Abstract
A GAA-repeat in the X25 gene is causing Friedreich's ataxia (FRDA), a common neurodegenerative disease and >20% of FRDA patients develop type II diabetes (T2D). Linkage has previously been detected between T2D and chromosome 9p13–q21, the region that harbours the X25 gene, but association studies of this gene in T2D have been contradicting. Here, we examined whether genetic variation in the X25 gene is associated with risk for T2D. The GAA-repeat and 18 single nucleotide polymorphisms (SNPs) covering the X25 gene were genotyped in 220 trios in which the affected offspring had abnormal glucose tolerance. Any nominally significant findings were examined in an independent sample consisting of 523 individuals with T2D and 326 healthy controls. Previously reported results were analysed together with our data using a meta-analysis approach. There was no association between the GAA-repeat and T2D susceptibility in our study, which was supported by the meta-analysis including all previous publications. One SNP (rs2498429), 8.2 kb downstream of X25, was nominally associated with T2D in the trios (P=0.02) and showed a trend of association in the same direction in the case–controls (P=0.08; combined permuted P=0.01). Further analysis showed that the nine-marker haplotype containing the rare allele of rs2498429 was nominally associated with T2D in the trios (P<0.01) as well as in the case–controls (P=0.03). In conclusions, this study excludes a role of genetic variation within the X25 gene, but instead suggests that genetic variation downstream the X25 gene, may increase risk for T2D.
Similar content being viewed by others
Introduction
Type II diabetes (T2D) is a common, multifactorial disorder characterized by chronic hyperglycaemia resulting from pancreatic β-cell dysfunction and insulin resistance.1 Friedreich's ataxia (FRDA, MIM 229300) is a common neurodegenerative disease and about 20% of the FRDA patients are known to develop T2D and 10% have milder disturbances in their glucose metabolism,2 suggesting an increased susceptibility to T2D in these individuals.
FRDA is inherited in an autosomal recessive mode and the mutated gene X25 encoding the frataxin protein is located in a region on chromosome 9q133 that has been suggested to be linked to T2D in several patient materials, including ours.4, 5, 6, 7, 8 Long homozygous trinucleotide repeats, (GAA)>66, in X25 are thought to cause about 98% of all FRDA cases while remaining cases are caused by a combination of point mutations and heterozygous expanded repeats.9 The extended trinucleotide repeats reduce X25 mRNA levels resulting in lower levels of the frataxin protein.3
Frataxin is localized in the inner mitochondrial membrane and expressed in tissues with high metabolic activity such as heart, liver, skeletal muscle and adipose tissue,10 that is, primary target tissues for T2D. The biological function of frataxin is still unknown but it has been suggested to play a role in iron metabolism.11 The identification of a set of genes involved in oxidative phosphorylation whose expression is coordinately decreased in human diabetic muscle suggests importance of mitochondrial genes that could be involved in the pathogenesis of T2D.12
The co-occurrence of glucose metabolism disturbances in FRDA patients, the suggested link between severity of glucose abnormalities and length of the trinucleotide repeat in FRDA13 and the localization of the X25 gene on chromosome 9q13, a region suggested to be linked to T2D,4, 5, 6, 7, 8 have prompted investigations of a possible association between the intronic X25 intermediate repeat and T2D.14, 15, 16, 17, 18, 19 Unfortunately, the results have been inconclusive; therefore we set out to investigate whether genetic variation at the X25 locus increases susceptibility to T2D.
