Abstract
Microsatellite markers were used to examine the population structure of Pemphigus bursarius, a cyclically parthenogenetic aphid. Substantial allele frequency differences were observed between populations on the primary host plant (collected shortly after sexual reproduction) separated by distances as low as 14 km. This suggested that migratory movements occur over relatively short distances in this species. However, the degree of allele frequency divergence between populations was not correlated with their geographical separation, indicating that isolation by distance was not the sole cause of spatial genetic structuring. Significant excesses of homozygotes were observed in several populations. Substantial allele frequency differences were also found between aphids on the primary host and those sampled from a secondary host plant after several parthenogenetic generations at the same location in two successive years. This could have been due to the existence of obligately parthenogenetic lineages living on the secondary host or genetically divergent populations confined to different secondary host plant species but sharing a common primary host.
Similar content being viewed by others
Introduction
Aphids are cyclically parthenogenetic herbivorous insects, properties that make them of interest for several reasons. Firstly, although most species are regarded as holocyclic (ie cyclical parthenogens, undergoing regular rounds of sexual reproduction, interspersed with parthenogenesis), modifications of the life cycle such as anholocycly (obligate parthenogenesis) are not uncommon (Moran, 1992). The existence of a variety of life cycles, sometimes within a single species, makes aphids valuable model organisms for studying the evolution and maintenance of sex (Simon et al, 2002). Secondly, the intimate association between aphids and their host plants makes them prone to the formation of specialised host races (eg Via, 1999; Via et al, 2000). Consequently, aphids are also valuable models for studying the conditions under which sympatric divergence and, ultimately, speciation can occur (Via, 2001). Thirdly, many aphid species are agricultural pests (Blackman and Eastop, 2000). An appreciation of the level of genetic diversity and the manner in which it is structured will be of value in developing pest management strategies. For example, studies of pest diversity could inform crop improvement programmes, helping to minimise the risk of pest genotypes able to attack new cultivars assumed to be pest resistant (Bournoville et al, 2000). Furthermore, studies of gene flow in insects are revealing the distances over which members of a given species typically disperse (Loxdale et al, 1993; Loxdale and Lushai, 2001). This information could be used to determine the spatial scales over which pest forecasting techniques might operate.
The lettuce root aphid, Pemphigus bursarius (L.) is one of several aphid species that feed on lettuce (Lactuca sativa, L.) (Blackman and Eastop, 2000) and can cause severe damage to crops (Dunn, 1959). Additionally, it is able to colonise a variety of non-crop species, largely within the Compositae (Dunn, 1959; Alleyne and Morrison, 1977). This aphid is regarded as holocyclic, alternating annually between sexual reproduction on a primary woody host, poplar (Populus nigra, L.) and parthenogenesis on its secondary hosts such as lettuce (see Dunn, 1959). Sexual reproduction occurs in the autumn and the resultant eggs remain in diapause throughout the winter. Egg hatch in spring gives rise to a foundress that feeds on the petiole of the primary host, causing a gall to form, in which she begins to reproduce parthenogenetically. The offspring of this foundress are winged and migrate to the secondary hosts, where their offspring in turn colonise the root systems. After several parthenogenetic generations on the secondary hosts, another winged form emerges in the autumn, which migrates back to poplar and gives rise to the sexual forms (Dunn, 1959).
Aphid populations frequently exhibit high levels of genetic structuring. In some species, this is due to limited dispersal capabilities, giving rise to an isolation by distance effect (Loxdale et al, 1993). Another common source of population structuring is host plant utilisation, with many aphid species being subdivided into host plant specific races (eg Via, 2001). A third source of population subdivision is the occurrence of lineages with different reproductive strategies (eg holocycly vs anholocycly) within a single species (Simon et al, 1999a).
The complex life cycle of P. bursarius suggests two areas of considerable interest: (1) the distance over which the regular migration between primary and secondary hosts promotes gene flow and, (2) the genetic relationship between populations found on the primary and secondary hosts. The aim of this paper was thus to use molecular markers (microsatellites) to investigate the degree of spatial population genetic structuring and to assess whether aphids occupying the primary and secondary host plants may be regarded as a single population.
