Abstract
Chinese hamster ovary cultured cells were transformed to continuously express wild-type and two mutant ornithine transcarbamylase genes, R141Q and R40H. In addition, these cells were transfected to transiently express the same genes. The R141Q mutation abolishes the enzymatic activity, and the amount of “mature” protein present in transfected cells is equivalent to the wild type. The R40H mutation causes a reduction of enzymatic activity to approximately 26 to 35% of wild type concomitant with a significant reduction in the amount of protein present. Transfection with wild-type and mutant genes together in various proportions did not reveal dominant negative effects of the two mutations studied. This expression system can be used to examine the deleterious effect of private mutations or lack thereof in families with ornithine transcarbamylase deficiency as well as evaluate the potential dominant negative effects of gene delivery for treatment of ornithine transcarbamylase deficiency.
Similar content being viewed by others
Main
The enzyme OTC (EC 2.1.3.3) catalyzes the conversion of ornithine and carbamyl phosphate to citrulline and inorganic phosphate. Human OTC is expressed from a gene on the X chromosome as an inactive precursor polypeptide containing a 32 amino acid N-terminal signal sequence that is cleaved upon transport into the mitochondria, where the active homotrimeric enzyme is formed(1). Mutations in the OTC gene result in varying degrees of OTCD, causing hyperammonemia due to decreased synthesis of urea in the liver.
More than 130 apparently deleterious “private” mutations of the human OTC gene have been observed in families affected by OTCD(2). Although some of the mutations are deletions or insertions that are readily identified as causing OTCD, over 80% are single-base substitutions, and only a few of these have been examined in expression studies to confirm and investigate altered biochemical and/or molecular mechanisms.
Following the initial isolation of human OTC cDNA and study of its expression in HeLa cells(3), it was demonstrated that a C to T transition in codon 141 (R141Q) observed in OTCD newborns with acute neonatal hyperammonemic coma resulted in a complete loss of enzyme activity when the mutant cDNA was expressed in Cos1 cells(4). Because arginine 141 is one of the active site residues that binds to one of the oxygen atoms of carbamyl phosphate(5), its replacement will directly affect substrate binding. On the other hand, the R40H mutation is associated with an extremely variable phenotype and the mechanism of its deleterious effect is unknown. When expressed in Cos1 cells, only 10% of control activity was observed(6). Subsequently, it was shown that mRNA transcribed from R40H cDNA was equal in both abundance and stability to that transcribed from wild-type cDNA, however, it was found that the R40H protein was less abundant than wild-type OTC(7). In our laboratory, an in vitro study of mature human OTC protein showed that the biochemical and physical properties of the R40H protein were indistinguishable from wild-type OTC(8).
A large proportion of mutant OTC appear to have a short half-life, as evidenced by low or absent cross-reactive material in deficient livers(9). If a wild-type OTC gene is expressed within the same cell, it is unknown whether the mutant polypeptide that is being simultaneously synthesized by the cell will interfere with the assembly of a wild-type trimer and thus exert a dominant negative effect. One previous study suggested that such a phenomenon may occur in the OTC protein(10).
In this work we used CHO cells to develop both stable and transient transfection systems, enabling the study of naturally occurring mutations in the human OTC gene, and we demonstrate the use of these systems by expressing wild-type, R141Q mutant, and R40H mutant OTC genes. We then conducted experiments to investigate the occurrence of dominant negative effects upon formation of the heterotrimeric enzyme using these two mutations. Our studies show that CHO cells can be used for stable transformation with the human OTC gene, that they are suitable for expression studies of mutations, and that no dominant negative effect is apparent for the two mutations studied.
METHODS
Cell lines.
CHO cells were either purchased from the American Type Culture Collection (Manassas, VA, U.S.A.) (CHO-K1, passage: approximately 400), or were a gift from the laboratory of Dr. P. Law (Department of Pharmacology, University of Minnesota, Minneapolis, MN, U.S.A.) (CHO passage unknown). Lines established in this study by hygromycin selection of stable transformants include CHO (pHOE1), expressing full-length wild-type human OTC; CHO (pHOE1R40H), expressing the R40H human OTC mutant; and CHO (pHOE1R141Q), expressing the R141Q human OTC mutant. Cells were grown in Ham's F12 medium supplemented with 10% fetal bovine serum (Atlanta Biologicals, Norcross, GA, U.S.A.), 100 U/mL penicillin G sodium, and 100 μg/mL streptomycin sulfate (Life Technologies, Rockville, MD, U.S.A.) in a humidified 5% CO2 atmosphere at 37°C. Stable transformant cultures were maintained in 250 μg/mL hygromycin B (Life Technologies).
