Abstract
Nine out of 57 bovine and caprine microsatellites investigated have proved polymorphic in roe deer populations from Central Europe. The polymorphism of four to nine microsatellites (with two to 16 alleles each) has been screened in 492 roe deer from 27 sample locations in Germany, the Netherlands and France, and 10 allozyme loci have been investigated in 118 roe deer from Germany. These studies have revealed a genetically homogeneous population, but with a local scatter of allele frequencies. The mean genetic distance among sample pairs, and the overall fixation index for the 27 population samples were D=0.1638 and GST=0.0972 for four microsatellite loci, and D=0.0598 and GST=0.1459 for 10 allozyme loci. No isolation-by-distance was observed. Roe deer from isolated habitats could be distinguished by various measures of genetic variability. The expected heterozygosity and the allelic diversity were higher in male than in female roe deer, but mean genetic distances and fixation indices were higher in females. The fixation indices of pairs of adjacent samples, and the genetic distance among these samples correlated highly significantly with the density of human settlement, measured by the percentage of land surface covered by roads and villages. The utility of allozymes and microsatellites for population genetic studies in cervids are compared.
Similar content being viewed by others
Introduction
Although the roe deer, Capreolus capreolus, is the largest free-living mammal in many regions throughout Europe, and a species of major importance in hunting, the understanding of its evolutionary genetics lags behind the insights gained for other cervids, especially the wapiti/red deer superspecies (Cervus elaphus), the moose (Alces alces) and the white-tailed deer (Odocoileus virginianus), which, to some extent, is the Nearctic ecological replacement form of the roe (Dratch & Pemberton, 1992; Pemberton & Slate, 1998). Several studies of allozyme variants in roe deer, mainly in populations in south-east Central Europe, showed it to be one of the most heterozygous and polymorphic cervid species (Hartl et al., 1991, 1993). The few studies of DNA variability have been confined to smaller population samples (mt-DNA: Jaeger et al., 1992; Randi et al., 1998; Wiehler & Tiedemann, 1998; fingerprints: Mörsch & Leibenguth, 1993; Volmer et al., 1995). Pemberton & Slate (1998) screened four polymorphic microsatellites in an island population from Norway. No large-scale polymorphism screening of German roe deer populations has been undertaken at either the protein or the DNA level.
The roe deer is abundant throughout the European temperate zone. Originally favouring the early regrowth stages of natural forest succession, it thrives in forests opened by silviculture, but is also found in intensively cultivated agricultural areas. The ecology and social behaviour of the roe deer are fairly well-known (Strandgaard, 1972; Ellenberg, 1978; Stubbe, 1990; Kurt, 1991). Many aspects of social organization and population ecology, including those which might be relevant for population genetics too, differ among regional populations using different habitats. Stable social systems, e.g. matrilineal clans, prevail in forests, whereas unstable social structures, e.g. aggregations, are commonly discerned in field habitats (Kurt, 1991). Kurt et al. (1993) claimed to have found significantly different FIS values between roe deer of the ‘forest’ and ‘field’ ecotypes. Moreover, intensive hunting pressure was thought to augment heterozygosity by disrupting social groups. However, Hartl et al. (1993) observed no elevated genetic distances when field- and forest-dwelling roe deer were compared. In general, roe deer are rather philopatric and maintain small home ranges for many years (Kurt, 1991). Dispersal is maximal in the age-classes of one or two years, and is sex-biased, with males tending to disperse earlier and further than females (Ellenberg, 1978; Stubbe, 1990; Kurt, 1991). The impact of sex-specific dispersal on the population genetics of a species not only throws light on a species’s population biology, but also complicates the interpretation of allelic evolution in fragmented populations. The latter aim is of primary interest for wildlife management, as discussed for red deer (Schreiber et al., 1994). Few publications refer to the genetic consequences of habitat fragmentation on roe deer: Wehner et al. (1991) found reduced allozyme variability in one of three populations studied in south-west Germany which had lived in isolation for 60 years, but even lower variability was observed in one seemingly non-isolated population sample. Fakler & Schreiber (1997) reported low genetic variability in the recently founded population of a Dutch nature reserve surrounded by city agglomerations.
The present study addresses the population structure of 27 roe deer population samples from Germany, the Netherlands and France, including 492 individuals. Microsatellite analysis compares samples using these highly polymorphic markers, which nevertheless permit the identification of homozygous and heterozygous genotypes. Ten allozyme loci were additionally screened in six population samples from which frozen tissue was available. We demonstrate an overall population homogeneity across Central Europe, with local scatter of allele frequencies. The genetic distinction of local populations correlated significantly with the intensity of human land-use. The subsamples of male and female roe deer differed, probably reflecting different space occupation of bucks and does. The utility of microsatellites and allozymes for detecting isolated roe deer populations is discussed.
Methods
Populations examined
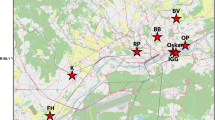
The total sample consisted of 492 roe deer from 27 collection sites (Fig. 1, Table 1). The specimens from the Vosges Mountains (VOS) were collected during 1980–90. Samples from Düsseldorf (DÜS) and Paderborn (PAD) were taken in 1993 and 1994, and the other samples from 1996 to 2000.