Methods
Sample collection
Samples of P. bursarius foundresses were collected from stands of poplar at various locations in England in July 1997 (Figure 1). The locations were: Horticulture Research International (hereafter referred to as HRI), Wellesbourne, Warwickshire; Warwick, Warwickshire; Second Drove, Cambridgeshire; Squires Drove, Norfolk and Sheeplands Farm, Berkshire. Three separate stands of poplar were sampled within Warwick: the north and south ends of the County Council staff sports ground (separated by 200 m) and St Nicholas' Park, 1 km from the sports ground.
A second sample of foundresses was collected from the same poplars at HRI in June 1998. Samples of adult aphids were also collected from the roots of lettuce grown at HRI (500 m from the poplar trees) in September 1996 and September 1997, around the time of return (autumn) migration to poplar and therefore expected to represent potential parents of the generation found on poplar the following year. Each adult was collected from a different lettuce plant to minimise the incidence of sampling multiple members of the same clone. All insects were stored at −20°C prior to genotypic testing.
Microsatellite genotyping
DNA was extracted from 27 to 36 individuals per sample using a PureGene kit (Gentra Systems) and re-suspended in 20–30 μl TE buffer. DNA extracts were used as PCR templates to amplify the P. bursarius microsatellite loci Pb 02, Pb 10, Pb 16, Pb 23 and Pb 29 (Miller et al, 2000). Amplified fragments were separated by electrophoresis using 4% polyacrylamide gels on an ABI Prism 377 DNA sequencer (Applied Biosystems) and their sizes estimated by comparison with GS350 standards (Applied Biosystems), co-loaded with each sample.
Data analysis
Observed and expected heterozygosities were calculated using GENEPOP 3.1d (Raymond and Rousset, 1995). Weir and Cockerham's (1984) θ and f estimators of FST and FIS were calculated using FSTAT 2.9.3 (Goudet, 1995). The significance of departures of FST and FIS from zero was tested using a randomisation procedure within FSTAT. Where tables of multiple tests were conducted, the significance level was adjusted by the sequential Bonferroni method (Sokal and Rohlf, 1995). However, since this procedure made little difference to the overall pattern of significant results, the unadjusted results are shown.
The method of Overall and Nichols (2001) was used to analyse excess homozygosity in the samples collected from poplar, which estimates the joint likelihood of substructure and selfing. Since this method has only been demonstrated successfully for 10 highly variable loci, it was necessary to validate the approach using simulated data sets with five loci and similar levels of polymorphism to those encountered in the present study. Simulated data sets were generated with allele frequencies comparable to those observed in each sample of foundresses. A total of 30 replicate simulations were carried out for a population with a selfing rate (S) of 0.6, a subdivided population with θ = 0.3 and a subdivided population with selfing within subpopulations (θ = 0.3, S = 0.6) using the same approach as Overall and Nichols (2001). Consensus likelihood surfaces were then produced from the 30 simulations. In all cases, the simulated parameter values were estimated correctly (data not shown).
Results
General levels of variability
All five loci tested were polymorphic with totals of seven, five, seven, eight and eight alleles observed at loci Pb 02, Pb 10, Pb 16, Pb 23 and Pb 29, respectively. However, no individual sample contained the full range of alleles (Table 1). Locus Pb 23 was monomorphic in two samples: Sheeplands Farm and the 1996 HRI lettuce sample. The observed and expected heterozygosities and number of alleles observed at each locus in each sample are given in Table 1.
Genetic differences between samples
Several pairs of samples exhibited significant allele frequency differences. Where this was the case, the difference was usually substantial as indicated by estimates of FST>0.3 (Table 2). The overall pattern of differences allowed the samples to be assigned to one of three groups:
-
1
Samples from poplar at Warwick and HRI (both years). No significant allele frequencies were seen between these samples.
-
2
Samples from poplar at Sheeplands Farm and Squires Drove and from lettuce at HRI (both years). Allele frequency differences between these samples were either not significant, or significant but relatively small (FST < 0.05). Furthermore, no differences were significant after correction for multiple tests (not shown).
-
3
Second Drove.
The relationship between geographical distance and genetic divergence between sites was generally poor. For example, there was substantial genetic divergence between Squires Drove and Second Drove (FST=0.422) which were separated by a distance of only 14 km. In contrast, Squires Drove was not significantly different from Sheeplands Farm, some 140 km away. An irregular pattern was also seen for the samples taken from Warwick and HRI. Allele frequencies in the samples from poplar at HRI did not differ from those in samples from poplar at Warwick, but were significantly different from the samples taken from lettuce at the same site. Although there were pronounced genetic differences between samples from the primary and secondary hosts at HRI, allele frequencies were not significantly different between years.
