Abstract
Sarcoidosis (SA) is a systemic granulomatous disorder of unknown etiology characterized by T helper 1-type inflammatory responses at sites of disease with signs of B cell hyperactivity. Like rheumatoid arthritis and diabetes, an infectious etiology has frequently been postulated but no single infectious trigger definitively identified. Polymorphic alleles at SLC11A1 have previously been associated with susceptibility to both the putative infectious agents and to these autoimmune disorders. We therefore investigated its candidacy as a genetic determinant of SA in Poland in an association-based study comparing 86 SA patients with 85 tuberculosis (TB) patients and 93 control subjects. The functional promoter (GT)n polymorphism and four of 10 other single nucleotide or insertion/deletion polymorphisms genotyped across SLC11A1 were informative in our sample. Consistent with previous autoimmune disease studies, allele 3 at the functional (GT)n promoter region repeat polymorphism was significantly associated with SA when compared with healthy controls (odds ratio 1.68; 95% CI: 1.01–2.81; P=0.04) or with TB patients (odds ratio 1.69; 95% CI: 1.042–0.78; P=0.03).
Similar content being viewed by others
Introduction
Sarcoidosis (SA) is a multisystem granulomatous disorder of unknown etiology. Diagnosis is firmly established when clinicoradiological findings are supported by histological evidence of noncaseating epithelioid cell granulomas found on tissue biopsy. Despite extensive research into the etiology of SA, no identifiable agent has been demonstrated to account for the granulomata. Chemicals, allergy, infectious antigens, autoimmunity and genetic factors have all been explored as potential causes.1 It is likely that genetically predisposed hosts are exposed to antigens that trigger an exaggerated cellular immune response with the production of antibodies, including self-antibodies, leading to granuloma formation.1, 2, 3, 4, 5, 6 A high frequency of endocrine autoimmunity in patients with SA has been reported, but the mechanism of this relationship is not known.7, 8, 9
The inflammatory response in SA is characterized by large numbers of activated macrophages that secrete proinflammatory cytokines/chemokines including tumor necrosis factor α (TNFα) and RANTES (regulation of activation, normal T-cell expression and secretion), and accumulation of CD4+ cells with a Th1-type interferon-γ (IFN-γ) and interleukin-2 (IL-2) cytokine profile.1 These features are shared in common with many inflammatory and autoimmune disorders and, like inflammatory bowel disease, rheumatoid arthritis and diabetes, an infectious etiology has frequently been postulated.1 Although no single infectious trigger has been definitively identified, indirect evidence10, 11, 12 implicates Mycobacterium tuberculosis and other mycobacteria as possible agents involved in triggering disease. Mycobacteria have also been implicated in triggering13, 14, 15, 16 or protecting individuals from15, 16, 17 autoimmune disease. Polymorphic alleles at SLC11A1 have previously been associated with susceptibility to both these putative infectious agents and to these autoimmune disorders (reviewed in Blackwell et al18). SLC11A1 encodes the natural resistance associated macrophage protein 1 that regulates divalent cation homeostasis in macrophages and has many pleiotropic effects on macrophage activation.18 It is also known to be involved in determining Th1 versus Th2 bias in immune response to infection or specific antigens. In humans, no functional variants across the coding region of the gene have been identified, but a functional (GT)n repeat polymorphism has been identified19 and characterized.20, 21, 22 The most common allele in all populations studied, designated allele 3 and characterized by T(GT)5AC(GT)5AC(GT)9 repeats, drives high levels of reporter gene expression and is associated with rheumatoid arthritis23 and juvenile rheumatoid arthritis.24 The second most common allele, designated allele 2 and characterized by T(GT)5AC(GT)5AC(GT)10 repeats, drives low expression and is associated with tuberculosis (TB) in multiple populations.25, 26, 27, 28 By these criteria, we predict that allele 3 would also be associated with susceptibility to SA and allele 2 with TB. To test this hypothesis we carried out a population-based study in Poland comparing SLC11A1 genotypes in SA patients with TB patients and control subjects. This is the first time that polymorphisms at SLC11A1 have been analyzed in patients with SA and TB in the same European Caucasoid ethnic group.
Materials and methods
Patients and controls
SA patients
In total, 86 outpatients (mean age 39.5±10 years; range 21–70 years; 34 women, 52 men) with documented pulmonary SA were studied. Diagnosis of SA was based on histological, clinical and radiological evidence.1 Chest radiographs were staged as stage I (bilateral hilar lymphadenopathy; 29 patients) or stage II (bilateral hilar lymphadenopathy and diffuse pulmonary infiltrationis; 57 patients). A negative PPD skin-test response was an additional criterion for inclusion in the study group. Microbiological and cytological examination of the sputum samples revealed no acid-fast bacilli, fungi or atypical cells.
