Abstract
Consanguineous unions permit the “reunion” of ancestral chromosomal segments in a pattern referred to as “autozygosity,” which is essentially a special form of homozygosity. This has long been exploited as a gene mapping tool because it is relatively easy to track a recessive mutation by the surrounding pattern of homozygous markers. The recent advent of single nucleotide polymorphism microarrays has rapidly replaced the historical use of microsatellites for this purpose. In this review, the author discusses other exciting opportunities offered by this unique arrangement of the human genome that range from pure clinical to functional genomic applications. The emerging field of whole genome sequencing promises to unlock much of the potential of the “autozygome.”
Similar content being viewed by others
Main
Evolution of sexual reproduction is viewed by many biologists as a means to increase genetic diversity and, consequently, fitness of the organism by introducing new genetic material to the offspring, the advantage of which recoups the “twofold cost of males.”1,2 An extension of this concept can be appreciated in the long rooted observation by agricultural societies that outbred crops and animals tend to be healthier than their inbred counterpart.3–5 Biologists attempted to explain this phenomenon in genetic terms since the early 20th century when the concept of “genetic load” was introduced.6 Genetic load refers to the relative decrease in the average fitness of a population with respect to the fitness it would have if all individuals had the genotype with the maximum fitness.7 Theoretically, the adverse effect of inbreeding on fitness can be explained on the basis of increased odds of possessing two copies of an ancestral allele (autozygosity that is essentially a special form of homozygosity) that is mutated, thus unmasking its pathogenic effect recessively, so called “mutational load.”8 Alternatively, autozygosity may simply be affecting reduced fitness by decreasing the “heterozygote advantage,” so called “segregation load.”8 The extension of genetic load to humans sparked an intense debate that continues to this day on the effect of inbreeding on human health.9–13 The clear demonstration of mutational load in many instances has widely shaped how the relationship of autozygosity to human health and disease is viewed.8,14–17 However, as I will show in this review, the “reunion” of pieces of human genome from common ancestors in their offspring also represents unparalleled opportunity to better understand the human genome. I will show that this opportunity is not only limited to Mendelian genetics, although that is the area that witnessed the most fruitful application of autozygosity in human genetics, but also extends to such areas as complex genetics and structural and functional genomics.
MECHANISM AND DIGITAL RENDERING
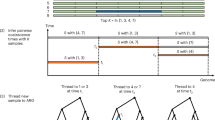
As shown in Figure 1, autozygosity is the appearance of two copies of a DNA segment that are identical by descent. Obviously, homozygosity can also be observed for any segment of DNA without being reflective of a common origin of the two copies, i.e., the two identical segments have been introduced to the genetic pool of the population independently (identical by state [IBS]). The distinction between identical by descent and IBS is critical for certain applications. Mathematically, it can be predicted that the longer the segment is, the lower the probability that the markers tagging that segment have the same calls by chance alone. This probability is the function of both the number of markers that reside on the segment and their degree of informativeness, which is expressed in terms of level of heterozygosity in the population.18 For example, a segment with only one marker that has a heterozygosity score of just 0.5 (one in two individuals taken at random from the target population will be heterozygous) has a 50% probability of being homozygous by chance, i.e., IBS. On the other hand, two markers that each has a heterozygosity level of 0.25 can be expected to be both homozygous by chance with a comparable probability of (1−0.25) × (1−0.25) = 56%. Thus, even markers with low heterozygosity scores can be informative when present in high density. It is this simple concept that gives single nucleotide polymorphisms (SNPs), despite their low heterozygosity score but high frequency in the human genome (1/kb), much higher power as markers, when compared with the more informative but less frequent microsatellites.19–21 If one adds the amenability of SNPs to high throughput genotyping, it becomes clear why SNPs are now the preferred choice for markers.22 This automation takes various forms, the most successful of which has been the ability to array a large set of SNPs on a miniaturized glass chip that, made possible by development of a hybridization-based color detection assay, allows the entire set to be genotyped simultaneously.23 Since their initial launch, the density of these chips has increased considerably from 10,000 to 1,000,000 SNPs and while higher coverage may be desired for other applications, a relevant question to this review is how much coverage is needed to capture autozygosity? To answer this question, one has to consider the crossing over events that led to the eventual autozygous block observed in Figure 1. The farther removed the common ancestor is from the parents of the proband, the higher the number of crossing over events that can break the ancestral haplotype (broken stick concept24,25). Thus, higher density is needed to capture an autozygous block when the parents are more distantly related. If we take 5% as an acceptable cutoff for type I error and an average SNP heterozygosity score of 0.35, then one would require 41 homozygous SNPs in a row on a 250,000 SNP platform to call a run of autozygosity reliably ([1−0.35]41 × 250,000 = 0.05).26,27 This translates to approximately 0.5 Mb of DNA. However, this equation assumes that SNPs are independent of one another, an incorrect assumption given the well-established phenomenon of linkage disequilibrium where SNPs that are in close proximity are likely to segregate together.28 Therefore, the 0.5-Mb detection limit represents an inflated sensitivity, and a 1–2 Mb cutoff represents a more realistic sensitivity limit for this particular chip.29
In reality, detecting that stretch of consecutive homozygous SNPs is not always straightforward. The sheer number of SNPs included on the chip makes it inevitable under the most stringent conditions to encounter missed or, more problematically, false genotypes. Therefore, a manual check of the genotypes is not only impractically slow but also error prone if the block of true autozygosity is interrupted by an apparently heterozygous SNP. To address this issue, numerous methods have been devised that take these limitations into account to tolerate missed or falsely heterozygous genotypes within blocks of autozygosity.18–20,30–33 Many such programs are available free of charge and some even offer a web-based interface. The two major manufacturers of SNP chips (Illumina and Affymetrix, CA) acknowledge the growing popularity of autozygosity analysis and provide loss of heterozygosity analysis as part of their software packages. In the author's experience, the longer the block of autozygosity, the higher the concordance between results from these different programs. When one pushes the sensitivity of these algorithms to their limits, the different programs diverge in their calls and a particular stretch of DNA may be called autozygous by one program but not by another.