Deviations from Hardy–Weinberg genotypic proportions within sexually derived samples
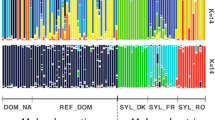
No cases of significant excess heterozygosity (FIS<0) were found in the samples collected from poplar. However, significant excess homozygosity (FIS > 0) at one or more loci was found at HRI (both years), Warwick sports ground (north), Squires Drove and Second Drove (Table 3). Maximum likelihood estimation of the rate of selfing and degree of substructuring (Overall and Nichols, 2001) indicated that at HRI and Squires Drove, homozygous excess was due to both selfing and substructuring, whereas at Second Drove, excess homozygosity appeared to be entirely due to substructuring (Table 3). However, in all cases, levels of population substructuring at or near zero were plausible (Figure 2).
Maximum likelihood estimates of S, the selfing rate and θ (FST) for samples with excess homozygosity. Contours encompass 95, 50 and 5% of the total likelihood. Only samples where a significant general trend towards excess homozygosity (‘All’ in Table 3) are shown.
Duplication of multilocus genotypes at HRI
Several individuals with identical genotypes at all five loci were seen on lettuce at HRI in 1996 and 1997. Within the 1996 sample, 62% of individuals shared their genotype with at least one other individual and 44% in 1997. Several of these multilocus genotypes were common to both years including the most abundant one (Table 4). In contrast, no individual in the HRI poplar samples shared its genotype with any other individual.
Discussion
Populations of P. bursarius foundresses collected from poplar often exhibited a substantial excess of homozygotes. Excess homozygosity has also been seen in holocyclic populations of Sitobion avenae (Simon et al, 1999a) and is thought to be due to ‘clonal selection’ acting to increase the frequency of relatively homozygous genotypes of high fitness (Sunnucks et al, 1997a). In the present study, clonal selection cannot explain the excess homozygosity on poplar since parthenogenetic reproduction had yet to occur. In contrast, a recent study of microsatellite variation in Myzus persicae (Sulzer) on its primary host found that populations generally conform to Hardy–Weinberg expectations (Wilson et al, 2002). Studies of allozyme variation in Rhopalosiphum padi (L.) on its primary host in France (Simon et al, 1996; Simon and Le Gallic, 1998) and Canada (Simon and Hebert, 1995) have also found populations that were apparently in Hardy–Weinberg equilibrium. However, a more recent study of R. padi on its primary host, using microsatellites, has revealed significant excess homozygosity (Delmotte et al, 2002). These authors identified null alleles, selection, inbreeding and the Wahlund effect as potential causes of excess homozygosity.
Ideally, null alleles can be detected by employing alternative PCR primers (eg Callen et al, 1993) or through apparent segregation distortions (eg Sefc et al, 1999). Unfortunately, neither approach is tenable in the present study due to limited quantities of template DNA and a lack of pedigree information. It is sometimes possible to discount the existence of null alleles when missing observations, interpreted as null homozygotes, are not encountered (eg Delmotte et al, 2002; Llewellyn et al, 2003) provided that null homozygotes not selected against (Van Treuren, 1998). Occasional missing observations were present in this study, but were not confined to sample and locus combinations exhibiting excess homozygosity, suggesting that other causes (eg template DNA quality) were at least partly responsible. The existence of null alleles may be inferred where excess homozygosity is confined to a single locus, or specific loci (eg Van Treuren, 1998), while the remainder consistently exhibit Hardy–Weinberg genotypic proportions. No such pattern is seen in the present study (Table 3) leading to the conclusion that there is no clear evidence for the existence of null alleles at the loci employed.
It is possible that selection against heterozygotes contributed the observed excess homozygosity. However, as has been noted for holocyclic R. padi, any such selection must be acting on the overwintering eggs or the foundresses and cannot be clonal selection (Delmotte et al, 2002). Inbreeding is a plausible source of excess homozygosity in P. bursarius since there is the possibility of matings between males and females of the same clone (effectively self-fertilisation) and has been suggested as the cause of homozygous excess in P. spyrothecae (Passerini) (Johnson, 2000). Maximum likelihood analysis of excess homozygosity seen in the present study supported the view that selfing was a contributing factor in several populations, in combination with a degree of population substructuring.