TB patients
In total, 85 unrelated patients (mean age 48.5±15 years; range 20–80 years; 25 women, 60 men) with newly detected active pulmonary TB at the Pulmonology Hospital in Sopot and Wejherowo, Poland, were studied. A diagnosis of TB was established using standard clinical, radiographic and bacteriological criteria. The patients studied were at a similar clinical stage and with similarly localized disease on the initial chest radiographs (ie infiltrates with cavitation in the one or two lung zones). The diagnosis of TB was confirmed in all patients by demonstration of acid-fast bacilli in sputum smears and by positive sputum culture of the M. tuberculosis strains. A positive PPD skin-test response was an additional criterion included in the studied group. Patients were also classified according to their response to chemotherapy. All TB patients included in this study responded to the first line treatment (rifampin, isoniazid, ethambutol, pyrazinamide) and were therefore classified as drug responders.
Controls
The sample studied consisted of 93 unrelated healthy blood donor volunteers (mean age 38.1±13 years; range 20–80 years; 39 women, 54 men) originating from the region of Gdansk in northern Poland. They had an unremarkable physical examination, normal blood and serum analysis as well as normal CXRs and no acid-fast bacilli in sputum. All of them were negative for the PPD skin test. The controls were of the same socioeconomic status and ethnic background as the patients. The Polish population is a relatively homogenous ethnic group. Patients and controls included people living in the Gdańsk area and in the neighboring districts. As a result of postwar movements, they come from east, north and central Poland; therefore, they may be considered as a homogeneous Caucasian group. None of the TB or SA patients or controls had a familial history of TB, SA or autoimmune disease. All patients and controls were vaccinated with BCG (bacillus Calmette-Guerin). In Poland, BCG vaccinations have been mandatory at birth and again at 7, 12 and 18 years of age since 1955.
Ethical approval for the study was obtained from the Independent Bioethics Committee for Scientific Researches, Medical University of Gdańsk, Poland, and informed consent was obtained for collection of 10 ml blood by venepuncture for preparation of DNA. Samples were anonymised for genetic analysis. A total of 86 SA patients, 85 TB patients and 93 control subjects were available for genotyping of SLC11A1 polymorphisms. All individuals included in this study were HIV negative.
Genotyping
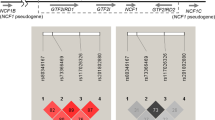
The functional promoter (GT)n repeat polymorphism and 10 single nucleotide (SNPs) (−237C/T; −86G/A; 274C/T; 469+14G/C; 823C/T; 1465–85A/G; D543N G/A; 1801+417C/T29, 30, 31) or insertion/deletion (IN/DEL) (3′UTR TGTG29; 3′UTR CAAA32) polymorphisms were genotyped across SLC11A1. The (GT)n repeat in the promoter region of SLC11A119 was PCR amplified using NED- and FAM-labelled forward primers and PCR products were analyzed by electrophoresis on 6% polacrylamide using a capillary ABI3100 Genetic Analyser (Applied Biosystems, Warrington, UK) as previously described.29 The two IN/DELs29, 32 were also typed using the ABI3100 as described.31 The (GT)n repeat polymorphism and IN/DELs were analyzed using ABI computer software GeneScan™ and Genotyper® v 3.7. For the SNPs, SNaPshot assays using the ABI PRISM® SNaPshot™ Multiplex System. SNP flanking primers were designed using Primer3 software (http://frodo.wi.mit.edu/) with extension primers designed according to the manufacturer's recommendations (Web Table 1). Following DNA amplification, PCR products were pooled and purified by the addition of 2 U exonuclease (New England Biolabs, Hitchin, UK) and 5 U Shrimp Alkaline Phosphatase (SAP; USB, Cleveland, OH, USA) per 15 μl of sample and incubated for 1 h at 37°C followed by 15 min at 72°C in order to remove unincorporated primers and dNTPs. After purification, 2 μl of PCR product was mixed with 0.5 μl of deionized water, 2.5 μl of SNaPshot ready Reaction Mix and 0.5 μl of pooled SNaPshot extension primers (with each primer at 0.4 μ M final concentration) and cycled according to the manufacturer's recommendations. SNaPshot products were then purified by the addition of 1 U SAP and incubated for 1 h at 37°C followed by 15 min at 72°C in order to remove unincorporated [F]ddNTPs. Following purification, 0.5 μl SNaPshot product was mixed with 9 μl Formamide Ultra (Bioproducts Ltd, UK) and 0.5 μl LIZ-120 Size Standard (ABI) then denatured at 95°C for 5 min. Fragments were run on the ABI 3100 Genetic Analyser and analyzed using GeneMapper® Software v 3.0 (Applied Biosystems).