Methods
Patients
All subjects in this study were participants of the Botnia study. 20, 21 The Botnia study was established 1990 to identify the genetic and metabolic factors contributing to the pathogenesis of T2D in families from the western coast of Finland. The collection was subsequently extended to other parts of Finland and Sweden. Subjects were classified into different stages of glucose tolerance based on their fasting and 2-h glucose concentrations.1
Altogether we have genotyped 743 affected individuals and 766 unaffected individuals in this study in a two-layered study design (Table 1). The first patient–material consisted of 660 individuals from 220 parent–offspring trios. The offspring had T2D (n=98), impaired glucose tolerance (IGT) (n=72) or impaired fasting glucose (IFG) (n=50) according to WHO’98 criteria.1 Since samples consisting of trios with DNA from parents available are biased towards patients of young age, we also chose to include prediabetic trios where the offspring was diagnosed with IGT and IFG in our study, since this strategy has proven successful in dissecting association with risk of T2D diabetes previously.22, 23 Only unrelated trios (one trio per family) were used to avoid confounding linkage in our test for association since this chromosomal region (9p13–q21) has already been suggested to be linked to T2D in our population.5
The second, patient material consisted of 849 individuals in a case–control design. This material was used to investigate the genetic variations found to be nominally associated to T2D in the initial trios. We included cases (n=523) with T2D according to WHO’98 guidelines and controls (n=326) with normal glucose tolerance.1
In both samples, individuals with genetically verified MODY, known type I diabetes and/or with fasting C-peptide levels <0.2 nmol/l were excluded. Affected (T2D) offspring had age-at-onset >20 years of age and case–controls had age-at-onset >35 years of age. All subjects gave their informed consent to the study, which was approved by local ethics committees.
Genotyping
A nested PCR was used to genotype the GAA-repeat. The initial PCR step was performed as previously described.13 The second PCR step was performed in a total volume of 20 μl using 5 μl from the initial PCR as template (3 mM MgCl2, 0.1 mM dNTP, 0.6 U Taq, 0.3 μ M GAA2F (5′ Hex), 0.3 μ M GAA2R. A total of 35 cycles of PCR was performed (94°C for 10 s, 59°C for 30 s, 72°C for 45 s) and 0.1 μl of each reaction was loaded and electrophoresed for 30 min on an ABI 3100 Sequencer (Perkin Elmer, Foster City, CA, USA) for fluorescent detection, using GS400HD ROX (Applied Biosystems, Foster City, CA, USA) as an internal standard. The product of GAA2F/GAA2R was used for size determinations.
A total of 40 SNPs in and around the X25 gene were chosen from NCBI dbSNP. Of the 40 SNPs, 10 were genotyped on an ABI3100 using the SnaPshot-technique according to manufacturer's instructions (PE Biosystems, Foster City, CA, US). Reactions were run on an ABI3100 using Gene Scan 120LIZ (Applied Biosystems, Foster City, CA, USA) as size standard. Two independent scorers read all results using GeneMapper™ 2.0.1 (Applied Biosystems, Foster City, CA, USA). In case of Mendelian inconsistencies, the genotyping for the entire trio was repeated. Five SNPs were excluded: one was found monomorphic, one failed in Hardy–Weinberg equilibrium test (P<0.01) and three assays were not working robustly (success rate <85%).
Remaining 30 SNPs were genotyped using matrix-assisted laser desorption/ionization time-of-flight (MALDI-TOF) mass spectrometry (SEQUENOM Inc., San Diego, CA, USA). The resulting mass spectra were analysed using the SpectroTYPER RT 2.0 software (Sequenome Inc., San Diego, CA, USA). Data from the parent–offspring trios were checked for Mendelian segregation using the PEDCHECK software.24
A total of 10 markers were found monomorphic, two failed in Hardy–Weinberg equilibrium test (P<0.01), four had a success rate of less than 85% and one showed excessive Mendel errors. These were subsequently excluded from the study. Four SNPs, which were genotyped on both ABI3100 and Sequenom showed an average genotype discrepancy of 1.4% (SD±0.7).
In total, 18 SNPs met our quality criteria and was used for subsequent analysis. PCR and extension primer sequences for both methods are found as Supplementary Information (web appendix, Supplementary Tables 1 and 2).
Statistical analysis
In order to calculate the number of trios required to evaluate the initial report of association between the X25 intermediate GAA repeat with T2D appropriately, we performed power calculations using the Genetic Power Calculator (http://statgen.iop.kcl.ac.uk/gpc/).25 In these calculations we assumed an intermediate GAA-repeat frequency of ∼30% as reported in Scandinavians15 and an odds ratio (OR) of 3.3 in carriers vs noncarriers, as reported in the initial positive study by Ristow et al.18 We further assumed a T2D disease prevalence of ∼6%. Under these parameters, we estimate that ∼130 trios and ∼150 cases and ∼100 controls would provide 95% power to reject the null hypothesis of no association at α<0.05 under a dominant model. Our 220 parent–offspring trios and 523 cases and 326 controls would provide >99.7% power to reject the null hypothesis of no association at α<0.05, respectively.