Several sampled populations are isolated from genetic exchange or have been artificially created: The 3400 hectares of the Amsterdamse Waterleidingduinen (AWD), North Holland, were stocked in 1952, with 10–12 roe deer transferred from the eastern Netherlands. For population status and development, see Fakler & Schreiber (1997). Flevoland (FLE), north-central Netherlands, was colonized spontaneously by an unknown number of founders when this polder area became dry in 1959. BR2, located at about 4 km from the non-isolated site BR1, denotes a park in Bremen city, which has been isolated since about 1980 and contains about 70 roes (H. Tempelmann, personal communication). Rügen Island (927 km2) in the Baltic Sea (RÜG) is separated from the mainland by a 1–2 km broad strait (A. Siefke, personal communication). BIE and KÜH refer to the nature reserves of Biedensand (525 hectares) and Kühkopf-Knoblochsaue (2100 hectares), Hessen, in the flood plains of the Rhine; they are semi-isolated, being surrounded by the Rhine and by old meanders sequestered from the Rhine. The samples from VOS originated from male roe deer trophies of a population which had passed bottlenecks prior to the early 1980s (G. Lang, personal communication). Roe deer from Münsingen (MÜN) were culled in a fenced military training area; the fence has probably become penetrable over the years (E. Hördler, personal communication).
Sampling
Samples from liver, spleen, kidney or muscle tissue were taken from hunted roe deer and conserved at either −70°C or at room temperature in 96% ethanol. The samples DÜS and PAD were received as dried tissue. Bone and antler samples from VOS and RÜG were stored at room temperature.
Microsatellites
Genomic DNA was extracted with the Qiamp tissue kit (Qiagen). The DNA extraction from bone and antler powder followed a modification of Launhardt et al. (1998). PCR was carried out in an reaction volume of 20 μL, using 50 ng of DNA, 10 μM primer (applying the primers listed in Table 2), 100 μM dNTP, and 2 U Taq polymerase. Five PCR cycles, as follows, were run after initial denaturation at 94°C for 2 min: 45 s at 94°C; 45 s at the annealing temperature of the primer (Table 2) plus 4°C; 45 s at 72°C. Another 30 cycles comprised the following steps: 45 s at 94°C; 45 s at the annealing temperature of the primer minus 1°C; 45 s at 72°C; and finally 2 min at 72°C. The primers McM505 and McM507 were amplified with a stepdown PCR as reported by Hulme et al. (1995). Amplicons were separated in an 8% polyacrylamide gel in an ALF Express sequencer.
Allozymes
Allozyme loci were assayed from homogenized liver tissue as presented before (Fakler & Schreiber, 1997). The alleles were designated by their electrophoretic mobilities in relation to the mobility of the most frequent variant which was defined as 100%.
Statistics
Recently, a number of statistics for the analysis of microsatellite data have been developed, assuming a stepwise mutation model (SMM). The proposed measures were designed for phylogenetic analyses, rating allele size as an evolutionary character. However, when divergence is low, as in intraspecific population comparisons, methods based on the infinite-allele model were considered appropriate (Takezaki & Nei, 1996). Additionally, the SMM is not compatible with numerous observations on microsatellite evolution (Wierdl et al., 1997; Colson & Goldstein, 1999). Therefore, we used standard measures (i.e. Nei distances, GST-values, etc.) for both allozyme and microsatellite data. For the analysis of the among-population component of genetic differentiation, fixation indices (GST) were corrected for small sample size, according to Nei & Chesser (1983). The hierarchical gene diversity analysis followed Nei (1973). Unbiased genetic distances were derived from the allele frequencies according to Nei (1978). Nei distances and fixation indices were correlated with geographical distances using Mantel’s test (Mantel, 1967), applying the software package NTSYS/PC 2.02f (Rohlf, 1998). Data transformation for this test, to adjust the different scales of geographical and genetic distances, followed Sokal (1979). χ2-tests of Hardy–Weinberg equilibrium used the Bonferroni and Yates corrections. Heterogeneity analysis of allele frequencies, testing deviations from hypothetical metapopulation expectations, followed Workman & Niswander (1970). Assignment indices (Favre et al., 1997) were used to compare the extent of male vs. female immigration into populations.
Results
Genetic variability
From 57 bovine and caprine microsatellite primers tested in four roe deer, 32 produced amplicons with roe deer DNA, and nine proved polymorphic (ILSTS005, ILSTS008, ILSTS058, OarAE129, OarCP26, OarHH51, McM131, McM505 and McM507). In order to examine the variability of these nine microsatellites, seven population samples, AWD, FLE, WES, KÜH, BIE, MÜN and BAD, were screened in the first part of the analysis. The number of alleles and the expected heterozygosities found are listed in Table 2. In the second part of the study, we screened all roe deer samples collected with four primers, amplifying from two to 16 alleles each (McM507: two alleles; OarCP26: three alleles; OarAE129: seven alleles; ILSTS058: 16 alleles). Allele frequencies (p), polymorphism (P), expected heterozygosities (He), and the allele numbers per locus (A.D.) obtained for each sample are listed in Table 3; the mean values amounted to P=0.990, He=0.545 and A.D.=4.04.