AUTOZYGOSITY AND HUMAN POPULATIONS
The common saying that “all humans are related” is probably justified by the extremely limited number of founders to whom contemporary human populations trace their ancestry. However, the numerous generations that separate us today from our common ancestors have allowed a sufficient amount of reshuffling of the chromosomes and introduction of enough mutations to abolish any meaningful autozygosity that can be traced all the way to the era of early humans. On the other hand, the very limited migrational activity of humans until recently, and the creation of many bottlenecks throughout the history of humanity by famines, warfare, and epidemics, have effectively reduced the mating pool, such that a more recent common ancestry can be expected for many of today's human populations.23 Indeed, this prediction was confirmed first in the Centre d'Etude du Polymorphisme Humain panel by microsatellites, and since the advent of SNP chip-based genotyping, a number of other studies followed that clearly demonstrate relatively frequent runs of autozygosity in populations that are viewed as outbred and in which consanguinity is widely unfavored or even outlawed.34–36 Several studies have confirmed that these runs are unlikely to be explained on the basis of uniparental disomy or deletions.37,38 Interestingly, this trend seems to be decreasing in more recent generations, consistent with the changing demographics of the reproductive pool both in size and diversity.39
However, what about inbred populations? Consanguinity is estimated to be practiced by 10% of the world's population.40 The most common form of consanguineous unions involves third-degree relatives (first cousins), which is widely practiced in the Middle East, North and Sub-Saharan Africa, Indian Subcontinent, and Brazil, whereas uncle-niece (second-degree relatives) is largely limited to certain communities in India where it is still legal.13 In these inbred populations, the average population inbreeding coefficient (percentage of the genome that is autozygous) is considerably higher than outbred populations.8,41 The difference is orders of magnitude higher than even in the Finnish population, a prime example of a bottleneck where the population of around 5 million can trace their ancestry to a small number of founders in the not too distant past but where consanguinity is uncommon.42
AUTOZYGOSITY AND HUMAN DISEASES
Autosomal recessive disorders
The a priori probability of both parents being carriers of an autosomal recessive mutation simply depends on the carrier frequency in the general population, which tends to be very low because of the selective pressure against these usually lethal or highly morbid alleles. However, in the consanguineous setting, the probability of one parent being a carrier is not independent of the other parent's. Indeed, first cousin parents share 1/8th of their genome on average (coefficient of relationship), which substantially increases the risk of autozygosity for a disease allele and the occurrence of recessive disorders in the offspring as a consequence. As a general rule, the rarer the recessive condition is, the more likely it is to be caused by autozygous rather than compound heterozygous mutations.43 This founder effect can manifest in two forms, one in which the founder is many generations removed (often referred to as the founder effect in population genetics) and another in which the founder is an immediate grandparent. Conceivably, consanguinity can unmask several rare recessive alleles for a given gene provided the mutation rate of the gene in question allows for their existence; otherwise, only one ancient founder mutation can be observed as is the case for the so-called Finnish diseases.44 Indeed, we have shown that in highly consanguineous populations, consanguinity plays a more important role for many of the rare diseases than the founder effect, and this leads to unexpected allelic heterogeneity even in genetic isolates.45 In both instances, i.e., allelic heterogeneity unmasked by consanguinity and allelic homogeneity caused by founder effect, autozygosity is the underlying mechanism, although the size of the autozygous block will vary considerably (unless the ancient founder mutation is observed in a consanguineous union).24–25,41
Complex disorders
The role played by autozygosity in more complex disorders is far less established than for simple Mendelian disorders.40 One area that was extensively studied in the past is whether autozygosity (as inferred by the level of consanguinity) is a risk factor for birth defects and the evidence is overwhelmingly in support of that and a doubling of the baseline risk is routinely quoted in prenatal genetic counseling sessions involving consanguineous couples.40,46 It is less clear, however, whether this increased risk is attributed to rare autosomal recessive forms of these defects or whether it represents unmasking of recessive alleles that merely reduce the threshold of occurrence of these defects under the multifactorial genetic load model.13 This uncertainty is similarly encountered when one interprets data on prereproductive mortality rates that are convincingly higher in offspring of consanguineous unions in a pattern compatible with the degree of consanguinity.8,15,47 Interestingly, this detrimental effect of autozygosity seems to be less pronounced in populations where consanguinity is common.8 It is possible that the attributed risk is lower in these populations because of the high baseline mortality due to many other nongenetic factors that act as confounding variables. There is also some debate as to whether populations that have been practicing consanguinity for many generations experience less adverse outcome of consanguinity compared with those with a more recent history of consanguinity, the hypothesis being that this could be the result of natural selection against detrimental alleles (allele purge) over an extended period of time.10,15,48–52
What about disorders that are usually adult in onset and so are under less negative selection pressure? It has been suggested that increased autozygosity results in increased risk not only for autosomal recessive diseases but also for such common disorders as diabetes, hypertension, cancer, and even susceptibility to infections.53–58 It is reasonable to speculate that homozygosity for weak recessive alleles can increase the predisposition to common diseases; after all, many of the risk SNPs identified on genome-wide associations studies (GWAS) have been shown to exert their risk more strongly when present in two copies, so in that sense the role of autozygosity is compatible with the “common variant common disease” hypothesis.40,59 Then again, it is also compatible with the increasingly popular “rare variant common disease” hypothesis, which is proposed as an alternative (not mutually exclusive) to explain the missing heritability of many of the common diseases that GWAS failed to uncover (the so-called dark matter of the genome).60 Autozygosity clearly has the potential to unmask those rare but high-risk alleles in a way similar to but less dramatic than Mendelian recessive diseases. As attractive as these speculations may sound, the evidence is largely lacking for the causal link between autozygosity and common diseases, and it is only recently that actual autozygosity is being tested instead of using epidemiological surveys with consanguinity as a proxy (see later).40
