Abstract
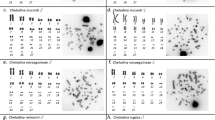
The analysis of C-banding, NOR and fluorochrome staining was carried out in three species of European mussel, Mytilus edulis, M. galloprovincialis and M. trossulus. The results obtained allow us to detect changes in the constitutive heterochromatin within the genus Mytilus. The existence of chromosomal markers permit us to identify and distinguish, at the cytogenetical level, these three types of mussel.
Similar content being viewed by others
Article PDF
References
Ahmed, M, and Sparks, A K. 1970. Chromosome number, structure and autosomal polymorphism in the marine mussels Mytilus edulis and Mytilus californianus. Biol Bull, 138, 1–13.
Cabrero, J, Alche, J D, and Camacho, J P M. 1987. Effects of B chromosomes on the activity of nucleolar organizer regions in the grasshopper Eyprepocnemis plorans: activation of a latent nucleolar organizer region on a B chromosome fused to an autosome. Genome, 29, 116–121.
Camacho, J P M, Cabrero, J, and Viseras, E. 1981. C-heterochromatin variation in the genus Eumigus (Orthoptera: Pamphagoidea). Genetica, 56, 185–188.
Caspersson, T, Faber, S, Foley, G E, Kudynowski, J, Modest, E J, Simonsson, F, Wagh, U, and Zech, L. 1968. Chemical differentiation along metaphase chromosomes. Exp Cell Res, 49, 219–222.
Dixon, D R, and Flavell, N. 1986. A comparative study of the chromosomes of Mytilus edulis and Mytilus galloprovincialis. J Mar Biol Ass UK, 66, 219–228.
Dixon, D R, and McFadzen, I R B. 1987. Heterochromatin in the interphase nuclei of the common mussel Mytilus edulis L. J Exp Mar Biol Ecol, 112, 1–9.
Dixon, D R, McFadzen, I R B, and Sisley, K. 1986. Heterochromatic marker regions (nucleolar organisers) in the chromosomes of the common mussel, Mytilus edulis (Mollusca: Pelecypoda). J Exp Mar Biol Ecol, 97, 205–212.
Fernández-Piqueras, J, Rodríguez Campos, A, Sentís Castaño, C, and Rojo Garcia, E. 1983. Chromosomal location of the active NORs in the Steropleurus martorelli complex. Genetica, 61, 9–12.
Gosling, E M. (ED.) 1992. The Mussel Mytilus. Elsevier Press, Amsterdam.
Howell, W M, and Black, D A. 1980. Controlled silver-staining of nucleolus organizer regions with a protective colloidal developer, a 1-step method. Experientia, 36, 1014–1015.
Ieyama, H, and Inaba, A. 1974. Chromosome numbers of ten species in four families of Pteriomorphia (Bivalvia). Venus, 33, 129–137.
John, B, King, M, Schweizer, D, and Mendelak, M. 1985. Equilocality of heterochromatin distribution and heterochromatin heterogeneity in acridid grasshoppers. Chromosoma, 91, 185–200.
King, M. 1980. C-banding studies on Australian hylid frogs: secondary constriction structure and the concept of euchromatin transformation. Chromosoma, 80, 191–217.
King, M, Honeycutt, R, and Contreras, N. 1986. Chromosomal repatterning in crocodiles: C, G and N-banding and the in situ hybridization of 18S and 26S rRNA cistrons. Genetica, 70, 191–201.
Koehn, R K. 1991. The genetics and taxonomy of species in the genus Mytilus. Aquaculture, 94, 125–145.
Macgregor, H C, and Sessions, S K. 1986. The biological significance of variation in satellite DNA and heterochromatin in newts of the genus Triturus: an evolutionary perspective. Phil Trans R Soc B, 312, 243–259.
Martínez-Expósito, M J, Pasantes, J J, and Méndez, J. 1994. NOR activity in larval and adult mussel Mytilus galloprovincialis L. J Exp Mar Biol Ecol, 94, 155–165.
Martínez-Lage, A, González-Tizón, A, and Méndez, J. 1994. Characterization of different chromatin types in Mytilus galloprovincialis L. after C-banding, fluorochrome and restriction endonuclease treatments. Heredity, 72, 242–249.
Mayr, B, Schweizer, D, Mendelak, M, Krutzler, J, Schleger, W, Kalat, M, and Auer, H. 1985. Levels of conservation and variation of heterochromatin and nucleolus organizers in the Bovidae. Can J Genet Cytol, 27, 665–682.
Méndez, J, Pasantes, J J, and Martínez-Expósito, M J. 1990. Banding pattern of mussel (Mytilus galloprovincialis) chromosomes induced by 2 × SSC/Giemsa-stain treatment. Mar Biol, 106, 375–377.
Moynihan, E P, and Mahon, G A T. 1983. Quantitative karyotype analysis in the mussel Mytilus edulis L. Aquaculture, 33, 301–309.
Pasantes, J, Martínez-Expósito, M J, Martínez-Lage, A, and Méndez, J. 1990. Chromosomes of Galician mussels. J Moll Stud, 56, 123–126.
Schweizer, D. 1976. Reverse fluorescent chromosome banding with chromomycin and DAPI. Chromosoma, 58, 307–324.
Schweizer, D. 1980. Simultaneous fluorescent staining of R bands and specific heterochromatic regions (DA-DAPI bands) in human chromosomes. Cytogenet Cell Genet, 27, 190–193.
Seed, R. 1968. Factors influencing shell shape in the mussel Mytilus edulis. J Mar Biol Ass UK, 48, 561–584.
Skibinski, D O F, and Beardmore, J A. 1979. A genetic study of intergradation between Mytilus edulis and Mytilus galloprovincialis. Experientia, 35, 1442–1444.
Sumner, A T. 1972. A simple technique for demonstrating centromeric heterochromatin. Exp Cell Res, 75, 304–306.
Thiriot-Quiévreux, C, and Ayraud, N. 1982. Les caryotypes de quelques espèces de bivalves et de gastéropodes marins. Mar Biol, 70, 165–172.
Väinölä, R, and Hvilsom, M M. 1991. Genetic divergence and a hybrid zone between Baltic and North Sea Mytilus populations (Mytilidae; Mollusca). Biol J Linn Soc, 43, 127–148.
Verma, R C, and Raina, S N. 1981. Cytogenetics of Crotolaria. V. Supernumerary nucleoli in C. agatiflora (Leguminosae). Genetica, 56, 75–80.
Yosida, T H, and Sagai, T. 1975. Variation of C-bands in several subspecies of Rattus rattus. Chromosoma, 50, 283–300.
Author information
Authors and Affiliations
Rights and permissions
About this article
Cite this article
Martínez-Lage, A., González-Tizón, A. & Méndez, J. Chromosomal markers in three species of the genus Mytilus (Mollusca: Bivalvia). Heredity 74, 369–375 (1995). https://doi.org/10.1038/hdy.1995.55
Received:
Issue Date:
DOI: https://doi.org/10.1038/hdy.1995.55
Keywords
This article is cited by
-
Molecular cytogenetics in the study of repetitive sequences helping to understand the evolution of heterochromatin in Melipona (Hymenoptera, Meliponini)
Genetica (2021)
-
Karyotypic diversification in Mytilus mussels (Bivalvia: Mytilidae) inferred from chromosomal mapping of rRNA and histone gene clusters
BMC Genetics (2014)