To calculate the number of case–controls required to evaluate our nominal association of the rare allele of rs2498429 with T2D susceptibility, we performed power calculations using the Genetic Power Calculator (http://statgen.iop.kcl.ac.uk/gpc/).25 We assumed a minor allele frequency of 6.6% and a genotype relative risk of 2.4, as observed in our trios. It showed that ∼290 cases and ∼180 controls would provide >95% power to reject the null hypothesis of no association at α<0.05. Having 523 cases and 326 controls would provide >99% power to reject the null hypothesis at α<0.05.
Differences in X25 GAA-repeat sizes were analysed either as separate alleles or dichotomized in groups normal (<10 repeats) or intermediate (⩾10<66 repeats) alleles as reported initially by Ristow et al.18 Affected offspring in trios and case–controls were divided into three groups depending on degree of glucose intolerance and analysed for differences in the GAA-repeat using a χ2 test as previously described.18 A meta-analysis of all published data14, 15, 16, 17, 18, 19 for the putative association between the X25 GAA-repeats and T2D diabetes was carried out using the Mantel–Haenszel test (M–H test) for estimating an OR (NCSS 2000, UT, USA).
To evaluate haplotype block patterns over the X25 locus, linkage disequilibrium (LD) between the SNPs was analysed using Haploview 2.05 and D′ values were calculated with 95% confidence intervals (CI).26 Also, data for 40 SNPs (between rs872077 and rs4744826) from the International HapMap project (http://www.hapmap.org/, public data release #11 September 2004) spanning the X25 locus was used as a ‘reference’ for the expected haplotype blocks and LD structure of the region. Five of our 18 SNPs (rs1800651, rs3793451, rs1411676, rs1411675 and rs953588) were the same as in the CEPH reference material from the HapMap data.
Transmission distortion from heterozygous parents were calculated using Transmission Disequilibrium Test (TDT)27 as implemented in Genehunter 2.1 R228 for both the GAA-repeat and for the SNP genotype analysis. Haplotype block transmission and frequency estimations in the trios were calculated using TRANSMIT.29 Haplotype block frequency estimations in the case–controls were calculated using GENECOUNTING.30, 31
All P-values are given as nominal P-values if not stated otherwise. The permuted pointwise P-value for the combined parent–offspring and case–control data for SNP rs2498429 was calculated by randomly transmitting alleles from the parents to the offspring, and permutating case status for the case–control data. This procedure was repeated 10 000 times for each of the two samples creating unique strata from each run. The first stratum from the trios and from the case–controls was combined in an M–H test, and this procedure was repeated for the rest of strata resulting in 10 000 test statistics. We compared our result with the empirical distribution of test statistics obtained from the permutation to achieve a pointwise permuted P-value for the combined data for the rs2498429 SNP.
Results
Assessment of statistical differences of X25 (GAA) repeats allele sizes; distribution between groups with different levels of glucose intolerance
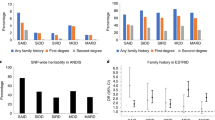
The X25 (GAA)n repeat was genotyped in the 220 parent–offspring trios and case–controls (523 and 326, respectively). Affected offspring were divided into three groups depending on their glucose tolerance (IFG, IGT or T2D). To investigate differences in distribution between normal (<10) and intermediate (⩾10<66) repeats in the three glucose tolerance groups the repeats were considered separately and dichotomized as normal or intermediate as in Ristow et al.18 Normal repeat-sizes were the most common in all three groups. Using a χ2-test neither a significant difference in repeat size distribution between these three groups, nor an association between intermediate trinucleotide repeats and T2D was found (data not shown). Allele distribution in the trios and case–controls are given as supplemental information (web appendix, Supplementary Figure 1a, b). A χ2-test was used to test for differences between normal and intermediate GAA-repeats in the case–controls, no significant differences were found (P=0.8).