AUTOZYGOSITY AND ANNOTATION OF HUMAN GENOME
Disease gene mapping
In no other area of genetics is the role of autozygosity more prominent than in mapping disease genes where it has served as the most successful mapping strategy in the recent history of human genetics.61 This method was first proposed by Smith and later verified by the convincing simulation published in the landmark article by Lander and Botstein.62,63 Historically, the method has faced several challenges: finding a consanguineous family with the disease in question, observing enough recombination events within the family to narrow the autozygous region surrounding the mutation, and prioritizing candidate genes within that region for sequencing. The initial reliance of this strategy on microsatellites greatly limited the number of laboratories that could apply it, and the very low throughput meant that a typical mapping project would take months if not years just to identify the culprit autozygous interval. The advent of SNP chips has completely revolutionized the way these projects are being conducted and, provided the appropriate family pedigree is available, a similar project can now be expected to be completed in weeks. Additionally, the high density of the genotypic data and their easy amenability to graphical rendering using any of the many software packages that are currently available (see earlier) means that a minimal linkage interval can often be identified by simple visual inspection of the genotypes on the screen, which makes logarithm of odds score calculation using linkage software often unnecessary and indeed redundant.22 It is important to note here that the ease with which the pattern of autozygosity between several affected members of a family can be compared with deduce the minimal shared autozygosity can also be extended to involve affected members from different families as long as the disease is genetically homogeneous. This has proven successful in the mapping of the critical interval for SCARB2 using three different patients from three unrelated families each with a different mutation.64 Using the same approach, it is also possible to narrow a previously published linkage interval with one single affected patient to identify the disease gene.65 It is worthwhile to highlight several pitfalls nonetheless. First, although quite unlikely, it is possible for an extended consanguineous family to harbor mutations in two or more different genes giving rise to the same phenotype, particularly when the phenotype is known to display genetic heterogeneity.66–72 Second, apparently shared autozygous blocks may in fact be IBS, which is particularly problematic when dealing with smaller intervals because the probability of sharing two haplotypes by chance is inversely correlated with their lengths (see earlier). Finally, the number of shared autozygous blocks between the different members of a given family is a function of the randomness of the crossing over events and their frequency. Although their randomness may not be predicted, the number of crossing over events correlates with the number of meiotic events separating the patient from the shared parental ancestor.73 Our successful mapping of FKBP10 as a novel disease gene for Bruck syndrome was based on just two brothers whose parents were very distant cousins that allowed us to easily identify only one small area of autozygous overlap between them.74 On the other hand, higher degrees of autozygosity than that would be expected based simply on the degree of relatedness of the parents is frequently encountered because often extensive background inbreeding is not reflected in abbreviated pedigrees. As a consequence, more blocks of autozygosity will be shared between the affected members by chance alone without being truly linked to this disease locus, which complicates the analysis and reduces the significance of the logarithm of odds score calculations that are based on the limited pedigree information.20,75,76 An important bottleneck remains, however, and that is sequencing. Our incomplete understanding of human biology is clearly reflected in how often the choice of a “good candidate” from the list of tens to hundreds of genes typically contained in the critical interval is wrong. Various tools have been devised to help researchers make their best educated guess by using data mining on protein-protein networks and expression data, but their usefulness in real life is limited.77–79 An exciting emerging alternative is next-generation sequencing based on massively parallel technologies, currently available on various platforms.80 Various approaches have been implemented successfully in this regard. One such approach involves the capture of the critical autozygosity interval by solid phase or fluid phase hybridization followed by massively parallel sequencing.81 This attractive option, which has proven useful recently in the identification of several disease genes, provides a means for highly focused analysis of the culprit locus with significantly lower demand on complex bioinformatics tools that are required to sift through variants encountered with genome-wide sequencing.81–84 The attractiveness of this option is offset by the time-consuming process of capture, currently weeks at best, and is challenged by the precipitous decreasing cost of nontargeted whole genome sequencing, which has been shown recently to identify Mendelian mutations effectively, although the need for sophisticated bioinformatic analysis remains a formidable challenge.85,86 We are currently investigating the potential to combine the best of autozygosity mapping and whole genome sequencing to show if there is a clear advantage in knowing the critical autozygosity interval before whole genome sequencing. In this approach, one only needs to reassemble the short reads generated by next-generation sequencing on a small reference “backbone” that is essentially the chromosomal region corresponding to the critical autozygosity interval, which greatly reduces the demand on bioinformatics. This may seem inefficient, but it is important to note that because the mutations are always going to be homozygous, the level of sequencing (depth of coverage) is so low theoretically (5X) that it does become an economically attractive option compared with the capture method or whole genome high-depth sequencing. Exome sequencing that essentially entails the capture of all known exons followed by next-generation sequencing is another method that has been devised and proven successful.87,88 However, the need for capture, the huge amount of variants that need to be filtered, and the choice to not screen for intronic mutations are factors that will probably keep autozygosity mapping followed by low-depth whole genome sequencing as an attractive alternative. Of course, this does not apply to compound heterozygous or to dominant heterozygous mutations.