Statistical analysis
Tests for deviation from Hardy–Weinberg equilibrium (HWE), allelic association between markers and disease, and linkage disequilibrium (LD) between markers (reported as the r2 statistic which ranges from 0 to 1 and takes allele frequencies into account), were performed within STATA v 8.0 (http://www.stata.com/) using the GenAssoc package (available from http://www-gene.cimr.cam.ac.uk/clayton/software/stata/). Power calculations were performed in excel using a script prepared in-house at CIMR by Dr Heather Cordell. Genotype and allele frequencies were compared using the χ2 statistic and odds ratios (OR) and 95% confidence intervals (CI) calculated using unconditional logistic regression analysis under a multiplicative model (ie two alleles contribute twice the effect of one allele). A likelihood ratio test was used to determine dominance effects by comparing one degree of freedom (df; allele-wise) and 2 df (genotype-wise) tests.
Results
Allele frequencies for markers in the healthy control group are shown in Table 1. Tests for deviation from HWE confirmed that all markers were in HWE in this control group. The variant alleles at −237C/T, −86G/A, 823C/T, D543N, 3′TGTG and 1801+417C/T were too low in frequency (<0.1) to be used to detect allelic association with disease, and were excluded from further analysis. Variant alleles for all other markers were ≥0.2. Power calculations demonstrated >80% power to detect a significant (P<0.05) effect at an odds ratio ≥2 for a marker with variant allele frequency ≥0.2.
Table 2 shows results of allelic association tests comparing SA patients, TB patients and controls for the five informative markers at SLC11A1. Allele 3 at the functional (GT)n repeat polymorphism was associated with higher risk of SA when compared to both control (OR=1.68; P=0.04) and TB patient (OR=1.69; P=0.03) groups. No dominance effects were observed for either comparison, consistent with a higher OR (3.47; 95% CI 1.04–11.60; P=0.04) for disease in SA patients homozygous for allele 3 at the (GT)n repeat compared to the TB patient group. No allelic associations between TB and SLC11A1 were observed in this population compared to the control group, and no other informative polymorphisms across SLC11A1 were associated with SA compared to either the control or TB patient groups, despite moderate LD between the (GT)n and 274C/T (r2=0.68) and between (GT)n and 469+14G/C (r2=0.58). There was little or no LD (r2=0.24–0.39) between any of the informative 5′ SLC11A1 polymorphic markers ((GT)n; 274C/T; 469+14G/C) and any of the informative 3′ markers (1465–85A/G; 3′ UTR CAAA), although the latter were in strong LD (r2=0.92) with each other.
Discussion
Previous studies of inflammatory and autoimmune diseases (reviewed in Blackwell et al18) and the clinical and immune response phenotypes associated with SA (reviewed1) provided a priori evidence that allele 3 at the (GT)n the functional repeat polymorphism in the promoter region of SLC11A1 would be associated with SA. Data presented here provide evidence to support this prediction, both when SA patients were compared to healthy controls and to TB patients, in this Polish population. This is consistent with previous etiological and epidemiological studies suggesting genetic predisposition to disease1 and, in particular, the hypothesis that mycobacteria might provide one of the specific environmental agents that triggers the inflammatory response in genetically susceptible individuals.12 In this context, we might have expected an ‘equal and opposite’ association between polymorphism at SLC11A1 and susceptibility to TB in this population, especially since the low expressing allele 220 had previously been shown to be associated with both susceptibility to TB25, 26, 27, 33 in multiple populations, and to high anti-inflammatory IL-10 responses33 in the Gambia. In the event, we observed no direct association between polymorphisms at SLC11A1 and TB in Poland, suggesting that the two diseases are under separate genetic control in this population. This does not, however, preclude the possibility that interaction with (environmental) mycobacteria/mycobacterial products could provide the trigger for heightened macrophage activation/proinflammatory responses associated with SLC11A1-regulation of susceptibility to SA in this population. Failure to observe an association between the (GT)n promoter polymorphism at SLC11A1 and pulmonary TB is consistent with previous reports showing population-to-population variation in demonstrable positive25, 27, 28, 33 versus no detectable34 association, the latter even in the presence of association with the 469+14G/C polymorhism in intron 4 of the gene. Hence, other etiological SLC11A1 variants are likely to contribute functionally to disease association with TB. There are a number of other studies that report linkage35, 36 or association37, 38, 39 with polymorphisms across SLC11A1 where the promoter (GT)n polymorphism has not been examined, and other reported failures40, 41 to detect linkage or association between SLC11A1 and TB. This is an expected outcome for diseases where complex inheritance of susceptibility leads to different genes influencing disease in different populations, but some caution in interpreting these data are necessary as samples sizes employed frequently have low power to detect allelic association even when present. Since the sample employed in our study is also small, further work is required both to confirm the positive association with SA and the negative finding for TB in this population.