Meta-analysis of published studies evaluates a putative association between the X25 trinucleotide repeat and T2D
Hereto, the association between the X25 repeat and T2D have been inconclusive.14, 15, 16, 17, 18, 19 To evaluate this apparently inconsistent data, we performed a meta-analysis32 of all published studies on the putative association of the trinucleotide repeat in the X25 gene and T2D.14, 15, 16, 17, 18, 19 Testing for heterogeneity between all different studies showed significant differences between results, (P<0.0001). This indicates that the meta-analysis cannot be used to provide a summary OR across the studies. When we repeated the heterogeneity test and excluded the initial positive results by Ristow et al, no heterogeneity between the ORs of remaining studies was seen (P=0.5). Thus this initial study was excluded from the meta-analysis, which shows no evidence for an association between the GAA-repeat in the X25 gene and T2D (OR=1.02, 95% CI (0.88–1.19), P=0.8, Figure 1) in the remaining studies.
OR (with 95% CI) for the X25 GAA repeat. For each study the open circle represents the OR for the intermediate allele and the line indicates its 95% CI. The dashed lines indicate the 95% CI for the estimated OR in this study. Data for the studies are presented for patients with normal or intermediate repeat/s for the case–control studies, and for transmission for the familial studies. Data are based on our M–H test calculations, combined OR=1.02, 95% CI (0.88–1.19), P=0.8.
Transmission disequilibrium tests (TDT) and linkage disequilibrium (LD)-mapping of genetic variants in the X25 gene and risk of T2D
We used a two-layered strategy, previously proven successful,32, 33 to evaluate whether there was an association between common genetic variation in and surrounding the X25 gene and T2D. First, to avoid false positives due to population stratification,34 we applied a family-based strategy to test both the GAA-repeat and 18 additional SNPs for association to T2D. One SNP (rs2498429) showed a nominal association with an increased risk of T2D (an overtransmission of the rare allele to affected offspring, P=0.02, Table 2).
We then evaluated the haplotype block structure at this locus and allelic association (LD) between the SNPs was calculated using HaploView v2.05© and D′ values were calculated with 95% CI.26 In total, 16 common haplotypes (frequency >5%, Figure 2a) were identified at this locus distributed into four different regions showing LD (Figure 2a,b). Data currently available at the International HapMap project (www.hapmap.org, release #11 September 2004) confirms, at least partly, our third and fourth haplotype blocks and their frequencies. We then tested whether any of these four blocks contained haplotypes that were associated with T2D. The fourth block, containing nine SNPs, spanning 17 kb, an excess transmission was observed when analysing our data using TRANSMIT.29 The haplotype harbouring the rare allele of rs2498429 was significantly overtransmitted to affected offspring in the trios (χ2=7.34, P<0.01, Figure 2a).
We observed four regions displaying strong LD and showing little evidence of historical recombination. (a) The major haplotype block structure (>5%) combinations with frequencies are shown and LD between the SNPs was analysed using Haploview 2.05 and D′ values were calculated with 95% CI.26 In the fourth block the haplotype containing the rare allele of rs2498429 was found significantly more often than expected in the diabetic offspring (P<0.01) using TRANSMIT and in the case–controls (P=0.03) using GENECOUNTING30, 31 (underlined). Only the haplotype containing the rare rs2498429 variant showed over transmission. (b) D′ plot each square depicts the magnitude of LD for a pair of markers, red colour indicating high D′ and white colour low D′. Single-point TDT27 indicates an overtransmission of rs2498429 SNP to diabetic offspring (permutated P=0.013). The X25 gene is depicted on top of the D′ plot, exons indicated by black boxes.
To evaluate our finding in the initial screening, the SNPs in the fourth haplotype block (Figure 2b) were genotyped in our case–control material. Here, the rare allele of rs2498429 showed a trend of association in the same direction as the initial finding (P=0.08, one-tailed).
A combined analysis (M–H test) of our trio and case–control data sets shows a significant association between rs2498429 and T2D susceptibility (OR=1.66, 95% CI (1.13–2.44), permutated P=0.01).