I have recently highlighted the clinical utility of autozygosity mapping in this journal, and the interested reader is referred to that article in which a strong case is made for the time and cost-saving benefits that are garnered by the implementation of SNP chip-based genotyping followed by determination of the pattern of autozygosity.89 Such an approach can obviate the need for specialized biochemical testing in the setting of metabolic disorders, minimize the need for sequencing multiple genes in the setting of genetically heterogeneous conditions, and uncover unsuspected deep intronic mutations. Failure to find the suspected gene for the autosomal recessive condition in question within a run of autozygosity (when parents are related) is also beneficial because it excludes this gene and obviates the need for its sequencing. It will also alert the clinician to revisit the initial diagnosis, but if the clinical diagnosis is certain, then it raises the possibility of potentially formerly unsuspected or additional genetic heterogeneity.89
Mapping disease genes for complex disorders is more problematic. Shortly after it was proposed that runs of autozygosity are more common than previously thought in outbred populations, enthusiasm built quickly on the utility of this approach to map genes for such common diseases as diabetes, Alzheimer, autism, etc.35,36,90 In theory, this method should work for both the “common variant common disease” model, where autozygosity for the risk allele should enhance its penetrance, and the “rare variant common disease” model where these rare recessive variants with major effect largely come into view by virtue of autozygosity. In reality, autozygosity holds more promise with the latter because the statistical burden of proof with the discovery of common variants by autozygosity is as yet so complex that it offers little advantage over the time-tested GWAS. If experience is any guide in this regard, the success story of autozygosity mapping in autism is a telltale example.91 In fact, this may well be the only true success story to date in the utility of autozygosity mapping in complex disorders because for other phenotypes the picture is much less clear. For instance, there was a great deal of enthusiasm that accompanied the finding that autozygosity may be associated with increased risk of cancer and colorectal cancer (CRC) in particular.92–94 This common cancer is known to have Mendelian forms usually in a dominant pattern, e.g., familial adenomatous polyposis and hereditary nonpolyposis CRC. MYH mutations are a notable exception because they exert their pathogenesis recessively to cause a uniquely recessive form of hereditary nonpolyposis CRC. Because MYH was uncovered by examining patients with CRC with autozygous blocks that overlap with that locus, it was only conceivable that autozygosity in patients with CRC may uncover other attractive recessively acting risk loci. Subsequent studies, however, have challenged the conclusions made by the original article, and it now seems unlikely that autozygosity is an important risk factor for CRC.95 A similar conclusion was also reached by a large study of autozygosity in pediatric leukemia.26 Two points warrant clarification. First, these negative data do not of course rule out the possibility of recessively acting Mendelian or near-Mendelian alleles in the pathogenesis of disease in some patients. However, they do suggest that this approach is unlikely to uncover them by simply comparing patients and normal controls because such recessive alleles are likely to be very rare, and a more traditional pedigree study of families with significant clustering has a better chance of success. This is in fact how MYH and, more recently, the cancer syndrome, in which multiple pediatric malignancies and café au lait spots are associated with homozygosity for a number of mismatch repair genes, were identified.96,97 It is also possible that the contribution of autozygosity may differ from one cancer to another. The next few years will be critical in defining the role of autozygosity in common diseases because until recently this link was largely based on epidemiological data that examined consanguinity as a proxy to autozygosity. Therefore, the current trend of directly dissecting the “autozygome” in the context of these disorders holds some promise.
DNA dispensability
The finding that human DNA is not uniformly diploid throughout its length is arguably one of the most important genomic discoveries since the completion of the Human Genome Project.98 Widespread copy number alteration results in segments of genomic DNA being duplicated and others being deleted. Importantly, a significant percentage of these copy number variable regions overlap with genes, and this has led to the correct assumption early on that copy number variants (CNVs) are likely to play a role in human health and disease.99 The cataloging of CNVs revealed the extremely interesting, and perhaps unexpected, finding that deletion of CNVs are not limited to hemizygosity, but in fact nullizygous deletions are also observed.100 This raises the intriguing question as to what parts of human DNA are tolerated in the nullizygous state, i.e., are dispensable? Another relevant question that follows is which of the “benign” hemizygous CNVs can be tolerated in the nullizygous state? The latter has been brought into focus by the finding that nullizygosity for some “benign” CNVs does indeed result in adverse clinical outcome akin to autosomal recessive inheritance.101 I argue that the answer to both questions can be pursued quite efficiently with an autozygosity-based approach. The approach is straightforward: by focusing the CNV analysis on offspring of first cousin parents, who share on average 1/8th of their genome including hemizygous CNVs, there is a much higher probability of encountering nullizygous CNVs that can be observed in blocks of autozygosity compared with offspring of unrelated parents whose chance of sharing the same hemizygous CNV is limited by their frequency in the general population. Another significant advantage of this approach is that one can systematically examine bias against nullizygosity throughout the human genome. In other words, if parents do share a hemizygous CNV and their normal offspring are consistently displaying the hemizygous or wild-type genotype for that locus in deviation from the expected Mendelian ratio, then one can quantify the degree of bias against the occurrence of nullizygosity at that locus. In a proof of concept article, we have recently demonstrated this bias that, surprisingly, was not only limited to genic DNA but also extended to nongenic DNA as well (unpublished data). With the mounting interest in nongenic DNA and its role in human biology, larger scale studies are urgently needed to follow-up on this unexpected result because the analysis of nongenic DNA that is biased against nullizygosity is likely to be critically important for normal human development.
