–Weinberg expectations. Instead, we suggest that the assumptions of the Hardy–Weinberg principle have been violated, in particular the assumptions of infinite population size and/or nonrandom mating.
The sample from Findhu Glen Burn consisted of both YOY as well as older individuals and was divided into these two age groups (ie one group consisting of YOY fish and one group encompassing the remaining fish). These two groups were also genetically differentiated. Since both groups showed deviations from Hardy–Weinberg expectations, we tested genetic differentiation by using the test by Goudet et al (1996). This test does not assume that the samples are in Hardy–Weinberg equilibrium. The test showed that the samples were differentiated (P<0.001). Hence, when performing the relatedness analyses, we used the allele frequencies of FIN0+ as background for the relatedness estimates for FIN0+ and the frequencies of FINOLD as background for the estimates of relatedness in FINOLD.
Relatedness simulations
Based on the allele frequencies of each of the two samples from the Findhu Glen Burn and the sample from Dollerup Møllebæk, respectively, we simulated pair-wise relatedness for 10 000 unrelated individuals (R=0), 10 000 half siblings (R=0.25) and 10 000 full siblings (R=0.5). This was done for each sample to estimate the differences in power of the relatedness analyses and the expected distribution of relatedness estimates for unrelated, half and full siblings among samples. No differences in the distribution of relatedness estimates were found among the three simulated samples in any of the relatedness groups (Kruskal–Wallis test: P>0.05 for all three tests).
We used a conservative threshold value for detecting closely related individuals by defining close relatives at a relatedness estimate of 0.25 or higher (corresponding to half and full siblings). This threshold value will exclude the large majority of unrelated individuals but also exclude c. 50% of true half siblings and a low percentage of true full siblings from the close relatives group. The simulations showed that 3.6% of the simulated unrelated individuals exhibited relatedness values of 0.25 or higher in FIN0+, 3.9% in FINOLD and 4.2% in Dollerup Møllebæk, respectively. The proportion of half and full siblings with relatedness values below 0.25 was 50.0 and 4.3% in FIN0+ sample and 49.9 and 4.5% in the FINOLD, respectively. The corresponding values were 49.9 and 4.8%, respectively, in the Dollerup Møllebæk sample.
Relatedness analysis
The pair-wise relatedness of individuals in the FIN0+ (N=89), FINOLD (N=31) and Dollerup Møllebæk (N=103) samples were estimated using allele frequencies of the respective samples as background (note that the background allele frequencies of FIN0+ are based on 93 individuals, FINOLD allele frequencies are based on 78 individuals and in Dollerup Møllebæk the estimates are based on 103 individuals). However, after this estimation, the Dollerup Møllebæk sample was divided into two groups. The first group consisted of all caught trout older than YOY (N=69, DOLOLD) and a second group consisted only of mature male parr and small mature resident males (N=51, DOLMAT, ie mature males from the DOLOLD sample and mature males caught in the additional 55 m section).
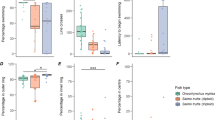
We analysed the average pair-wise metric distances between individuals according to their pair-wise relatedness. Two groups were created in each sample, where the first group encompassed all pairs of trout with pair-wise relatedness above 0.25 and the second group consisted of all pairs of trout with relatedness of less than 0.25. Significant differences in pair-wise metric distances among the two groups were found in both FIN0+ and FINOLD (Kruskal–Wallis test, P=0.000 for both tests) but not in DOLOLD or DOLMAT, with trout in the first group (R⩾0.25) found closer together than trout in the less related group (R<0.25) (Table 2).
A second, more refined, approach to study how pair-wise relatedness and pair-wise metric distances interacted was performed by analysing the average regression slopes of pair-wise relatedness on pair-wise metric distances within samples. These average slopes ranged from −0.00064 in FIN0+ to −0.00007 in the DOLMAT sample, respectively (Figure 1, Table 3). Both samples FIN0+ and FINOLD showed an average individual regression slope of pair-wise relatedness on pair-wise metric distances that was significantly different from zero (Student's one sample t-test, P=0.009 and P=0.006, respectively), indicating that related fish were found closer together than unrelated fish in these two samples. The average individual regression slopes of pair-wise relatedness on pair-wise metric distances were not significantly different from zero in the DOLOLD or DOLMAT (Student's one sample t-test, P=0.253 and P=0.551, respectively).
To detect differences in the number of full sibling families (ie R=0.5) between samples, the number of full sibling families within each sample was estimated with the SGP software. The analyses suggested that 11.7% of trout in the DOLMAT sample belonged to a full sibling group, whereas 14.5, 38.7 and 64.0% of trout belonged to full sibling family in the DOLOLD, FIN0+ and FINOLD samples, respectively.