No clear relationship between the genetic divergence and geographical separation between samples was found. Hence, processes other than isolation by distance were important in determining the observed population structure. Frequent local bottlenecks could produce an irregular population structure, which casual observations made in the course of sample collection for the present study indicate may occur in P. bursarius. Attempts were made in 1998 to resample aphids from all the stands of poplar sampled in 1997. However, with the exception of HRI, samples were not collected in 1998 as aphids were scarce or entirely absent. Mature galls were often found, but did not contain any aphids, characteristic of anthocorid (Heteroptera: Anthocoridae) predation (Dunn, 1960), occasionally anthocorids were found in these galls.
Perhaps the most striking aspect of the irregular population structure was the high level of genetic divergence between aphid samples from poplar and lettuce at the same location (HRI). A number of possible causes for this difference between the primary and secondary host plants can be identified.
The population sampled from lettuce had been reproducing parthenogenetically throughout the summer. During this process, it is likely that some clones will become more abundant than others, either by chance or because they have high fitness (clonal selection). There is, therefore, the possibility that several members of the same clone (with identical multilocus genotypes) were sampled, distorting the observed allele frequencies in the lettuce samples. This does appear to have been the case with duplicate genotypes, being common within the 1996 and 1997 HRI lettuce samples but not the 1997 and 1998 HRI poplar samples. The distorting effects of sampling multiple members of the same clone can be assessed by conducting genetic analyses with an edited data set in which each multilocus genotype is represented only once (Sunnucks et al, 1997a). Editing the data from the lettuce samples made no difference to the general pattern of population structure in this case (not shown), indicating that clonal amplification was not responsible for the difference between aphids from lettuce and poplar at HRI.
Aphids are rather weak fliers, and will be strongly influenced by the wind in the direction they travel in flight (Loxdale et al, 1993). Consequently, many individuals leaving their primary host may fail to find a suitable secondary host, as seen in R. padi (Ward et al, 1998). This may lead to a founder effect during the transition from primary to secondary host, resulting in allele frequency differences. However, the consistent allele frequencies observed between years on lettuce would not be expected if appreciable founder effects were operating.
If founder effects and the consequences of clonal amplification can be discounted as sources of population differences between poplar and lettuce at HRI, it seems likely that it is due to some form of population subdivision. Subdivision could arise if the aphids sampled from lettuce went on to colonise other, unsampled poplar trees and those sampled from poplar later colonised unsampled secondary hosts. However, the similarity between samples from poplar trees at HRI and Warwick make this hypothesis of microgeographic population structuring seem unlikely.
An alternative scenario is that the aphids sampled from lettuce did not go on to colonise poplar at all but represented an anholocyclic population. The occurrence of both holocyclic and anholocyclic lineages in aphid populations has been demonstrated or inferred in several species (Loxdale and Brookes, 1990; Moran, 1991; Simon et al, 1999a, 1999b). Allele frequency differences between holocyclic and anholocyclic aphids have been shown to be important in driving genetic divergence between populations of Sitobion avenae in France (Simon et al, 1999a) and Aphis pomi (de Geer) in Canada (Singh and Rhomberg, 1984) and have been suggested as a cause for genetic differences between populations of S. fragariae (Walker) on its primary and secondary hosts in south-east England (Loxdale and Brookes, 1990). The fact that the same multilocus genotypes were seen on lettuce in successive years suggests that some P. bursarius clones do persist throughout the winter and recolonise lettuce without an intervening sexual cycle. However, it is unclear whether these clones are anholocyclic or engage in a mixed strategy of both sexual reproduction and asexual overwintering as do some clones of R. padi (Simon et al, 1991).
A key component of an aphid's environment is the host plants upon which it feeds and several species are thought to consist of a number of specialised host races. The best documented example of host races in an aphid is the separation of the pea aphid, Acyrthosiphon pisum (Harris) into alfalfa and clover host races. It has been known for some time that pea aphid clones originating from clover and alfalfa perform better on their original host than the alternative host (Via, 1991a, 1991b). More recently, it has been shown that A. pisum colonising new host plants strongly prefer their original host. Since A. pisum carries out its sexual cycle on these hosts, this behaviour leads to reproductive isolation between sympatric populations (Via, 1999). Furthermore, hybrids between the two host races have lower fitness than either parent on its original host (Via et al, 2000).