SNP annotation
When a novel sequence variant is identified in a disease gene, it is a common practice to screen an ethnically controlled cohort for the presence of this variant to verify its pathogenicity on a statistical basis. The logic here is that failure to find the variant in 180 normal controls, i.e., 1/180 controls, makes it likely to be pathogenic based on the assumption that a more common variant is unlikely to be disease causing, otherwise the frequency of the disease must be much higher than what the epidemiological evidence suggests. This assumption of course is only appropriate for rare autosomal recessive diseases and is not applicable to the study of common diseases. However, it is not uncommon to encounter the variant in question in the heterozygous state in a very small percentage of the normal cohort, say 1 of the 180 controls. Because the variant is rare, it is less likely to exist in homozygous state in an outbred population. However, I argue that consanguineous populations offer a unique opportunity to identify homozygous sequence variants whose presence in normal controls virtually rules out their involvement in the pathogenesis of the highly penetrant autosomal recessive diseases. In contrast, rare sequence variants that are deposited in SNP databases may in fact be pathogenic in the homozygous state as we have recently shown,102 further emphasizing the clear advantage of sequencing the “autozygome” and depositing the resulting data in public databases.
Mutation dating and frequency calculation
Another exciting utility of autozygosity analysis is the ability to date mutations, which in turn can be very helpful in understanding human history. Several mathematical approaches have been devised that exploit the “broken stick” concept in dating the mutation (see earlier).103,104 In addition, although apparently heterozygous SNPs that interrupt a run of autozygosity are often reflective of a genotyping calling error, some are in fact bona fide mutations. Therefore, sequencing of the autozygome promises to be an efficient method of understanding SNP mutation rate by quantifying these occurrences (Table 1).
CONCLUSION
I have shown in this review the unique opportunities that the presence of autozygous regions of DNA offers. The ease with which these regions can currently be mapped at a genome-wide level has suddenly renewed interest in the study of relationship of autozygosity to human health and disease. The historical role of autozygosity mapping in the study of autosomal recessive disorders can be extrapolated on a clinical basis. Autozygosity offers an attractive means with which the human genome can be annotated functionally by mapping dispensable versus indispensable segments and by revealing variants that are tolerated in the homozygous state. Despite some success in the use of autozygosity as a tool to understand the genetics of complex disorders, significant challenges remain. Creative statistical solutions have to be devised to harness better the potential of autozygosity in this setting. The rapid reduction in cost of next-generation sequencing is likely to help unlock the potential of this approach by fully exploring sequence and copy variants in the autozygome and their contribution to common diseases. No doubt, the next few years promise to shed more light on the autozygome and its role in health and disease.
References
Doncaster CP, Pound GE, Cox SJ . The ecological cost of sex. Nature 2000; 404: 281–285.
Hurst LD, Peck JR . Recent advances in understanding of the evolution and maintenance of sex. Trends Ecol Evol 1996; 11: 46–52.
Crnokrak P, Roff DA . Inbreeding depression in the wild. Heredity 1999; 83: 260–270.
Keller L, Waller D . Inbreeding effects in wild populations. Trends Ecol Evol 2002; 17: 230–241.
Kristensen TN, Sørensen AC, Sorensen D, Pedersen KS, Sørensen JG, Loeschcke V . A test of quantitative genetic theory using Drosophila—effects of inbreeding and rate of inbreeding on heritabilities and variance components. J Evol Biol 2005; 18: 763–770.
Muller HJ . Our load of mutations. Am J Hum Genet 1950; 2: 111–176.
Crow JF . Some possibilities for measuring selection intensities in man. Hum Biol 1958; 30: 1–13.
Khoury MJ, Cohen BH, Chase GA, Diamond EL . An epidemiologic approach to the evaluation of the effect of inbreeding on prereproductive mortality. Am J Epidemiol 1987; 125: 251–262.
Roberts DF . Consanguineous marriages and calculation of the genetic load. Ann Hum Genet 1969; 32: 407–410.
Chakraborty R, Chakravarti A . On consanguineous marriages and the genetic load. Hum Genet 1977; 36: 47–54.
Sanghvi LD . 1. The concept of genetic load: a critique. Am J Hum Genet 1963; 15: 298–309.
Crow JF . 2. The concept of genetic load: a reply. Am J Hum Genet 1963; 15: 310–315.
Khlat M, Khoury M . Inbreeding and diseases: demographic, genetic, and epidemiologic perspectives. Epidemiol Rev 1991; 13: 28–41.
Morton NE . The mutational load due to detrimental genes in man. Am J Hum Genet 1960; 12: 348–364.
Khoury MJ, Cohen BH, Newill CA, Bias W, McKusick VA . Inbreeding and prereproductive mortality in the Old Order Amish. II. Genealogic epidemiology of prereproductive mortality. Am J Epidemiol 1987; 125: 462–472.
Elisbão T, Freire-Maia N . Inbreeding effect on morbidity: II. Analyses of a third survey including and excluding infant-juvenile mortality among Brazilian whites and Negroes. Am J Med Genet 1984; 18: 387–391.
Ruas PM, Freire-Maia N . Inbreeding effect on morbidity: I. Three analyses (one with cousin and sib controls) of two surveys among Brazilian whites and Negroes. Am J Med Genet 1984; 18: 381–386.
Carr IM, Sheridan E, Hayward BE, Markham AF, Bonthron DT . IBDfinder and SNPsetter: tools for pedigree-independent identification of autozygous regions in individuals with recessive inherited disease. Hum Mutat 2009; 30: 960–967.
Polasek O, Hayward C, Bellenguez C, et al. Comparative assessment of methods for estimating individual genome-wide homozygosity-by-descent from human genomic data. BMC Genomics 2010; 11: 139.
Carr IM, Flintoff KJ, Taylor GR, Markham AF, Bonthron DT . Interactive visual analysis of SNP data for rapid autozygosity mapping in consanguineous families. Hum Mutat 2006; 27: 1041–1046.