All major haplotypes as well as frequencies and patterns of LD, were the same as observed in the trios. The overtransmitted haplotype was shown to be significantly associated (permuted P=0.03) in the case–controls when analysing our data using GENECOUNTING.30, 31
Discussion
Our study represents the hereto largest genetic evaluation of the role of the Friedreich's ataxia gene, X25, in T2D susceptibility. Previous studies evaluating the X25 GAA-repeat association to T2D have been conflicting14, 15, 16, 17, 18, 19 and we therefore investigated the association between the GAA-repeat in two different nonoverlapping data sets, using a screening approach previously shown to be successful.32 In addition, we extended our analysis with additional SNP genotyping and haplotype analysis in the trios in an initial screen. The trios were chosen as the screening sample to avoid false-positive signals due to population stratification. Nominally significant findings in the initial screen were further investigated for a possible replication in a case–control sample from the same population.
Genetic association studies have traditionally been hampered by initial jackpot effects and underpowered investigations. One way to increase power and circumvent this is to perform a meta-analysis of all published studies for a given association. This strategy has proven highly efficient confirming the association of the Pro12Ala variant in the PPARγ gene and the E23K variant in the KCNJ11 gene with T2D.32, 33
Evaluating our two sets of samples independently and in a meta-analysis together with all previous published studies (totally including 1724 cases, 948 controls and 375 parent–offspring trios in analysis) failed to detect any association between the GAA-repeat and T2D. Population stratification as the cause of these negative results is unlikely considering the result in our and other family-based studies17 and the large meta-analysis should provide more than enough power to detect true association according to our power calculations. It is more likely that the initial report with a surprisingly high OR for T2D18 represented a jackpot effect and that the other study reporting an association19 was underpowered (supported by their 95% CI, Figure 1). It is worth noticing that the result from the initial study by Ristow et al has not been replicated independently despite showing strong, internally replicated, genomewide P-values. Importantly, this highlights the question of how to interpret genetic data from initial studies and the need for replication in independent populations using well-powered study designs before any far-reached conclusions are drawn.
To investigate whether other common genetic variation in the X25 gene could contribute to the variable association results and predispose to T2D, we studied additional common genetic variation in and around the X25 gene and their haplotypes in our patient material. One SNP (rs2498429, Table 2), located 8.2 kb downstreams of the X25 gene, was found to be nominally associated to T2D in the initial screen of the TDT trios. The rare allele of rs2498429 has an OR of 2.4 in our trios, as compared to an OR of 1.4 in our case–control samples, suggesting that a larger case–control sample would be required for a replication to be significant on its own. However, the trend in the same direction in the case–controls as in the trios provide partial support of an association and the results were further strengthened by the combined analysis, which is significant after permutation.
This SNP was found to be a part of a haplotype block residing in the 3′ UTR region of the X25 gene and extending distally from the gene region, which was confirmed by the HapMap data. Subsequent analysis of these haplotypes showed association between the haplotype containing the rare allele of rs2498429 and T2D in our parent–offspring trios. This finding was further supported by the case–control haplotype data and the association to T2D susceptibility is significant after permutation in our combined analysis.
In the chromosomal region defined by the haplotype block containing rs2498429, no characterized genes are currently known. However, in the defined area there are a number of Expression Sequence Tag (EST) that would be interesting to characterize further and one of these, EST (BM022009), contains the putatively associated SNP (rs2498429) and was recently discovered in a human pancreatic cDNA library.35
In conclusion, our study excludes a role of genetic variation within the X25 gene, but instead suggests that genetic variation downstream the X25 gene, may increase risk for T2D. Replication in larger data sets from different populations will further clarify the potential role for this SNP and the surrounding genetic variation in the risk of T2D.
References
Alberti KG, Zimmet PZ : Definition, diagnosis and classification of diabetes mellitus and its complications. Part 1: diagnosis and classification of diabetes mellitus provisional report of a WHO consultation. Diab Med 1998; 15: 539–553.
Finocchiaro G, Baio G, Micossi P, Pozza G, di Donato S : Glucose metabolism alterations in Friedreich's ataxia. Neurology 1988; 38: 1292–1296.