Six allozyme loci, out of 10 (Ak-1*, Ak-2*, Dia-1*, Dia-2*, Me-1*, Me-2*, Mpi*, Pgm-1*, Pgm-2* and Sod*) screened in 118 roe deer from WES, KÜH, BIE, LAN, GEF and OBE, proved polymorphic, having two (Ak-1*, Dia-2*, Me-2*, Pgm-1*) or three alleles (Mpi*, Pgm-2*), respectively. Mean P, He and A.D. for all 10 allozyme loci were P=0.583, He=0.213 and A.D.=1.63. The corresponding values for single population samples and the allele frequencies are listed in Table 4.
Population structure
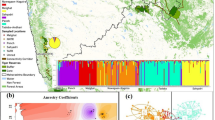
Neither the geographical distribution of microsatellite or protein alleles nor their frequencies correlated with the geographical arrangement of the collection sites. Nor did a principal component analysis (PCA, Fig. 2) of allele frequencies reveal any regional groupings (with the weights of PC1, PC2 and PC3 of 14, 13 and 10% in microsatellites and 41, 31 and 15% in allozymes, respectively). Mean unbiased genetic distances and fixation indices were calculated for each sample, by averaging D or GST values over all possible pairings of a respective sample (Table 3). The overall mean microsatellite Nei distance was D=0.1638 and the overall fixation index for all populations was GST=0.0972. The mean allozyme Nei distance was D=0.0584, and overall GST amounted to GST=0.1459. Using a Mantel test, neither the fixation indices nor the genetic distances from microsatellite or allozyme data correlated with the geographical distances separating the sample sites (microsatellite GST: r=−0.076, t=−1.015, P=0.8650; microsatellite D: r=−0.064, t=−0.871, P=0.8082; allozyme GST: r=−0.289, t=−1.208, P=0.8601; allozyme D: r=−0.244, t=−1.028, P=0.8480). A hierarchical gene diversity analysis divided the overall GST-value into percentages, which explain the influence of local and regional groupings on population subdivision. The following sample groups were defined for microsatellites: Netherlands (AWD, FLE), north German lowlands (DAN, HOR, AHR, BR1, BR2), east German lowlands (UCK, SCH, RÜG), central Germany (DÜS, PAD, BIK, WES), south-west German highlands and Rhine basin (KÜH, BIE, MÜK, VOS, LAN, BAD, BÖB, MÜN), and Bavaria (OBE, GEF, TRE, FIB, BAY). Only 27.2% (GCS(T)=0.0264) of the overall genetic differentiation (GST=0.0972) was due to these regional groupings. In conclusion, 72.8% of GST was explained by small-scale population diversity, i.e. on the spatial level of local samples. For allozymes, the groupings for the hierarchical gene diversity analysis were: Oberfranken (OBE, GEF), Rhine basin (KÜH, BIE, LAN) and Westerwald (WES). Of the overall gene diversity, 24.8% (GCS(T)=0.0362) was caused by allozyme variation between these regional groups; thus 75.2% remains for the small-scale variation on the level of single samples.
Principal component analyses of the allele frequencies of 27 roe deer population samples analysed for four microsatellite loci (a), and of six samples screened for 10 allozyme loci (b). The numbers (a) and abbreviations (b) refer to the samples defined in Table 1
Significant deviations from the Hardy–Weinberg expectations, expressed by a deficiency of heterozygotes, were revealed occasionally in χ2-tests comparing the frequencies of homo- and heterozygotes: FLE (ILSTS058: χ2=18.56, d.f.=1, P=0.0002); BIK (OarAE129: χ2=6.45, d.f.=1, P=0.0444); KÜH (ILSTS058: χ2=17.40, d.f.=1, P=0.0003); and WES (Me-2*: χ2=16.48, d.f.=1, P=0.0003).