Evans DM, Cardon LR . Guidelines for genotyping in genomewide linkage studies: single-nucleotide-polymorphism maps versus microsatellite maps. Am J Hum Genet 2004; 75: 687–692.
Gibbs JR, Singleton A . Application of genome-wide single nucleotide polymorphism typing: simple association and beyond. PLoS Genet 2006; 2: e150.
Weber JL . Clinical applications of Genome Polymorphism Scans. Biol Direct 2006; 1: 16.
Barton D, David F . Some notes on ordered random intervals. J R Stat Soc 1956; 18: 79–94.
Macarthur RH . On the relative abundance of bird species. Proc Natl Acad Sci USA 1957; 43: 293–295.
Hosking FJ, Papaemmanuil E, Sheridan E, et al. Genome-wide homozygosity signatures and childhood acute lymphoblastic leukemia risk. Blood 2010; 115: 4472–4477.
Enciso-Mora V, Hosking FJ, Houlston RS . Risk of breast and prostate cancer is not associated with increased homozygosity in outbred populations. Eur J Hum Genet 2010; 18: 909–914.
International HapMap Consortium The International HapMap Project. Nature 2003; 426: 789–796.
Hildebrandt F, Heeringa SF, Rüschendorf F, et al. A systematic approach to mapping recessive disease genes in individuals from outbred populations. PLoS Genet 2009; 5: e1000353.
Seelow D, Schuelke M, Hildebrandt F, Nürnberg P . HomozygosityMapper—an interactive approach to homozygosity mapping. Nucleic Acids Res 2009; 37: W593–W599.
Woods CG, Valente EM, Bond J, Roberts E . A new method for autozygosity mapping using single nucleotide polymorphisms (SNPs) and EXCLUDEAR. J Med Genet 2004; 41: e101.
Schäffer AA . Computing probabilities of homozygosity by descent. Genet Epidemiol 1999; 16: 135–149.
Forshew T, Johnson CA . SCAMP: a spreadsheet to collate autozygosity mapping projects. J Med Genet 2004; 41: e125.
Broman KW, Weber JL . Long homozygous chromosomal segments in reference families from the centre d'Etude du polymorphisme humain. Am J Hum Genet 1999; 65: 1493–1500.
Li LH, Ho SF, Chen CH, et al. Long contiguous stretches of homozygosity in the human genome. Hum Mutat 2006; 27: 1115–1121.
McQuillan R, Leutenegger AL, Abdel-Rahman R, et al. Runs of homozygosity in European populations. Am J Hum Genet 2008; 83: 359–372.
Curtis CD, Likhite VS, McLeod IX, Yates JR, Nardulli AM . Interaction of the tumor metastasis suppressor nonmetastatic protein 23 homologue H1 and estrogen receptor alpha alters estrogen-responsive gene expression. Cancer Res 2007; 67: 10600–10607.
Simon-Sanchez J, Scholz S, Fung HC, et al. Genome-wide SNP assay reveals structural genomic variation, extended homozygosity and cell-line induced alterations in normal individuals. Hum Mol Genet 2007; 16: 1–14.
Nalls MA, Simon-Sanchez J, Gibbs JR, et al. Measures of autozygosity in decline: globalization, urbanization, and its implications for medical genetics. PLoS Genet 2009; 5: e1000415.
Bittles AH, Black ML . Evolution in health and medicine Sackler colloquium: consanguinity, human evolution, and complex diseases. Proc Natl Acad Sci USA 2010; 107( suppl 1): 1779–1786.
Mueller RF, Bishop DT . Autozygosity mapping, complex consanguinity, and autosomal recessive disorders. J Med Genet 1993; 30: 798–799.
Norio R . Finnish Disease Heritage I: characteristics, causes, background. Hum Genet 2003; 112: 441–456.
King RJ, Rotter JI, Motulsky AG Approach to genetic basis of common diseases. In: King RJ, Rotter JI, Motulsky AG (eds) The genetic basis of common diseases, 2nd ed. NY: Oxford University Press, 2002; 1076.
Feingold J . Multiple mutations in a specific gene in a small population. C R Acad Sci III 1998; 321: 553–555.
Aldahmesh MA, Abu-Safieh L, Khan AO, et al. Allelic heterogeneity in inbred populations: the Saudi experience with Alström syndrome as an illustrative example. Am J Med Genet A 2009; 149A: 662–665.
Stoltenberg C, Magnus P, Skrondal A, Lie RT . Consanguinity and recurrence risk of birth defects: a population-based study. Am J Med Genet 1999; 82: 423–428.
Edmond M, De Braekeleer M . Inbreeding effects on prereproductive mortality: a case-control study in Saguenay-Lac-Saint-Jean (Québec, Canada) based on a population registry 1838–1971. Ann Hum Biol 1993; 20: 535–543.
Sanghvi LD . The genetic consequences of inbreeding and outbreeding: the role of natural selection in human evolution. A Burg Wartenstein Symposium. Vol series no. 63.
Centerwall WR, Savarinathan G, Mohan LR, Booshanam V, Zachariah M . Inbreeding patterns in rural south India. Soc Biol 1969; 16: 81–91.
Sanghvi LD . Inbreeding in India. Eugen Q 1966; 13: 291–301.
Ghosh AK . Inbreeding in the Kota of Nilgiri Hills, Madras. Soc Biol 1972; 19: 289–291.
Rao PS, Inbaraj SG . Trends in human reproductive wastage in relation to long-term practice of inbreeding. Ann Hum Genet 1979; 42: 401–413.
Rudan I, Smolej-Narancic N, Campbell H, et al. Inbreeding and the genetic complexity of human hypertension. Genetics 2003; 163: 1011–1021.