In previous functional studies, allele 3 at the (GT)n promoter polymorphism was shown to drive high levels of reporter gene activity, even in the absence of exogenous stimuli. By implication, this allele would therefore be associated with high expression of SLC11A1 protein, high activation status of macrophages, and concomitant enhancement of proinflammatory responses including TNF-α, interleukin-1β and chemokine responses. All these are features of immunopathology associated with SA.1 In reporter gene studies the enhanced activity of allele 3 is reversed in the presence of the variant T allele at −237C/T polymorphism.21 This variant was at low frequency (<0.06) in this Polish population. Hence, it was neither possible nor necessary to stratify by the −237C/T polymorphism to detect the allele 3 effect. Nor did we observe any other allelic associations with SA across SLC11A1, suggesting that the (GT)n repeat is the functional variant associated with disease in this population. This is consistent with a previous study of SLC11A1 polymorphisms and SA in an African-American population, where four polymorphisms were studied but only the (GT)n repeat showed significant association with disease.42 In this case the authors pooled the rarer alleles at the (GT)n with allele 2, and reported that this combination of alleles was significantly protective for SA compared with the more common allele 3. Another study of SA in a Japanese population examined only a single polymorphism (1703G/A) at the 3′ end of the gene, and failed to demonstrate an association.43 Similarly, we and others42 found no evidence for association with markers in the 3′ end of the gene in. There was, in any case, no LD between polymorphic markers at 5′ and 3′ ends of SLC11A1, so there was no prior expectation that we should find allelic association with markers in the 3′ region of SLC11A1 in our population.
In summary and consistent with previous autoimmune disease studies, our study demonstrates that allele 3 at the functional (GT)n promoter region repeat polymorphism of SLC11A1 is associated with susceptibility to SA in a European Caucasoid population. This provides further support for the notion of a possible infectious trigger for SA, and adds to the cumulative evidence that exposure to infectious agents has shaped the gene-by-environment interactions that now contributes to susceptibility to autoimmune-related diseases in man.
References
Statement on sarcoidosis: Joint Statement of the American Thoracic Society (ATS), the European Respiratory Society (ERS) and the World Association of Sarcoidosis and Other Granulomatous Disorders (WASOG) adopted by the ATS Board of Directors and by the ERS Executive Committee, February 1999. Am J Respir Crit Care Med 1999; 160: 736–755.
Hunninghake GW, Crystal RG : Mechanisms of hypergammaglobulinemia in pulmonary sarcoidosis. Site of increased antibody production and role of T lymphocytes. J Clin Invest 1981; 67: 86–92.
Weinberg I, Vasiliev L, Gotsman I : Anti-dsDNA antibodies in sarcoidosis. Semin Arthritis Rheum 2000; 29: 328–331.
Negri C, Scovassi AI, Cerino A et al: Autoantibodies to poly(ADP-ribose)polymerase in autoimmune diseases. Autoimmunity 1990; 6: 203–209.
Pilatte Y, Tisserand EM, Greffard A et al: Anticarbohydrate autoantibodies to sialidase-treated erythrocytes and thymocytes in serum from patients with pulmonary sarcoidosis. Am J Med 1990; 88: 486–492.
DeRemee RA : Sarcoidosis and Wegener's granulomatosis: a comparative analysis. Sarcoidosis 1994; 11: 7–18.
Karlish AJ, MacGregor GA : Sarcoidosis, thyroiditis, and Addison's disease. Lancet 1970; 2: 330–333.
Seinfeld ED, Sharma OP : TASS syndrome: unusual association of thyroiditis, Addison's disease, Sjogren's syndrome and sarcoidosis. J Roy Soc Med 1983; 76: 883–885.