However, we also wanted to estimate the proportion of trout in each sample not having any close relatives. The permutation analysis of proportion of trout without close relatives, R<0.25 (ie less related then half siblings), showed that 35% (SE=0.010) in the DOLOLD sample had no close relatives. The corresponding values were 28% (SE=0.008) in the DOLMAT sample followed by 11% (SE=0.006) in the FIN0+ sample. Significant differences were found between DOLOLD and DOLMAT, DOLOLD and FIN0+ as well as between DOLMAT and FIN0+ (Mann–Whitney U-test: P<0.000 for all three tests). The proportion of individuals without close relatives in the FINOLD sample was 13%.
Discussion
We present data showing significant differences in metric distance between highly related trout (R⩾0.25) as compared to weakly or unrelated trout (R<0.25), with highly related trout found closer together in the Findhu Glen Burn. Moreover, the average individual regression slope of relatedness on metric distance was weak but significantly lower than zero in both YOY and older trout from the Findhu Glen Burn, indicating that related trout were found in closer proximity than expected by chance. If redd location was the only factor leading to kin-biased distribution, we would expect that older trout in the Findhu Glen Burn showed a weaker kin-biased distribution as compared to YOY trout. As this was not the case, we suggest that kin-selected advantages were at least partly responsible for the kin-biased distribution observed in the Findhu Glen Burn.
Even though there should be ample opportunities for kin-biased behaviours among mature male parr and small mature resident males in the Dollerup Møllebæk, we did not observe any correlation between relatedness and metric distance. Nor was there any correlation between relatedness and metric distance for older than YOY trout in this stream. Hence, it is unlikely that kin-biased behaviours play an important role in the Dollerup Møllebæk. We discuss the possible mechanisms responsible for the observed pattern in terms of dispersal from redds, the actual number of families present at the two localities and the possibility of kin recognition.
Mechanisms for nonrandom distribution
Trout hatch and emerge from redds in close proximity to relatives and as they disperse, passively (eg Elliott, 1987) or actively (eg Kalleberg, 1958), they might be more likely to acquire territories close to related trout (McCormick et al, 1998). Trout unable to acquire and control a territory are assumed to die (Elliott, 1994). This would lead to a pattern where the degree of relatedness would decrease with increasing metric distance between individuals. Hence, this pattern would reflect where each individual was spawned (Hansen et al, 1997).
Alternatively, kin-selected benefits may be responsible for kin-biased distribution. These benefits could consist of reduced aggressiveness towards related individuals and smaller territory sizes (reviews by Brown and Brown, 1996; Olsén, 1999) as well as increased feeding opportunities for subordinates when in close proximity with dominant kin (Griffiths and Armstrong, 2002). Hamilton (1964) suggested that kin-biased behaviours could evolve if the dispersal of relatives was predictable in space and time. However, the association we observed in the Findhu Glen Burn is probably too weak to allow for kin-biased behaviours as a large proportion of an individual's neighbours are only weakly related or unrelated. Hence, many of the kin-biased behaviours would be misdirected and unrelated individuals would gain fitness from the behaviours. However, experimental studies have demonstrated that many salmonids, including brown trout, have the ability to discriminate between related and nonrelated individuals (reviews by Brown and Brown, 1996; Olsén, 1999). Hence, the proportion of misdirected behaviours could be minimized, and kin-biased behaviours mediated through kin recognition could be subject to positive selection.
Other factors that can influence kin-biased distribution are kin competition and inbreeding avoidance. Closely related individuals share large parts of their genome and may have larger overlaps in resource utilization than less related individuals leading to more fierce competition among kin than among nonkin (Waldman, 1988). Association with highly related conspecifics could under these conditions lead to individuals reducing their inclusive fitness. The fiercest competition with related individuals could however be reduced by avoiding association with related individuals. Carlsson and Carlsson (2002) also reported that related juvenile brown trout were found further apart then expected if trout were randomly distributed.