Campuzano V, Montermini L, Molto MD et al: Friedreich's ataxia: autosomal recessive disease caused by an intronic GAA triplet repeat expansion. Science 1996; 271: 1423–1427.
Hanis CL, Boerwinkle E, Chakraborty R et al: A genome-wide search for human non-insulin-dependent (type 2) diabetes genes reveals a major susceptibility locus on chromosome 2. Nat Genet 1996; 13: 161–166.
Lindgren CM, Mahtani MM, Widen E et al: Genomewide search for type 2 diabetes mellitus susceptibility loci in Finnish families: the Botnia study. Am J Hum Genet 2002; 70: 509–516.
Luo TH, Zhao Y, Li G et al: A genome-wide search for Type II diabetes susceptibility genes in Chinese Hans. Diabetologia 2001; 44: 501–506.
Pratley RE, Thompson DB, Prochazka M et al: An autosomal genomic scan for loci linked to prediabetic phenotypes in Pima Indians. J Clin Invest 1998; 101: 1757–1764.
Imperatore G, Hanson RL, Pettitt DJ, Kobes S, Bennett PH, Knowler WC : Sib-pair linkage analysis for susceptibility genes for microvascular complications among Pima Indians with type 2 diabetes. Pima Diabetes Genes Group. Diabetes 1998; 47: 821–830.
Delatycki MB, Knight M, Koenig M, Cossee M, Williamson R, Forrest SM : G130 V, a common FRDA point mutation, appears to have arisen from a common founder. Hum Genet 1999; 105: 343–346.
Campuzano V, Montermini L, Lutz Y et al: Frataxin is reduced in Friedreich ataxia patients and is associated with mitochondrial membranes. Hum Mol Genet 1997; 6: 1771–1780.
Babcock M, de Silva D, Oaks R et al: Regulation of mitochondrial iron accumulation by Yfh1p, a putative homolog of frataxin. Science 1997; 276: 1709–1712.
Mootha VK, Lindgren CM, Eriksson KF et al: PGC-1alpha-responsive genes involved in oxidative phosphorylation are coordinately downregulated in human diabetes. Nat Genet 2003; 34: 267–273.
Filla A, De Michele G, Cavalcanti F et al: The relationship between trinucleotide (GAA) repeat length and clinical features in Friedreich ataxia. Am J Hum Genet 1996; 59: 554–560.
t’Hart LM, Ruige JB, Dekker JM, Stehouwer CD, Maassen JA, Heine RJ : Altered beta-cell characteristics in impaired glucose tolerant carriers of a GAA trinucleotide repeat polymorphism in the frataxin gene. Diabetes 1999; 48: 924–926.
Dalgaard LT, Hansen T, Urhammer SA, Clausen JO, Eiberg H, Pedersen O : Intermediate expansions of a GAA repeat in the frataxin gene are not associated with type 2 diabetes or altered glucose-induced beta-cell function in Danish Caucasians. Diabetes 1999; 48: 914–917.
Dupont S, Dubois D, Vionnet N et al: No association between the Friedreich's ataxia gene and NIDDM in the French population. Diabetes 1998; 47: 1654–1656.
Lynn S, Hattersley AT, McCarthy MI, Frayling TM, Turnbull DM, Walker M : Intermediate expansions of a X25/frataxin gene GAA repeat and type II diabetes: assessment using parent–offspring trios. Diabetologia 2000; 43: 384–385.
Ristow M, Giannakidou E, Hebinck J et al: An association between NIDDM and a GAA trinucleotide repeat polymorphism in the X25/frataxin (Friedreich's ataxia) gene. Diabetes 1998; 47: 851–854.
Shadrina MI, Miloserdova OV, Slominskii PA, Balabolkin MI, Limborskaiia SA : Association of polymorphic trinucleotide repeat (GAA)n of the Frataxin gene with diabetes mellitus type 2 in the Moscow population. Mol Biol (Mosk) 2002; 36: 37–39.
Groop L, Forsblom C, Lehtovirta M et al: Metabolic consequences of a family history of NIDDM (the Botnia study): evidence for sex-specific parental effects. Diabetes 1996; 45: 1585–1593.