Papadopoulos KI, Hallengren B : Polyglandular autoimmune syndrome type III associated with coeliac disease and sarcoidosis. Postgrad Med J 1993; 69: 72–75.
Hanngren A, Odham G, Eklund A et al: Tuberculostearic acid in lymph nodes from patients with sarcoidosis. Sarcoidosis 1987; 4: 101–104.
Almenoff PL, Johnson A, Lesser M, Mattman LH : Growth of acid fast L forms from the blood of patients with sarcoidosis. Thorax 1996; 51: 530–533.
Lyons DJ, Fielding JF : What's in a relationship? Sarcoidosis and tuberculosis. Irish Med J 1990; 83: 76–79.
Rook G, McCulloch J : HLA-DR4, mycobacteria, heat-shock proteins, and rheumatoid arthritis. Arthritis Rheum 1992; 35: 1409–1412.
Rook GA, Adams V, Hunt J et al: Mycobacteria and other environmental organisms as immunomodulators for immunoregulatory disorders. Springer Semin Immunopathol 2004; 25: 237–255.
Classen JB, Classen DC : Immunization in the first month of life may explain decline in incidence of IDDM in The Netherlands. Autoimmunity 1999; 31: 43–45.
Classen DC, Classen JB : The timing of pediatric immunization and the risk of insulin-dependent diabetes mellitus. Infect Dis Clin Pract 1997; 6: 449–454.
Sanjeevi CB, Das AK, Shtauvere-Brameus A : BCG vaccination and GAD65 and IA-2 autoantibodies in autoimmune diabetes in southern India. Ann NY Acad Sci 2002; 958: 293–296.
Blackwell JM, Searle S, Mohamed H, White JK : Divalent cation transport and susceptibility to infectious and autoimmune disease: continuation of the Ity/Lsh/Bcg/Nramp1/Slc11a1 gene story. Immunol Lett 2003; 85: 197–203.
Blackwell JM, Barton CH, White JK et al: Genomic organization and sequence of the human NRAMP gene: identification and mapping of a promoter region polymorphism. Mol Med 1995; 1: 194–205.
Searle S, Blackwell JM : Evidence for a functional repeat polymorphism in the promoter of the human NRAMP1 gene that correlates with autoimmune versus infectious disease susceptibility. J Med Genet 1999; 36: 295–299.
Zaahl MG, Robson KJ, Warnich L, Kotze MJ : Expression of the SLC11A1 (NRAMP1) 5′-(GT)n repeat: opposite effect in the presence of −237C → T. Blood Cells Mol Dis 2004; 33: 45–50.
Donninger H, Cashmore TJ, Scriba T et al: Functional analysis of novel SLC11A1 (NRAMP1) promoter variants in susceptibility to HIV-1. J Med Genet 2004; 41: e49.
Shaw M-A, Clayton D, Atkinson SE et al: Linkage of rheumatoid arthritis to the candidate gene NRAMP1 on 2q35. J Med Genet 1996; 33: 672–677.
Sanjeevi CB, Miller EN, Dabadghao P et al: Polymorphism at NRAMP1 and D2S1471 loci associated with juvenile rheumatoid arthritis. Arthritis Rheum 2000; 43: 1397–1404.
Bellamy R, Ruwende C, Corrah T et al: Variation in the NRAMP1 gene is associated with susceptibility to tuberculosis in West Africans. N Engl J Med 1998; 338: 640–644.
Hoal van Helden EG, Lewis L-A, Tanzer F et al: SLC11A1 (NRAMP1) but not SLC11A2 (NRAMP2) polymorphisms are associated with susceptibility to tuberculosis in a high incidence community in South Africa. Int J Tuberc Lung Dis 2004; 8: 1464–1471.
Gao PS, Fujishima S, Mao XQ et al: Genetic variants of NRAMP1 and active tuberculosis in Japanese populations. International Tuberculosis Genetics Team. Clin Genet 2000; 58: 74–76.
Ma X, Dou S, Wright JA et al: 5′ dinucleotide repeat polymorphism of NRAMP1 and susceptibility to tuberculosis among Caucasian patients in Houston, Texas. Int J Tuberc Lung Dis 2002; 6: 818–823.
Liu J, Fujiwara TM, Buu NT et al: Identification of polymorphisms and sequence variants in human homologue of the mouse natural resistance-associated macrophage protein gene. Am J Hum Genet 1995; 56: 845–853.