Inbreeding depression has been well documented in the literature and many animals also actively avoid breeding with close relatives (reviewed by Pusey and Wolf, 1996). Dispersal of individuals from natal areas, even more so if dispersal predominantly involves one sex, can separate related individuals (Pusey and Wolf, 1996) and hence allows animals to avoid inbreeding (Hamilton and May, 1977). However, we find it unlikely that kin competition and/or inbreeding avoidance are responsible for the observed kin-biased distribution of trout in the Findhu Glen Burn or the lack of kin-biased distribution of the Dollerup Møllebæk. If kin competition would be of importance at the time of sampling, we would expect highly related trout to be found further apart than by chance as observed by Carlsson and Carlsson (2002). Moreover, inbreeding avoidance would be reflected in the distribution of mature individuals, however, we did not observe any negative or positive kin-biased distribution of mature trout in the Dollerup Møllebæk. Hence, we do not favour kin competition or inbreeding avoidance as explanations for the kin-biased structure in the Findhu Glen Burn or lack of kin-biased structure in the Dollerup Møllebæk.
An explanation for discrepancies between localities?
No correlation was observed between metric distance and relatedness in the Dollerup Møllebæk, whereas this was observed in the Findhu Glen Burn. This discrepancy is best explained by the apparent lack of highly related (eg half siblings and full siblings) individuals in the DOLOLD and DOLMAT samples. A less likely explanation, however plausible, is the context-dependent nature of kin-biased behaviours.
The observation of much smaller family sizes in the Dollerup Møllebæk compared to the Findhu Glen Burn is interesting. There are at least two possible explanations, which are not necessarily mutually exclusive: (1) The number of spawners could be higher in Dollerup Møllebæk than in Findhu Glen Burn and (2) variance in survival among families could be higher in Findhu Glen Burn than in Dollerup Møllebæk.
Concerning the first explanation, a large number of trout spawning within a limited area, such as in the Dollerup Møllebæk, would result in many, but small families among the offspring due to density-dependent mortality (Elliott, 1994), particularly if the variance in mortality among families is not too high. Conversely, if the spawning population is small (for instance due to high postsmolt mortality), but good nursery areas are available and mortality among juveniles is density dependent (Elliott, 1994), then it is expected that a sample of juvenile fish may be represented by a relatively small number of families (as in Findhu Glen Burn). There is no direct evidence supporting this explanation, but direct observations suggests that the number of lake-migratory spawners in the Dollerup Møllebæk is high, whereas only five adult sea trout spawners were caught in Findhu Glen Burn in the 1999–2000 field season (J Carlsson, unpublished results).
Concerning the second explanation, there are several factors that could lead to high variance of mortality among families, including time of hatching (eg Einum and Fleming, 2000), location of redds (eg Lisle and Lewis, 1992) and egg size (eg Thorpe et al, 1984). However, we see no obvious reason why these factors should be more important in Findhu Glen Burn than in Dollerup Møllebæk. Consequently, we tend to favour the first explanation, ie the spawning population size is higher in Dollerup Møllebæk than in Findhu Glen Burn.
An alternative to family size is the context-dependent nature of kin selection that is evident from inspection of Hamilton's rule; rb−c>0, where b is the benefit in fitness for the recipient, c is the cost in fitness for the actor and r is the coefficient of relatedness between them (Hamilton, 1964). Brown and Brown (1996) also predicted that kin-biased territorial behaviours directed towards half siblings would decrease, or cease sooner than towards full siblings as habitat quality decreases (ie food availability and increased risk of predation) since the benefits in inclusive fitness for the actor of kin-biased behaviours is reduced by decreasing relatedness to the recipient of kin-biased behaviours. Consequently, even if trout in the Dollerup Møllebæk are able to discriminate between kin and non-kin, the proportion of highly related trout (ie half and full siblings) might be too small and the relatedness between individuals might be too low for kin-biased behaviours to receive positive selection.
Regardless of the explanation, the small family sizes observed in Dollerup Møllebæk reduces the possibilities for interactions between closely related individuals considerably. Hence, we favour the arguments of family size over context-dependent kin-biased behaviours as explanation for differences in kin-biased distribution between the two streams. If this is a widespread phenomenon also in other brown trout populations, the implication is that kin selection is, in practise, only likely to be operational at very early life stages, that is before density-dependent mortality has reduced average family size, or in populations maintained by a small spawning population.
Conclusions
In conclusion, our results do not allow for unequivocally rejecting the hypothesis that association between highly related trout merely reflects dispersal from the same spawning redds. There are some indications, however, in favour of kin recognition as an explanation for the kin-biased distribution of trout in Findhu Glen Burn: the effect of spawning redd location is expected to decay as fish age; however, we observed similar individual regression slopes of pair-wise relatedness on pair-wise metric distances in older trout. Finally, the small family sizes in Dollerup Møllebæk suggest that at least in some populations, the opportunities for kin selection are limited, simply because there are very few closely related individuals left to interact.