Tripathy D, Carlsson M, Almgren P et al: Insulin secretion and insulin sensitivity in relation to glucose tolerance: lessons from the Botnia Study. Diabetes 2000; 49: 975–980.
Edelstein SL, Knowler WC, Bain RP et al: Predictors of progression from impaired glucose tolerance to NIDDM: an analysis of six prospective studies. Diabetes 1997; 46: 701–710.
Shaw JE, Zimmet PZ, de Courten M et al: Impaired fasting glucose or impaired glucose tolerance. What best predicts future diabetes in Mauritius? Diabetes Care 1999; 22: 399–402.
O’Connell JR, Weeks DE : PedCheck: a program for identification of genotype incompatibilities in linkage analysis. Am J Hum Genet 1998; 63: 259–266.
Purcell S, Cherny SS, Sham PC : Genetic power calculator: design of linkage and association genetic mapping studies of complex traits. Bioinformatics 2003; 19: 149–150.
Gabriel SB, Schaffner SF, Nguyen H et al: The structure of haplotype blocks in the human genome. Science 2002; 296: 2225–2229.
Spielman RS, McGinnis RE, Ewens WJ : Transmission test for linkage disequilibrium: the insulin gene region and insulin-dependent diabetes mellitus (IDDM). Am J Hum Genet 1993; 52: 506–516.
Kruglyak L, Daly MJ, Reeve Daly MP, Lander ES : Parametric and nonparametric linkage analysis: a unified multipoint approach. Am J Hum Genet 1996; 58: 1347–1363.
Clayton D : A generalization of the transmission/disequilibrium test for. Am J Hum Genet 1999; 65: 1170–1177.
Zhao JH, Lissarrague S, Essioux L, Sham PC : GENECOUNTING: haplotype analysis with missing genotypes. Bioinformatics 2002; 18: 1694–1695.
Zhao JH : 2LD, GENECOUNTING and HAP: computer programs for linkage disequilibrium analysis. Bioinformatics 2004; 20: 1325–1326.
Altshuler D, Hirschhorn JN, Klannemark M et al: The common PPARgamma Pro12Ala polymorphism is associated with decreased risk of type 2 diabetes. Nat Genet 2000; 26: 76–80.
Gloyn AL, Weedon MN, Owen KR et al: Large-scale association studies of variants in genes encoding the pancreatic beta-cell KATP channel subunits Kir6.2 (KCNJ11) and SUR1 (ABCC8) confirm that the KCNJ11 E23K variant is associated with type 2 diabetes. Diabetes 2003; 52: 568–572.
Jorde LB : Linkage Disequilibrium and the Search for Complex Disease Genes. Genome Res 2000; 10: 1435–1444.
Melton D, Brown J, Kenty G et al: Endocrine Pancreas Consortium, 2000, http://www.ncbi.nlm.nih.gov/, Accession number: BM022009.
Acknowledgements
We thank the Botnia families for their support and willingness to participate and for making this project possible. This work was supported by grants from the Sigrid Juselius Foundation, the JDF Wallenberg Foundation, the Ingabritt and Arne Lundbergs Foundation, the Finnish Diabetes Research Foundation, The Swedish Medical Research Council, the Novo-Nordisk Foundation, and a European Commission grant to Genome Integrated Force in Type 2 Diabetes (GIFT), the Royal Physiographic Society in Lund, ‘Diabetesföreningen i Malmö med omnejd’, the Medical Faculty of Lund University. The Royal Physiographic Society, Novartis Foundation and The Swedish Medical Research Council support CML.
Author information
Authors and Affiliations
Corresponding author
Additional information
Supplementary Information accompanies the paper on European Journal of Human Genetics website (http://www.nature.com/ejhg)
Supplementary information
Rights and permissions
About this article
Cite this article
Holmkvist, J., Almgren, P., Parikh, H. et al. Haplotype construction of the FRDA gene and evaluation of its role in type II diabetes. Eur J Hum Genet 13, 849–855 (2005). https://doi.org/10.1038/sj.ejhg.5201413
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/sj.ejhg.5201413