Sex-specific genetic population structure
In 13 population samples, those comprising at least four males and four females, the microsatellite data were analysed separately for the sexes (populations FLE, DÜS, UCK, BIK, WES, KÜH, BIE, MÜK, LAN, BÖB, GEF, OBE, BAY). These test groups included 145 males and 157 females. In males the mean polymorphism, expected heterozygosity and allelic diversity were P=1.000, He=0.579 and A.D.=3.976, and in females P=0.991, He=0.554 and A.D.=3.707. Assignment indices, calculated for males and females (see Table 5), were significantly lower in bucks, indicating a higher share of male dispersal among our population samples (Mann–Whitney U-test: U=38; N=13; P=0.0171). The sexes also differed, almost significantly (Friedeman ANOVA: χ2=3.769, N=13, d.f.=1, P=0.052), in the mean genetic distances between pairs of single-sex population samples. The mean among-sample distance value was D=0.1377 in males, and D=0.1807 in females. The total fixation indices differed markedly between the sexes too (GST=0.0695 in males and GST=0.1034 in females), as did the amount of population subdivision into local or regional groupings (hierarchical gene diversity analysis of the groups defined above, but including only the 13 population samples mentioned): in males, 86.7% of the total gene diversity was due to small-scale variation on the level of single samples, and in females 94.5%. Heterogeneity analyses of allele frequencies, testing if male and female samples from one sample site belong to the same population, revealed significant differences between sexes in three cases: BIK (Nm=14, Nf=12; OarCP26 χ2=6.03, d.f.=1, P=0.0425; OarAE129 χ2=17.86, d.f.=2, P=0.0005), BÖB (Nm=11, Nf=10; ILSTS058: χ2=9.10, d.f.=2, P=0.0424) and MÜK (Nm=7, Nf=7; OarAE129: χ2=9.29, d.f.=1, P=0.0093). Limited availability of undenatured frozen tissue samples prohibited the separate analysis of allozyme data from males and females.
Population isolation
Certain population samples, chiefly those from more or less isolated sites, were distinguished: remarkably low microsatellite variability (see Table 3) characterized the roe deer from BR2 (He and A.D.), RÜG (A.D.) and PAD (P, He and A.D.). The allele frequencies of primer locus OarAE129 differed markedly from those of neighbouring sites in the sample from city park BR2, the nature reserve BIE, the island of Rügen, VOS and PAD. The PCA of allele frequencies separated roe deer from BR2 and MÜN from the remaining samples (Fig. 2). The separation is obvious from PC2, which is chiefly determined by the allele frequencies of the OarCP26 locus (respective weights: allele 134: 74.9%, allele 136: 87.0%, allele 138: 57.1%). High mean genetic distances (Table 3), compared with the mean value calculated from microsatellite data (D=0.1638), were observed for the samples BR2 (D=0.5074), RÜG (D=0.2275), BIE (D=0.2202), and MÜN (D= 0.2254). An extremely high mean microsatellite fixation index, of GST=0.1694, was observed only for BR2. For allozymes, the sample BIE showed a markedly higher mean value of GST=0.172 than did the others, with values ranging from GST=0.057 to GST=0.098 (Table 4). High allozymic Nei distances distinguished the roe deer from BIE (D=0.1232) from the other populations (distances: D=0.0197 to D=0.0678). PCA of allozymic allele frequencies likewise separated the BIE sample (see Fig. 2), mainly due to the alleles at the Pgm-1* locus, whose weights were 94.9% each (PC 2), followed by the alleles of Me-2*, with a weight of 83.7%. The frequency of allele Pgm-1*-16, p=0.900, was much higher at BIE than at all other sample sites (p=0.000 to p=0.125). The allele Me-2* 123 was more frequent at BIE (p=0.682) than in all other samples (p=0.156 to p=0.523).
In order to test if GST-fixation indices, or mean Nei distances between samples, correlated with urbanization, microsatellite data for six different geographical groupings of population samples were compared, containing sample sites separated by about 60 km each (except for the grouping BR1 and BR2, separated by only 4 km). The GST-values and Nei distances varied markedly between the groups: BR1 and BR2: GST= 0.2121, mean D=0.6865; HOR and AHR: GST= 0.0328, mean D=0.0862; UCK and SCH: GST= 0.0000, mean D=0.0000; BIK and WES: GST=0.0565, mean D=0.1528; KÜH, BIE and MÜK: GST=0.0983, mean D=0.2429; OBE, TRE and GEF: GST=0.0311, mean D=0.0721. The fixation indices and genetic distances correlated significantly with urbanization in the habitats of roe deer (Spearman rank test for both GST and D: N=6, R=0.9856, t(n − 2)=11.662, P=0.0003, see Fig. 3), which was calculated from a computer satellite atlas (D-Sat, Scout Systems GmbH, München 1998) by quantifying the percentage areas which houses, villages and streets occupied within a square of 60 × 60 km2 around the sample sites.
The correlation (Spearman rank test for both GST and D: N=6, R=0.9856, t (n−2)=11.662, P=0.0003) between mean genetic distances (D) and fixation indices (GST) of local groups of roe deer populations (separated by about 60 km or less) with the percentage of the urbanized area, including villages and roads within a perimeter of about 60 × 60 km2. Population groups are: 1, BR1 and BR2; 2, KÜH, BIE and MÜK; 3, BIK and WES; 4, HOR and AHR; 5, OBE, TRE and GEF; and 6, SCH and UCK.
Discussion
Genetic variability
From 57 bovine and caprine microsatellite primers tested in four animals, 32 (56%) amplified roe deer DNA and nine of these (28%) produced polymorphic microsatellite markers in roe deer, with an average of eight alleles. Allelic diversity seems considerable higher than in other cervids (e.g. Wilson et al., 1997; Marshall et al., 1998; Pemberton & Slate, 1998). However, such comparisons may be misleading when markers of different variability are selected.