We suggest that in future studies of kin-biased distributions of salmonid fishes in the wild, hypotheses concerning the role of kin recognition should be critically evaluated against the null hypothesis that association of related individuals reflects dispersal from spawning redds. Also, any suggestions of the presence and role of kin selection should take specific conditions of the populations into consideration, in particular if there are a sufficient number of highly related individuals to allow for kin-biased interactions among individuals.
References
Bohlin T, Hamrin S, Heggberget TG, Rasmussen G, Saltveit SJ (1989). Electrofishing – theory and practice with special emphasis on salmonids. Hydrobiologia 173: 9–43.
Broberg MM, Nielsen EE, Dieperink C (2000). Incidence of physical injury of mature male parr in a natural population of brown trout. J Fish Biol 57: 1610–1612.
Brown GE, Brown JA (1996). Kin discrimination in salmonids. Rev Fish Biol Fish 6: 201–219.
Cairney M, Taggart JB, Høyheim B (2000). Characterization of microsatellite and minisatellite loci in Atlantic salmon (Salmo salar L. and cross-species amplification in other salmonids. Mol Ecol 9: 2175–2187.
Carlsson J, Carlsson JEL (2002). Micro-scale distribution of brown trout: an opportunity for kin-selection? Ecol Freshwater Fish 11: 234–240.
Einum S, Fleming IA (2000). Selection against late emergence and small offspring in Atlantic salmon (Salmo salar). Evolution 54: 628–639.
Elliott JM (1987). The distances travelled by downstream-moving trout fry, Salmo trutta, in a lake district stream. Freshwater Biol 17: 491–499.
Elliott JM (1994). Quantitative Ecology and the Brown Trout, Oxford University Press: Oxford.
Estoup A, Largiader CR, Perrot E, Chourrout D (1996). Rapid one-tube DNA extraction for reliable PCR detection of fish polymorphic markers and transgenes. Mol Mar Biol Biotechnol 5: 295–298.
Estoup A, Presa P, Krieg F, Vaiman D, Guyomard R (1993). (CT)n and (GT)n microsatellites: a new class of genetic markers for Salmo trutta L. brown trout). Heredity 71: 488–496.
Estoup A, Rousset F, Michalakis Y, Cornuet J-M, Adriamanga M, Guyomard R (1998). Comparative analysis of microsatellite and allozyme markers: a case study investigating microgeographic differentiation in brown trout (Salmo trutta). Mol Ecol 7: 339–353.
Fontaine P-M, Dodson JJ (1999). An analysis of the distribution of juvenile Atlantic salmon (Salmo salar) in nature as a function of relatedness using microsatellites. Mol Ecol 8: 189–198.
Goudet J (1995). FSTAT, version 1.2: a computer program to calculate F-statistics. J Hered 86: 485–486.
Goudet J, Raymond M, deMeeus T, Rousset F (1996). Testing differentiation in diploid populations. Genetics 144: 1933–1940.
Gowan C, Young MK, Fausch KD, Riley SC (1994). Restricted movement in resident stream salmonids: A paradigm lost? Can J Fish Aquat Sci 51: 2626–2637.
Griffiths SW, Armstrong JD (2002). Kin-biased territory overlap and food sharing among Atlantic salmon juveniles. J Anim Ecol 71: 480–486.
Gross MR (1984). Sunfish, salmon, and the evolution of alternative reproductive strategies and tactics in fishes. In: Wootton R, Potts G (eds) Fish Reproduction: Strategies and Tactics, Academic Press: London pp 55–75.
Guo SW, Thompson EA (1992). Performing the exact test for Hardy–Weinberg proportion for multiple alleles. Biometrics 48: 361–372.
Hamilton WD (1964). The genetical evolution of social behaviour I & II. J theor Biol 7: 1–16.
Hamilton WD, May RM (1977). Dispersal in stable habitats. Nature 269: 578–581.
Hansen MM, Nielsen EE, Mensberg K-LD (1997). The problem of sampling families rather than populations: relatedness among individuals in samples of juvenile brown trout (Salmo trutta L.). Mol Ecol 6: 469–474.
Holmes WG, Sherman PW (1983). Kin recognition in animals. Am Sci 71: 46–55.
Kalleberg J (1958). Observations in a stream tank of territoriality and competition in juvenile salmon and trout (Salmo salar L. and S. trutta. L). Rep Inst Freshwater Res Drottningholm 39: 55–98.
Knight ME, Van Oppen MJH, Smith HL, Rico C, Hewitt GM, Turner GF (1999). Evidence for male-biased dispersal in Lake Malavi cichlids from microsatellites. Mol Ecol 8: 1521–1527.