Jaber L, Shohat T, Rotter JI, Shohat M . Consanguinity and common adult diseases in Israeli Arab communities. Am J Med Genet 1997; 70: 346–348.
Bener A, Hussain R, Teebi AS . Consanguineous marriages and their effects on common adult diseases: studies from an endogamous population. Med Princ Pract 2007; 16: 262–267.
Rudan I, Rudan D, Campbell H, et al. Inbreeding and risk of late onset complex disease. J Med Genet 2003; 40: 925–932.
Rudan I, Skarić-Jurić T, Smolej-Narancić N, et al. Inbreeding and susceptibility to osteoporosis in Croatian island isolates. Coll Antropol 2004; 28: 585–601.
Campbell H, Carothers AD, Rudan I, et al. Effects of genome-wide heterozygosity on a range of biomedically relevant human quantitative traits. Hum Mol Genet 2007; 16: 233–241.
Schork NJ, Murray SS, Frazer KA, Topol EJ . Common vs. rare allele hypotheses for complex diseases. Curr Opin Genet Dev 2009; 19: 212–219.
Manolio TA, Collins FS, Cox NJ, et al. Finding the missing heritability of complex diseases. Nature 2009; 461: 747–753.
Altshuler D, Daly MJ, Lander ES . Genetic mapping in human disease. Science 2008; 322: 881–888.
Lander ES, Botstein D . Homozygosity mapping: a way to map human recessive traits with the DNA of inbred children. Science 1987; 236: 1567–1570.
Smith CAB . Detection of linkage in human genetics. J R Stat Soc B 1953; 15: 153–192.
Berkovic SF, Dibbens LM, Oshlack A, et al. Array-based gene discovery with three unrelated subjects shows SCARB2/LIMP-2 deficiency causes myoclonus epilepsy and glomerulosclerosis. Am J Hum Genet 2008; 82: 673–684.
Faden M, Al-Zahrani F, Arafah D, Alkuraya FS . Mutation of CANT1 causes Desbuquois dysplasia. Am J Med Genet A 2010; 152A: 1157–1160.
Benayoun L, Spiegel R, Auslender N, et al. Genetic heterogeneity in two consanguineous families segregating early onset retinal degeneration: the pitfalls of homozygosity mapping. Am J Med Genet A 2009; 149A: 650–656.
Lezirovitz K, Pardono E, de Mello Auricchio MT, et al. Unexpected genetic heterogeneity in a large consanguineous Brazilian pedigree presenting deafness. Eur J Hum Genet 2008; 16: 89–96.
Frishberg Y, Ben-Neriah Z, Suvanto M, et al. Misleading findings of homozygosity mapping resulting from three novel mutations in NPHS1 encoding nephrin in a highly inbred community. Genet Med 2007; 9: 180–184.
Ducroq D, Shalev S, Habib A, Munnich A, Kaplan J, Rozet JM . Three different ABCA4 mutations in the same large family with several consanguineous loops affected with autosomal recessive cone-rod dystrophy. Eur J Hum Genet Dec 2006; 14: 1269–1273.
Laurier V, Stoetzel C, Muller J, et al. Pitfalls of homozygosity mapping: an extended consanguineous Bardet-Biedl syndrome family with two mutant genes (BBS2, BBS10), three mutations, but no triallelism. Eur J Hum Genet 2006; 14: 1195–1203.
Miano MG, Jacobson SG, Carothers A, et al. Pitfalls in homozygosity mapping. Am J Hum Genet 2000; 67: 1348–1351.
Pannain S, Weiss RE, Jackson CE, et al. Two different mutations in the thyroid peroxidase gene of a large inbred Amish kindred: power and limits of homozygosity mapping. J Clin Endocrinol Metab 1999; 84: 1061–1071.
Génin E, Todorov AA, Clerget-Darpoux F . Optimization of genome search strategies for homozygosity mapping: influence of marker spacing on power and threshold criteria for identification of candidate regions. Ann Hum Genet 1998; 62: 419–429.
Shaheen R, Al-Owain M, Sakati N, Alzayed ZS, Alkuraya FS . FKBP10 and Bruck syndrome: phenotypic heterogeneity or call for reclassification? Am J Hum Genet 2010; 87: 306–307; author reply 308.
Leutenegger AL, Labalme A, Genin E, et al. Using genomic inbreeding coefficient estimates for homozygosity mapping of rare recessive traits: application to Taybi-Linder syndrome. Am J Hum Genet 2006; 79: 62–66.
Woods CG, Cox J, Springell K, et al. Quantification of homozygosity in consanguineous individuals with autosomal recessive disease. Am J Hum Genet 2006; 78: 889–896.
Tiffin N, Kelso JF, Powell AR, Pan H, Bajic VB, Hide WA . Integration of text- and data-mining using ontologies successfully selects disease gene candidates. Nucleic Acids Res 2005; 33: 1544–1552.
Köhler S, Bauer S, Horn D, Robinson PN . Walking the interactome for prioritization of candidate disease genes. Am J Hum Genet 2008; 82: 949–958.
Chen J, Bardes EE, Aronow BJ, Jegga AG . ToppGene Suite for gene list enrichment analysis and candidate gene prioritization. Nucleic Acids Res 2009; 37: W305–W311.
Shendure J, Ji H . Next-generation DNA sequencing. Nat Biotechnol 2008; 26: 1135–1145.
Mamanova L, Coffey AJ, Scott CE, et al. Target-enrichment strategies for next-generation sequencing. Nat Methods 2010; 7: 111–118.
Volpi L, Roversi G, Colombo EA, et al. Targeted next-generation sequencing appoints c16orf57 as clericuzio-type poikiloderma with neutropenia gene. Am J Hum Genet 2010; 86: 72–76.