The roe deer studied by us seem to exhibit similar allozyme variability measures as did the roe deer from Switzerland, Austria, Slovenia, Slovakia, Bulgaria, France, Italy, and England studied by Hartl & Reimoser (1988), Hartl et al. (1993), Lorenzini et al. (1993) and Hewison (1995). Clearly, the overall variability measures of this investigation surpass those reported by previous studies (ranging from P=0.178 to P=0.133, and He=0.059 to He=0.032), but this is because we screened only a few of the proteins known to be monomorphic in roe deer.
Population structure
Principle component analysis of allele frequencies, among-sample genetic distances and fixation indices of both microsatellite and allozyme data confirmed that there is a genetically rather homogeneous roe deer population throughout Central Europe (cf. Fig. 1). There is no evidence from recent taxonomic revisions that our study areas were the home of different roe deer subspecies or contained a contact belt of populations having expanded from different Pleistocene refugia (Lehman & Sägesser, 1986; Stubbe, 1990). Thus, our genetic data agree with the taxonomy and phylogeography of Central European roe deer. Allozyme and microsatellite differentiation did not correlate with the geographical distances between local samples also; thus isolation-by-distance is not evident in the study area.
Although significant regional differentiation has not been found, allele frequencies, even between adjacent population samples, still showed a small-scale scatter. Hierarchical gene diversity analysis confirmed a low level of population subdivision, predominantly at the local scale. In view of the roe deer’s phylogeographic homogeneity throughout Central Europe, this scatter might be explained by ecological and behavioural determinants specific to individual populations. The general philopatry of roe deer, characterized by short dispersal distances of usually less than 5 km, and small home ranges, which may be stable for a lifetime (Stubbe, 1990), probably support small-scale genetic differentiation. Moreover, the translocation of roe deer for hunting purposes, or a temporal rather than spatial stratification of genotype distribution could be considered. However, there are no indications that our sample populations have been affected by translocations, except for the Amsterdamse WaterleidingDuinen (cf. above). Few cases of roe deer introductions or translocations for hunting purposes have been reported from Germany, including a very limited number of animals being introduced mainly in the beginning 20th century (Beninde, 1941; Niethammer, 1963; Stubbe, 1990). Beninde (1941) recorded only four successful translocations out of 32 attempts comprising only four to five specimens each; none of these were close to our collection sites. Considering the abundance of roe deer throughout Central Europe, with an annual German hunting bag of just over one million specimens, a few successfully translocated specimens cannot have significantly altered the natural genetic population patterns.
Social system and genetic differentiation
When microsatellite data were analysed separately for bucks and does, an (insignificantly) lower variability was observed in females (P=1.000, He=0.579 and A.D.=3.976 in males and P=0.991, He=0.554 and A.D.=3.707 in females). Mean genetic distances between local subsamples of females surpassed those between male subsamples almost significantly (mean D=0.1377 in males against D=0.1807 in females). The female among-population variability (GST= 0.1034) surpassed the male value (GST=0.0695) too. Females exhibited a higher among-sample differentiation (94.5% of total differentiation) at the small, local scale than did males (86.7%). Moreover, assignment indices indicated a higher share of genetically distinctive males in local samples. These results match the sex bias of roe deer dispersal: males disperse further than females, and occupy larger home ranges (Ellenberg, 1978; Stubbe, 1990; Kurt, 1991). Additionally, twice as many males as females disperse (Ellenberg, 1978). Male dispersal increases with population density and resource scarcity, promoted by the presence of adult males (Strandgaard, 1972; Hewison & Gaillard, 1996), whereas conversely female dispersal decreases when resource competition in neighbouring clans is high (Ellenberg, 1978). In conclusion, our population samples may have contained genetically more heterogeneous male subsamples, based on more dispersers than the female subsamples. Sex-biased dispersal may also explain the significant allele frequency differences between males and females in three (BIK, BÖB, MÜK) of the 13 population samples.
This effect might be generally typical for samples of species with sex-biased (i.e. male-biased) dispersal, but it has been considered only rarely in sampling schemes for mapping genetic variation. Schreiber et al. (1994) observed the influence of sex-biased dispersal on the genotype and frequency distribution of the transferrin polymorphism in red deer. In their study population, mean dispersal distances of 19 km in males and 2.55 km in females had produced a patchy spatial mosaic of transferrin genotypes, and a Wahlund effect (heterozygote deficiency) within a forested hunting reserve of 2600 hectares of continuous, unrestricted habitat. The sex bias in dispersal distances appears to be lower in roe deer than in red deer: From raw data of Ellenberg (1978) we estimated mean dispersal distances of 2.61 km for male, and of 2.28 km for female roe deer of two years of age, or more. Heterozygote deficiency was confined to single microsatellites in only three population samples. GIS values of microsatellites and allozymes (Tables 4 and 5) did not indicate a general heterozygote deficiency either. Consequently, sex-biased dispersal does not seem to cause more than marginal Wahlund effects in roe deer, which is thus different from red deer (Schreiber et al., 1994).