Lewis L-A, Victor T, Hoal-van Helden E et al: Identification of a C to T mutation at position −236 bp in the human NRAMP1 promoter. Immunogenetics 1996; 44: 309–311.
Mohamed HS, Ibrahim ME, Miller EN et al: SLC11A1 (formerly NRAMP1) and susceptibility to visceral leishmaniasis in The Sudan. Eur J Hum Genet 2004; 12: 66–74.
Buu NT, Cellier M, Gros P, Schurr E : Identification of a highly polymorphic length variant in the 3′UTR of NRAMP1. Immunogenetics 1995; 42: 428–429.
Awomoyi AA, Marchant A, Howson JM et al: Interleukin-10, polymorphism in SLC11A1 (formerly NRAMP1), and susceptibility to tuberculosis. J Infect Dis 2002; 186: 1804–1814.
Cervino AC, Lakiss S, Sow O, Hill AV : Allelic association between the NRAMP1 gene and susceptibility to tuberculosis in Guinea-Conakry. Ann Hum Genet 2000; 64: 507–512.
Greenwood CM, Fujiwara TM, Boothroyd LJ et al: Linkage of tuberculosis to chromosome 2q35 loci, including NRAMP1, in a large aboriginal Canadian family. Am J Hum Genet 2000; 67: 405–416.
Shaw M-A, Collins A, Peacock CS et al: Evidence that genetic susceptibility to Mycobacterium tuberculosis in a Brazilian population is under oligogenic control: linkage study of the candidate genes NRAMP1 and TNFA. Tuberc Lung Dis 1997; 78: 35–45.
Ryu S, Park YK, Bai GH et al: 3′UTR polymorphisms in the NRAMP1 gene are associated with susceptibility to tuberculosis in Koreans. Int J Tuberc Lung Dis 2000; 4: 577–580.
Delgado JC, Baena A, Thim S, Goldfeld AE : Ethnic-specific genetic associations with pulmonary tuberculosis. J Infect Dis 2002; 186: 1463–1468.
Liu W, Cao WC, Zhang CY et al: VDR and NRAMP1 gene polymorphisms in susceptibility to pulmonary tuberculosis among the Chinese Han population: a case–control study. Int J Tuberc Lung Dis 2004; 8: 428–434.
Liaw YS, Tsai-Wu JJ, Wu CH et al: Variations in the NRAMP1 gene and susceptibility of tuberculosis in Taiwanese. Int J Tuberc Lung Dis 2002; 6: 454–460.
El Baghdadi J, Remus N, Benslimane A et al: Variants of the human NRAMP1 gene and susceptibility to tuberculosis in Morocco. Int J Tuberc Lung Dis 2003; 7: 599–602.
Maliarik MJ, Chen KM, Sheffer RG et al: The natural resistance-associated macrophage protein gene in African Americans with sarcoidosis. Am J Respir Cell Mol Biol 2000; 22: 672–675.
Akahoshi M, Ishihara M, Remus N et al: Association between IFNA genotype and the risk of sarcoidosis. Hum Genet 2004; 114: 503–509.
Acknowledgements
Our study was funded by Polish State Committee for Scientific Researches – Grant No. 3PO5B 15522. Genotyping and genetic analysis in Cambridge was supported by grants to JB from The Wellcome Trust.
Author information
Authors and Affiliations
Corresponding author
Additional information
Supplementary Information accompanies the paper on European Journal of Human Genetics website (http://www.nature.com/ejhg)
Supplementary information
Rights and permissions
About this article
Cite this article
Dubaniewicz, A., Jamieson, S., Dubaniewicz-Wybieralska, M. et al. Association between SLC11A1 (formerly NRAMP1) and the risk of sarcoidosis in Poland. Eur J Hum Genet 13, 829–834 (2005). https://doi.org/10.1038/sj.ejhg.5201370
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/sj.ejhg.5201370
Keywords
This article is cited by
-
Diminished Susceptibility of African–Americans to Non-tuberculous Mycobacterial Disease
Lung (2018)
-
Genomics of Human Pulmonary Tuberculosis: from Genes to Pathways
Current Genetic Medicine Reports (2017)
-
The functional SLC11A1 gene polymorphisms are associated with sarcoidosis in Turkish population
Molecular Biology Reports (2012)
-
Tuberculosis as a complex trait: impact of genetic epidemiological study design
Mammalian Genome (2011)
-
Genetic polymorphisms and susceptibility to lung disease
Journal of Negative Results in BioMedicine (2006)