Philippe O, Rio M, Carioux A, et al. Combination of linkage mapping and microarray-expression analysis identifies NF-kappaB signaling defect as a cause of autosomal-recessive mental retardation. Am J Hum Genet 2009; 85: 903–908.
Rehman AU, Morell RJ, Belyantseva IA, et al. Targeted capture and next-generation sequencing identifies C9orf75, encoding taperin, as the mutated gene in nonsyndromic deafness DFNB79. Am J Hum Genet 2010; 86: 378–388.
Roach JC, Glusman G, Smit AF, et al. Analysis of genetic inheritance in a family quartet by whole-genome sequencing. Science 2010; 328: 636–639.
Lupski JR, Reid JG, Gonzaga-Jauregui C, et al. Whole-genome sequencing in a patient with Charcot-Marie-Tooth neuropathy. N Engl J Med 2010; 362: 1181–1191.
Ng SB, Buckingham KJ, Lee C, et al. Exome sequencing identifies the cause of a Mendelian disorder. Nat Genet 2010; 42: 30–35.
Ng SB, Turner EH, Robertson PD, et al. Targeted capture and massively parallel sequencing of 12 human exomes. Nature 2009; 461: 272–276.
Alkuraya FS . Homozygosity mapping: one more tool in the clinical geneticist's toolbox. Genet Med Apr 2010; 12: 236–239.
Wang S, Haynes C, Barany F, Ott J . Genome-wide autozygosity mapping in human populations. Genet Epidemiol 2009; 33: 172–180.
Morrow EM, Yoo SY, Flavell SW, et al. Identifying autism loci and genes by tracing recent shared ancestry. Science 2008; 321: 218–223.
Assié G, LaFramboise T, Platzer P, Eng C . Frequency of germline genomic homozygosity associated with cancer cases. JAMA 2008; 299: 1437–1445.
Bacolod MD, Schemmann GS, Wang S, et al. The signatures of autozygosity among patients with colorectal cancer. Cancer Res 2008; 68: 2610–2621.
Bacolod MD, Schemmann GS, Giardina SF, Paty P, Notterman DA, Barany F . Emerging paradigms in cancer genetics: some important findings from high-density single nucleotide polymorphism array studies. Cancer Res 2009; 69: 723–727.
Spain SL, Cazier JB, Houlston R, Carvajal-Carmona L, Tomlinson I . Colorectal cancer risk is not associated with increased levels of homozygosity in a population from the United Kingdom. Cancer Res 2009; 69: 7422–7429.
De Vos M, Hayward BE, Picton S, Sheridan E, Bonthron DT . Novel PMS2 pseudogenes can conceal recessive mutations causing a distinctive childhood cancer syndrome. Am J Hum Genet 2004; 74: 954–964.
Sampson JR, Dolwani S, Jones S, et al. Autosomal recessive colorectal adenomatous polyposis due to inherited mutations of MYH. Lancet 2003; 362: 39–41.
Feuk L, Carson AR, Scherer SW . Structural variation in the human genome. Nat Rev Genet 2006; 7: 85–97.
Zhang F, Gu W, Hurles ME, Lupski JR . Copy number variation in human health, disease, and evolution. Annu Rev Genomics Hum Genet 2009; 10: 451–481.
Itsara A, Cooper GM, Baker C, et al. Population analysis of large copy number variants and hotspots of human genetic disease. Am J Hum Genet 2009; 84: 148–161.
Knijnenburg J, Oberstein SA, Frei K, et al. A homozygous deletion of a normal variation locus in a patient with hearing loss from non-consanguineous parents. J Med Genet 2009; 46: 412–417.
Shamseldin H, Al-Dosari M, Rahbeeni Z, Qari A, Hashem M, Alkuraya FS . Study of consanguineous populations can improve the annotation of SNP databases [published online ahead of print October 27, 2010]. Eur J Med Genet doi: 10.1016/j.ejmg.2010.10.009.
Genin E, Tullio-Pelet A, Begeot F, Lyonnet S, Abel L . Estimating the age of rare disease mutations: the example of Triple-A syndrome. J Med Genet 2004; 41: 445–449.
Rannala B, Bertorelle G . Using linked markers to infer the age of a mutation. Hum Mutat 2001; 18: 87–100.
Acknowledgements
The author would like to thank the William K. Bowes, Jr. Award committee for their generosity and support of medical geneticists. The author would also like to thank the members of his laboratory for their hard work and dedication and for their critical review of this manuscript.
Author information
Authors and Affiliations
Corresponding author
Additional information
Disclosure: The author declares no conflict of interest.
*Each year Genetics in Medicine is honored to publish an invited article by that year's recipient of the William K. Bowes, Jr. Award in Medical Genetics, awarded each year to an individual who has made significant contributions to the field.
Rights and permissions
About this article
Cite this article
Alkuraya, F. Autozygome decoded. Genet Med 12, 765–771 (2010). https://doi.org/10.1097/GIM.0b013e3181fbfcc4
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1097/GIM.0b013e3181fbfcc4
Keywords
This article is cited by
-
A novel homozygous missense substitution p.Thr313Ile in the PDE6B gene underlies autosomal recessive retinitis pigmentosa in a consanguineous Pakistani family
BMC Ophthalmology (2023)
-
Mitochondrial “dysmorphology” in variant classification
Human Genetics (2022)
-
Runs of homozygosity: windows into population history and trait architecture
Nature Reviews Genetics (2018)
-
Novel phenotypes and loci identified through clinical genomics approaches to pediatric cataract
Human Genetics (2017)
-
Clinical genomics expands the morbid genome of intellectual disability and offers a high diagnostic yield
Molecular Psychiatry (2017)