Possibly, sex-biased dispersal might produce sex-specific population genetic patterns in white-tailed deer (Odocoileus virginianus), too. Manlove et al. (1975) and Ramsey et al. (1979) found genotype frequencies to differ between males and females, but Ramsey et al. (1979) attributed these differences to sex-different selection pressures in the social hierarchy. Purdue et al. (2000) found differently fixed mtDNA-haplotypes between white-tailed deer populations separated by only 50 km, indicating different maternal lineages, while biparently inherited allozymes revealed no differentiation. Purdue et al. (2000) explain this difference with the male-biased dispersal of the white-tailed deer.
Habitat fragmentation
Urbanization correlated highly significantly with the mean genetic distances among adjacent population samples, and with their fixation indices. This correlation followed an exponential function (Fig. 3). Specifically, the city park population in Bremen (BR2) displayed a remarkably reduced microsatellite variability of 44.3 and 39.4% of the mean allelic diversity and expected heterozygosity of all samples, and the highest mean D- and GST-values, when compared to the remaining populations; PCA supported this population’s genetic distinction. The approximately 70 roe deer from this urban site have been effectively isolated for 15–20 years (H. Tempelmann, personal communication). In the sample RÜG, isolation effects were evident only from reduced allelic diversity (31.9% of the mean) and from the mean Nei’s distances. The large size of Rügen Island (927 km2) and the observation of occasional exchange of animals with the mainland through a 1–2 km broad seaway (A. Siefke, personal communication) probably decrease the stringency of isolation. Fakler (1999) described reduced RAPD-DNA variability of badgers (Meles meles) from Rügen Island, compared with mainland badgers. Roe deer from the nature reserve Biedensand (BIE), comprising 525 ha, were distinguished by elevated genetic distances (microsatellites and allozymes), fixation indices (allozymes), and by PCA (allozymes). There are only about 50 hectares of forest in this nature reserve, surrounded by flood plain meadows, fields, and reed beds, and large parts of it are annually flooded for several weeks in spring. The genetic distinction of the roe deer from BR2, RÜG and BIE is also supported by markedly deviating microsatellite and allozyme (only BIE) allele frequencies, relative to adjacent sites (Tables 3 and 4). The reduced genetic variability of roe deer from PAD (P=0.750, He=0.380, and A.D.=2.50) does not correlate with any recognizable habitat isolation.
The documented effects of habitat isolation need not necessarily imply the consequences of prolonged inbreeding. Rather, a founder effect might have led to initially only few genetic lineages. If so, their reduced variability measures are consistent with the intra-lineage variation of single demes of larger, non-isolated populations.
Microsatellites and allozymes as population markers
The allozyme and microsatellite analyses supplied non-identical, though largely overlapping, information. Microsatellites exhibited far higher allelic diversity and heterozygosity than did allozymes. Allele numbers ranged between two to 16 per microsatellite locus in roe deer, but the 10 allozyme loci investigated in six population samples included just six bi- or triallelic polymorphisms. In red deer (Cervus elaphus) Marshall et al. (1998) found six to 13 alleles in nine microsatellites, whereas only two alleles were observed in three polymorphic proteins.
In the present microsatellite study, genetic distance values emerged as the most sensitive indicator for detecting population isolation. Allelic diversity, and finally expected heterozygosities and fixation indices, came next. Markers with few alleles (e.g. McM507, two alleles) displayed an overall population homogeneity throughout the study area. Isolated populations were detected only by polyallelic markers, such as the seven-allele locus OarAE129. Apparently, the greater microsatellite heterozygosity is less sensitive to mild genetic drift (or Wahlund effects) than are allozymic heterozygosities. Likewise, Barker et al. (1997) found that bottlenecks chiefly increased allozyme genetic distances, whereas in microsatellites their main effect was to reduce allelic diversity. As rare alleles are lost preferentially, heterozygosity and genetic distance values were less affected (Barker et al., 1997). Also in our study, population subdivision was best reflected by fixation indices derived from the more sensitive allozymic heterozygosity measures. Genetic distances derived from microsatellites and from allozymes seem to resolve population structure equally well.
References
Barker, J. S. F., Moore, S. S., Hetzel, D. J. S., Evans, D. et al. (1997). Genetic diversity of Asian water buffalo (Bubalus bubalis): microsatellite variation and a comparison with protein-coding loci. Anim Genet, 28: 103–115.
Beninde, J. (1941). Die Einkreuzung bei Rehwild. Z Jagdk, 3: 57–79.
Colson, I. and Goldstein, D. B. (1999). Evidence for complex mutations at microsatellite loci in Drosophila. Genetics, 152: 617–627.
Dratch, P. A. and Pemberton, J. M. (1992). Application of biochemical genetics to deer management: What the gels tell. In: Brown, R. D. (ed.) Biology of Deer, pp. 367–383. Springer, New York, Berlin.
Ede, A. J., Pierson, C. A. and Crawford, A. M. (1995). Ovine microsatellites at the OarCP9, OarCP16, OarCP20, OarCP21, OarCP23 and OarCP26 loci. Anim Genet, 26: 129–130.