In addition to the pea aphid, the use of molecular markers has revealed probable host races in several other aphid species. Clones of Schizaphis graminum (Rondani) are classified into several biotypes on the basis of their feeding behaviour. A recent study of mitochondrial DNA variation has revealed that these biotypes are probably host adapted races (Shufran et al, 2000). Populations of S. avenae, confined to specific host plants, have been identified using RAPD (De Barro et al, 1995; Lushai et al, 2002) and microsatellite (Sunnucks et al, 1997a) markers, as well as generalists occupying several different hosts (Sunnucks et al, 1997a; Lushai et al, 2002). RAPD markers, have also revealed distinct populations of Aphis gossypii (Glover) on curcurbit and non-curcurbit hosts (Vanlerberghe-Masutti and Chavigny, 1998) and, in conjunction with mitochondrial DNA, Therioaphis trifolii (Monell) on clover and lucerne (Sunnucks et al., 1997b).
The existence of distinct, secondary host-specific races in P. bursarius might account for the allele frequency differences seen between poplar and lettuce if a significant proportion of the aphids sampled on poplar at HRI were derived from some unsampled, non-lettuce race. Maximum likelihood analysis of the excess homozygosity seen in the HRI poplar samples detected some population subdivision, compatible with the hypothesis of more than one genetically distinct secondary host race being present on poplar. Secondary host races might also explain the spatial population genetic structure seen on poplar in 1997. Allele frequencies on poplar would be determined by the relative proportions of the different host races, which in turn would be strongly influenced by the local abundance of different hosts. Such a scenario may explain the similarity between the populations on lettuce at HRI and on poplar at Sheeplands Farm and Squire's Drove, both in regions where lettuce is cultivated commercially. However, it should be noted that for secondary host races to coexist on poplar would require some, unknown, mechanism to restrict gene flow between them (Via, 2001; Drès and Mallet, 2002).
This study attempted to elucidate the population genetic structure of P. bursarius, using microsatellites. Two clear results were apparent. Firstly, isolation by distance is an inadequate explanation for the spatial structure of this species. Secondly, there are pronounced genetic differences between P. bursarius populations on poplar, its primary host and lettuce, one of its secondary hosts. No firm conclusions can, as yet, be drawn as to the processes involved in generating this irregular population structure. However, the existence of both obligate and cyclical parthenogenetic lineages or of several divergent, secondary host races are both possible explanations. A detailed study of P. bursarius populations isolated from poplar and several secondary hosts is currently underway to investigate these ideas.
References
Alleyne EH, Morrison FO (1977). The lettuce root aphid, Pemphigus bursarius (L.) (Homoptera: Aphidoidea) in Quebec, Canada. Ann Entomol Soc Quebec 22: 171–180.
Blackman RL, Eastop VF (2000). Aphids on the World's Crops. John Wiley & Sons: Chichester, UK.
Bournoville R, Simon JC, Badenhausser I, Girousse C, Guilloux T, Andre S (2000). Clones of pea aphid, Acyrthosiphon pisum (Hemiptera: Aphididae) distinguished using genetic markers, differ in their damaging effect on a resistant alfalfa cultivar. Bull Entomol Res 90: 33–39.
Callen DF, Thompson AD, Shen Y, Phillips HA, Richards RI, Mulley JC et al (1993). Incidence and origin of ‘null’ alleles in the (AC)n microsatellite markers. Am J Hum Genet 52: 922–927.
De Barro PJ, Sherratt TN, Brookes CP, David O, Maclean N (1995). Spatial and temporal genetic variation in British field populations of the grain aphid Sitobion avenae studied using RAPD-PCR. Proc Roy Soc Lond Series B 262: 321–327.
Delmotte F, Leterme N, Gauthier JP, Rispe C, Simon JC (2002). Genetic architecture of sexual and asexual populations of the aphid Rhopalosiphum padi based on allozyme and microsatellite markers. Mol Ecol 11: 711–723.
Drès M, Mallet J (2002). Host races in plant-feeding insects and their importance in sympatric speciation. Philos Trans R Soc Lond Ser B 357: 471–492.