Komdeur J, Hatchwell BJ (1999). Kin recognition: function and mechanism in avian societies. Trends Ecol Evol 14: 237–241.
Laird PW, Zijderveld A, Linders K, Rudnicki MA, Jaenisch R, Berns A (1991). Simplified mammalian DNA isolation procedure. Nucleic Acids Res 19: 4293.
Lisle TE, Lewis J (1992). Effects of sediment transport on survival of salmonid embryos in a natural stream: a simulation approach. Can J Fish Aquat Sci 49: 2337–2344.
McCormick SD, Hansen LP, Quinn TP, Saunders RL (1998). Movement, migration, and smolting of Atlantic salmon (Salmo salar). Can J Fish Aquat Sci 55(Suppl. 1): 77–92.
Mjølnerød IB, Refseth UH, Hindar K (1999). Spatial association of genetically similar Atlantic salmon juveniles and sex bias in spatial patterns in a river. J Fish Biol 55: 1–8.
Olsén KH (1999). Present knowledge of kin discrimination in salmonids. Genetica 104: 295–299.
O'Reilly PT, Hamilton LC, McConnel SK, Wright JM (1996). Rapid analysis of genetic variation in Atlantic salmon (Salmo salar) by PCR multiplexing of dinucleotide and tetranucleotide microsatellites. Can J Fish Aquat Sci 53: 2292–2298.
Pusey A, Wolf M (1996). Inbreeding avoidance in animals. Trends Ecol Evol 11: 201–205.
Queller DC, Goodnight KF (1989). Estimating relatedness using genetic markers. Evolution 43: 258–275.
Raymond M, Rousset F (1995). GENEPOP (version 1.2) population genetics software for exact test and ecumenicism. J Hered 86: 248–249.
Rice WR (1989). Analyzing tables of statistical tests. Evolution 43: 223–225.
Slettan A, Olsaker I, Lie Ø (1995). Atlantic salmon, Salmo salar, microsatellites at the SSOSL25, SSOSL85, SSOSL311, SSOSL417 loci. Anim Genet 26: 281–282.
Slettan A, Olsaker I, Lie Ø (1996). Polymorphic Atlantic salmon, Salmo salar L., microsatellites at the SSOSL438, SSOSL439 and SSOSL444 loci. Anim Genet 27: 57–58.
Smith BR, Herbinger CM, Merry HR (2001). Accurate partition of individuals into full-sib families from genetic data without parental information. Genetics 158: 1329–1338.
Thorpe JE, Miles MS, Keay DS (1984). Developmental rate, fecundity and egg size in Atlantic salmon, Salmo salar L. Aquaculture 43: 289–305.
Waldman B (1987). Mechanisms of kin recognition. J Theor Biol 128: 159–185.
Waldman B (1988). The ecology of kin recognition. Ann Rev Ecol Syst 19: 543–571.
Acknowledgements
We thank Andy Walker, Alan Walker and Jeff Graves for help during several phases of the project, Struan Robertson, Marcus Åkerstedt, Mikael Jakobsson and Ola Ohlsson for voluntary assistance in the field, and Dorte Bekkevold, Einar Eg Nielsen and Anti Vasemägi for discussions and comments on the manuscript. We also thank two anonymous referees for significantly improving earlier versions of the manuscript. We also thank The Baroness Willoughby de Eresby and her Ancaster Estate staff for their interest and cooperation in the Findhu Glen Burn studies. This research has been supported by a Marie Curie Fellowship of the European Community programme Quality of Life under contract number QLK5-CT-2000-52073 (JC), the Swedish Council of Forestry and Agricultural Research (JC and KHO), the Swedish University of Agricultural Sciences (JC and JELC), the Nilsson-Ehle Foundation (JC) and the Swedish Institute (JC).
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Carlsson, J., Carlsson, J., Olsén, K. et al. Kin-biased distribution in brown trout: an effect of redd location or kin recognition?. Heredity 92, 53–60 (2004). https://doi.org/10.1038/sj.hdy.6800376
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/sj.hdy.6800376
Keywords
This article is cited by
-
Long-term monitoring of a brown trout (Salmo trutta) population reveals kin-associated migration patterns and contributions by resident trout to the anadromous run
BMC Ecology and Evolution (2021)
-
MHC-mediated spatial distribution in brown trout (Salmo trutta) fry
Heredity (2012)
-
Dispersal and demography of brown trout, Salmo trutta, inferred from population and family structure in unstable Mediterranean streams
Hydrobiologia (2011)
-
Sibship within samples of brown trout (Salmo trutta) and implications for supportive breeding
Conservation Genetics (2005)