Ellenberg, H. (1978). Zur Populationsökologie des Rehes (Capreolus capreolus L., Cervidae) in Mitteleuropa. Spixiana, 2: München.
Fakler, P. (1999). Genetische Differenzierung des Eurasischen Dachses in Mittel-, Nord- und Nordwest-Europa. PhD Thesis, University of Heidelberg, Germany.
Fakler, P. and Schreiber, A. (1997). Allozyme heterozygosity in two isolated populations of roe deer (Capreolus capreolus) in the Netherlands. Neth J Zool, 47: 1–8.
Favre, L., Balloux, F., Goudet, J. and Perrin, N. (1997). Female-biased dispersal in the monogamous mammal Crocidura russula: evidence from field data and microsatellite patterns. Proc R Soc B, 264: 127–132.
Hartl, G. B. and Reimoser, F. (1988). Biochemical variation in roe deer (Capreolus capreolus L.): are r-strategists among deer genetically less variable than K-strategists? Heredity, 60: 221–227.
Hartl, G. B., Reimoser, F., Willing, R. and Köller, J. (1991). Genetic variability and differentiation in roe deer (Capreolus capreolus L) of Central Europe. Génét Sél Évol, 23: 281–299.
Hartl, G. B., Markov, G., Rubin, A., Findo, S. et al. (1993). Allozyme diversity within and among populations of three ungulate species (Cervus elaphus, Capreolus capreolus, Sus scrofa) of Southeastern and Central Europe. Z Säugetierk, 58: 352–361.
Hewison, A. J. M. (1995). Isozyme variation in roe deer in relation to their population history in Britain. J Zool, Lond, 235: 279–288.
Hewison, A. J. M. and Gaillard, J. M. (1996). Birth-sex ratios and local resource competition in roe deer, Capreolus capreolus. Behav Ecol, 7: 461–464.
Hulme, D. J., Smith, A. J., Silk, J. P., Redwin, J. M. et al. (1995). Polymorphic sheep microsatellites at the McM2, McM131, McM135, McM136, McM140, McM200, McM214, McM373, McM505, McM507 and McM512 loci. Anim Genet, 26: 365–374.
Jaeger, F., Hecht, W. and Herzog, A. (1992). Investigation of mitochondrial DNA from Hessian roe deer Capreolus capreolus. Z Jagdwiss, 38: 26–33.
Kemp, S. J., Hishida, O., Wambugu, J., Rink, A. et al. (1995). A panel of polymorphic bovine, ovine and caprine microsatellite markers. Anim Genet, 26: 299–306.
Kurt, F. (1991). Das Reh in der Kulturlandschaft. Parey, Hamburg, Berlin.
Kurt, F., Hartl, G. B. and Völk, F. (1993). Breeding strategies and genetic variation in European roe deer Capreolus capreolus populations. Acta Theriologica, 38: (Suppl. 2) 187–194.
Launhardt, K., Epplen, C., Epplen, J. T. and Winkler, P. (1998). Amplification of microsatellites adapted from human systems in faecal DNA of wild Hanuman langurs (Presbytis entellus). Electrophoresis, 19: 1356–1361.
Lehman, E. V. O. N. and Sägesser, H. (1986). Capreolus capreolus Linnaeus, 1758 – Reh. In: Niethammer, J. and Krapp, F. (eds) Handbuch der Säugetiere Europas, vol. 2/II. Aula-Verlag, Wiesbaden, 233–268.
Lorenzini, R., Patalano, M., Appolonio, M. and Mazzarone, V. (1993). Genetic variability of roe deer Capreolus capreolus in Italy: electrophoretic survey on populations of different origin. Acta Theriologica, 38: (Suppl. 2), 141–151.
Manlove, M. N., Avise, J. C., Hillestad, H. O., Ramsey, P. R. et al. (1975). Starch gel electrophoresis for the study of population genetics in white-tailed deer. Proc Ann Conf Southeast Assoc Game Fish Comm, 29: 392–403.
Mantel, N. (1967). The detection of disease clustering and a generalized regression approach. Cancer Res, 27: 209–220.
Marshall, T. C., Slate, J., Kruuk, L. E. and Pemberton, J. M. (1998). Statistical confidence for likelihood-based paternity inference in natural populations. Mol Ecol, 7: 639–655.
Mörsch, G. and Leibenguth, F. (1993). DNA fingerprinting of the roe deer Capreolus capreolus L. Comp Biochem Physiol B, 104: 229–233.
Nei, M. (1973). Analysis of gene diversity in subdivided populations. Proc Nat Acad Sci USA, 70: 3321–3323.
Nei, M. (1978). Estimation of average heterozygosity and genetic distance for a small number of individuals. Genetics, 89: 583–590.
Nei, M. and Chesser, R. K. (1983). Estimation of fixation indices and gene diversities. Ann Hum Gen, 47: 253–259.
Niethammer, G. (1963). Die Einbürgerung von Säugetieren und Vögeln in Europa. Verlag Paul Parey, Hamburg, Berlin.