Dunn JA (1959). The biology of the lettuce root aphid. Ann Appl Biol 47: 475–491.
Dunn JA (1960). The natural enemies of the lettuce root aphid, Pemphigus bursarius (L.). Bull Entomol Res 51: 271–277.
Goudet J (1995). FSTAT (Version 1.2): A computer program to calculate F- statistics. J Hered 86: 485–486.
Johnson PCD (2000). Genetic variation in the aphid Pemphigus spyrothecae. PhD Thesis, University of Cambridge.
Llewellyn KS, Loxdale HD, Harrington R, Brookes CP, Clark SJ, Sunnucks P (2003). Migration and genetic structure of the grain aphid (Sitobion avenae) in Britain related to climate and clonal fluctuation as revealed using microsatellites. Mol Ecol 12: 21–34.
Loxdale HD, Brookes CP (1990). Temporal genetic stability within and restricted migration (gene flow) between local populations of the blackberry-grain aphid Sitobion fragariae in south-east England. J Anim Ecol 59: 497–514.
Loxdale HD, Hardie J, Halbert S, Foottit R, Kidd NAC, Carter CI (1993). The relative importance of short-range and long-range movement of flying aphids. Biol Rev Cambridge Philos Soc 68: 291–311.
Loxdale HD, Lushai G (2001). Use of genetic diversity in movement studies of flying insects. In: Woiwod IP, Reynolds DR, Thomas CD (eds) Insect Movement: Mechanisms and Consequences, CAB International: Wallingford. pp 361–386.
Lushai G, Markovitch O, Loxdale HD (2002). Host-based genotype variation in insects revisited. Bull Entomol Res 92: 159–164.
Miller NJ, Birley AJ, Tatchell GM (2000). Polymorphic microsatellite loci from the lettuce root aphid, Pemphigus bursarius. Mol Ecol 9: 1951–1952.
Moran NA (1991). Phenotype fixation and genotypic diversity in the complex life- cycle of the aphid Pemphigus betae. Evolution 45: 957–970.
Moran NA (1992). The evolution of aphid life-cycles. Annu Rev Entomol 37: 321–348.
Overall ADJ, Nichols RA (2001). A method for distinguishing consanguinity and population substructure using multilocus genotype data. Mol Biol Evol 18: 2048–2056.
Raymond M, Rousset F (1995) Genepop (Version 1.2)–Population genetics software for exact tests and ecumenicism. J Hered 86: 248–249.
Sefc KM, Regner F, Turetschek E, Glossl J, Steinkellner H (1999). Identification of microsatellite sequences in Vitis riparia and their applicability for genotyping of different Vitis species. Genome 42: 367–373.
Shufran KA, Burd JD, Anstead JA, Lushai G (2000). Mitochondrial DNA sequence divergence among greenbug (Homoptera: Aphididae) biotypes: evidence for host-adapted races. Insect Mol Biol 9: 179–184.
Simon JC, Baumann S, Sunnucks P, Hebert PDN, Pierre JS, Le Gallic JF et al. (1999a). Reproductive mode and population genetic structure of the cereal aphid Sitobion avenae studied using phenotypic and microsatellite markers. Mol Ecol 8: 531–545.
Simon JC, Blackman RL, Le Gallic JF (1991). Local variability in the life-cycle of the bird cherry-oat aphid, Rhopalosiphum padi (Homoptera, Aphididae) in western France. Bull Entomol Res 81: 315–322.
Simon JC, Carrel E, Hebert PDN, Dedryver CA, Bonhomme J, Le Gallic JF (1996). Genetic diversity and mode of reproduction in French populations of the aphid Rhopalosiphum padi L. Heredity 76: 305–313.
Simon JC, Hebert PDN (1995). Patterns of genetic variation among Canadian populations of the bird cherry-oat aphid, Rhopalosiphum padi L (Homoptera, Aphididae). Heredity 74: 346–353.
Simon JC, Le Gallic JF (1998). Patterns of allozyme variation among French populations of the cereal aphid Rhopalosiphum padi (Aphididae). In: Nieto Nafrìa JM, Dixon AFG (eds) Aphids in Natural and Managed Ecosystems, Universidad de León: León. pp 105–111.