Pemberton, J. M. and Slate, J. (1998). Genetic studies of wild deer populations: a technical revolution. In: Milne, J. A. (ed.) Recent Developments in Deer Biology. Proc. 3rd. Internat. Congr. Biol. Deer, Macaulay Land Use Research Institute and Moredun Research Institute, 64–72.
Penty, J. M., Henry, H. M., Ede, A. J. and Crawford, A. M. (1993). Ovine microsatellites at the OarAE16, OarAE54, OarAE57, OarAE119 and OarAE129 loci. Anim Genet, 24: 219–219.
Pierson, C. A., Ede, A. J. and Crawford, A. M. (1994). Ovine microsatellites at the OarHH30, OarHH51, OarVH54, OarCP88, OarCP93, OarCP134 loci. Anim Genet, 25: 294–295.
Purdue, J. R., Smith, M. H. and Patton, J. C. (2000). Female philopatry and extreme spatial genetic heterogeneity in white-tailed deer. J Mammal, 81: 179–185.
Ramsey, P. R., Avise, J. C., Smith, M. H. and Urbston, D. F. (1979). Biochemical variation and genetic heterogeneity in South Carolina deer populations. J Wildl Management, 43: 136–142.
Randi, E., Pierpaoli, M. and Danilkin, A. (1998). Mitochondrial DNA polymorphism in populations of Siberian and European roe deer (Capreolus pygargus and Capreolus capreolus). Heredity, 80: 429–437.
Rohlf, F. J. (1998). NTSYS/PC Numerical taxonomy and multivariate analysis system, version 2.02f. Exeter Software, Setauket, NY.
Schreiber, A., Klein, F. and Lang, G. (1994). Transferrin polymorphism of red deer in France: evidence for spatial genetic microstructure of an autocthonous herd. Génét Sél Évol, 26: 187–203.
Sokal, R. R. (1979). Testing statistical significance of geographic variation patterns. Syst Zool, 28: 227–232.
Strandgaard, H. (1972). The roe deer (Capreolus capreolus) population at Kalø and the factors regulating its size. Danish Rev Game Biol, 7: 1–205.
Stubbe, C. (1990). Rehwild, 3rd. edn. Deutscher Landwirtschaftsverlag, Berlin.
Takezaki, N. and Nei, M. (1996). Genetic distances and reconstruction of phylogenetic trees from microsatellite DNA. Genetics, 144: 389–399.
Volmer, K., Hecht, W., Herzog, A. and Faltings, V. (1995). Genetische Untersuchungen an Rehen (C. capreolus L.). Z Jagdwiss, 41: 241–247.
Wehner, J., Müller, H. P. and Kierdorf, H. (1991). Untersuchungen zur genetischen Situation ausgewählter rheinischer Rehwild-Populationen unter besonderer Berücksichtigung isolationsbedingter Veränderungen. Z Jagdwiss, 37: 40–48.
Wiehler, J. and Tiedemann, R. (1998). Phylogeography of the European roe deer Capreolus capreolus as revealed by sequence analysis of the mitochondrial control region. Acta Theriologica, Suppl. 5, 187–197.
Wierdl, M., Dominska, M. and Petes, T. D. (1997). Microsatellite instability in yeast: dependence on the length of the microsatellite. Genetics, 146: 769–779.
Workman, P. L. and Niswander, J. D. (1970). Population studies on Southwestern Indian tribes. Am J Hum Genet, 22: 24–49.
Acknowledgements
Prof. Dr Noelle Cockett (College of Agriculture, Utah State University, U.S.A.) kindly provided test aliquots of microsatellite primers from cattle and sheep. We thank all colleagues named in Table 1 for providing samples, and in particular Dr G. Heidemann (Hamburg), Dr H. Kilias (Hummeltal) and Dr G. Lang (Gries) who organized sampling in North Germany, Bavaria and the Vosges mountains, respectively. Dr W. Lutz (Bonn) provided dried tissue from Düsseldorf and Paderborn. M. W. was supported by a Ph.D. scholarship (Graduiertenförderung Baden-Württemberg).
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Wang, M., Schreiber, A. The impact of habitat fragmentation and social structure on the population genetics of roe deer (Capreolus capreolus L.) in Central Europe. Heredity 86, 703–715 (2001). https://doi.org/10.1046/j.1365-2540.2001.00889.x
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1046/j.1365-2540.2001.00889.x
Keywords
This article is cited by
-
Human-driven genetic differentiation in a managed red deer population
European Journal of Wildlife Research (2021)
-
Connectivity of mule deer (Odocoileus hemionus) populations in a highly fragmented urban landscape
Landscape Ecology (2019)
-
Temporal variation of the genetic diversity of a German red deer population between 1960 and 2012
European Journal of Wildlife Research (2016)
-
A genetic discontinuity in moose (Alces alces) in Alaska corresponds with fenced transportation infrastructure
Conservation Genetics (2015)
-
Roe deer population structure in a highly fragmented landscape
European Journal of Wildlife Research (2014)