Simon JC, Leterme N, Latorre A (1999b). Molecular markers linked to breeding system differences in segregating and natural populations of the cereal aphid Rhopalosiphum padi L. Mol Ecol 8: 965–973.
Simon JC, Rispe C, Sunnucks P (2002). Ecology and evolution of sex in aphids. Trends Ecol Evol 17: 34–39.
Singh RS, Rhomberg L (1984). Allozyme variation, population structure, and sibling species in Aphis pomi. Can J Genet Cytol 26: 364–373.
Sokal RR, Rohlf FJ (1995). Biometry. Freeman: New York.
Sunnucks P, DeBarro PJ, Lushai G, Maclean N, Hales D (1997a). Genetic structure of an aphid studied using microsatellites: cyclic parthenogenesis, differentiated lineages and host specialization. Mol Ecol 6: 1059–1073.
Sunnucks P, Driver F, Brown WV, Carver M, Hales DF, Milne WM (1997b). Biological and genetic characterization of morphologically similar Therioaphis trifolii (Hemiptera: Aphididae) with different host utilization. Bull Entomol Res 87: 425–436.
Van Treuren R (1998). Estimating null allele frequencies at a microsatellite locus in the oystercatcher (Haematopus ostralegus). Mol Ecol 7: 1413–1417.
Vanlerberghe-Masutti F, Chavigny P (1998). Host-based genetic differentiation in the aphid Aphis gossypii Glover, evidenced from RAPD fingerprints. Mol Ecol 7: 905–914.
Via S (1991a). The genetic structure of host plant adaptation in a spatial patchwork – Demographic variability among reciprocally transplanted pea aphid clones. Evolution 45: 827–852.
Via S (1991b). Specialized host plant performance of pea aphid clones is not altered by experience. Ecology 72: 1420–1427.
Via S (1999). Reproductive isolation between sympatric races of pea aphids. I. Gene flow restriction and habitat choice. Evolution 53: 1446–1457.
Via S (2001). Sympatric speciation in animals: the ugly duckling grows up. Trends Ecol Evol 16: 381–390.
Via S, Bouck AC, Skillman S (2000). Reproductive isolation between divergent races of pea aphids on two hosts. II. Selection against migrants and hybrids in the parental environments. Evolution 54: 1626–1637.
Ward SA, Leather SR, Pickup J, Harrington R (1998). Mortality during dispersal and the cost of host-specificity in parasites: how many aphids find hosts? J. Anim Ecol 67: 763–773.
Weir BS, Cockerham CC (1984). Estimating F-statistics for the analysis of population structure. Evolution 38: 1358–1370.
Wilson ACC, Sunnucks P, Blackman RL, Hales DF (2002). Microsatellite variation in cyclically parthenogenetic populations of Myzus persicae in south-eastern Australia. Heredity 88: 258–266.
Acknowledgements
Nick Miller was supported by a Biotechnology and Biological Sciences Research Council CASE studentship, in conjunction with Horticulture Research International. Hugh Loxdale, Neil Kift, Rosemary Collier, James Lynn and two anonymous reviewers provided valuable comments on earlier drafts of the manuscript.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Miller, N., Birley, A., Overall, A. et al. Population genetic structure of the lettuce root aphid, Pemphigus bursarius (L.), in relation to geographic distance, gene flow and host plant usage. Heredity 91, 217–223 (2003). https://doi.org/10.1038/sj.hdy.6800331
Received:
Published:
Issue Date:
DOI: https://doi.org/10.1038/sj.hdy.6800331
Keywords
This article is cited by
-
The effect of urban–rural gradient on black poplar endophagous herbivorous insects
Arthropod-Plant Interactions (2023)
-
The influence of geographic distance on life-history traits of the carob moth, Ectomyelois ceratoniae (Zeller) (Lepidoptera: Pyralidae)
International Journal of Tropical Insect Science (2022)
-
Genetic structure and diversity of a newly invasive species, the codling moth, Cydia pomonella (L.) (Lepidoptera: Tortricidae) in China
Biological Invasions (2013)
-
Maintaining genetic diversity and population panmixia through dispersal and not gene flow in a holocyclic heteroecious aphid species
Heredity (2012)
-
Host-associated divergence and incipient speciation in the yucca moth Prodoxus coloradensis (Lepidoptera: Prodoxidae) on three species of host plants
Heredity (2010)