OTC expression constructs.
Plasmid pHOE1 was constructed by inserting the 1.2 Kb Hin FI human OTC cDNA fragment from plasmid pHO731(3) into pIREShyg (CLONTECH, Palo Alto, CA) cleaved with Bam HI. Both fragments were blunt-ended by Klenow enzyme fill-in before ligation. Sequencing of pHOE1 revealed the loss of a single base pair from the upstream end of the Hin FI fragment during construction but no mutations within the reading frame. The desired single base pair changes were introduced into the OTC gene in pHOE1 by megaprimer PCR site-directed mutagenesis(11). Mutations were introduced using the mutagenic primers for R40H (GAGAAGGTCAtGGCCCTTCAGCTGCAC) and R141Q (TTATACACTtGAGCCAATA), where lowercase letters represent mutagenic nucleotides. In accordance with the nomenclature of Seraphin and Kandels-Lewis(11), we used the following enzymes and primers for site-directed mutagenesis of pHOE1. To construct pHOE1R40H we used enzyme A (Bam HI), enzyme B (Aoc I), and the mutated primer (R40H); to construct pHOE1R141Q we used enzyme A (Aoc I), enzyme B (Bst EII), and the mutated primer (R141Q). The opposing primers were pIRES-F (CCCACTGCTTACTGGCTTATCG) and pIRES-R (CTGCTTCCTTCACGACATTCAAC) for R40H and R141Q, respectively. All constructs were confirmed by DNA sequencing.
Transfection.
Plasmids were introduced into cells by lipofection. Cells were plated 24 h before transfection to reach 50 to 80% confluence in 35-mm dishes. Plasmid DNA (4 μg), prepared using an endotoxin-free plasmid purification kit (QIAGEN Inc., Valencia, CA, U.S.A.), was mixed with 20 μL GenePorter reagent (Gene Therapy Systems, San Diego, CA, U.S.A.) and incubated for 30 min at room temperature. A luciferase expression plasmid, pGL3-control (Promega, Madison, WI), was included in transient transfections to determine relative transformation efficiency. The DNA/lipid mixture was combined with 1 mL of serum-free medium and applied to cells. Following 5 h incubation, 1 mL medium containing 20% fetal bovine serum was added. To establish stable transformation of cell lines, the cells were harvested 48 h posttransfection, serially diluted, and incubated in complete medium containing 600 μg/mL hygromycin B until individual colonies could be lifted on cloning discs and expanded.
OTC assays.
Cells were harvested by treatment with 0.05% trypsin and 0.53 mM EDTA (Life Technologies) for 3–5 min, washed with phosphate-buffered normal saline, suspended in mitochondrial lysis buffer [MLB: 0.5% Triton X100, 10 mM HEPES, 0.5 mM DTT, 2 mM EDTA, 1% protease inhibitor cocktail (P8340; Sigma Chemical Co., St. Louis, MO, U.S.A.), pH 7.4], and incubated for 30 min on ice. Cell lysis was accomplished by two rounds of rapid freezing and thawing in liquid nitrogen and a 37°C water bath. Cell lysates were chilled on wet ice, centrifuged at 13,000 g for 10 min at 4°C, and the supernatant fluid was transferred to clean tubes on ice.
OTC activity was measured as described previously(12). Luciferase activity was measured by luminometry using Luciferase Assay Reagent (Promega). Protein concentration was determined using a bicinchoninic acid assay kit (Pierce, Rockford, IL, U.S.A.). In transient transfection experiments, OTC activities were normalized for variance due to transfection. Normalization factors were determined by dividing each sample's luciferase-specific activity by the mean luciferase-specific activity of all samples. The OTC specific activity for each sample was then multiplied by its normalization factor.
Western blot analyses were performed using anti-OTC serum from a rabbit immunized with pure recombinant human OTC. Proteins were precipitated from cell lysate supernatants in 80% acetone and resuspended at 1.75 mg/mL. Each well was loaded with 35 μg protein and separation was achieved by SDS PAGE through a 12% slab gel. After electrophoretic transfer to a nitrocellulose membrane, the blot was incubated with a 1:500 dilution of OTC antiserum and then with alkaline phosphatase-conjugated goat anti-rabbit IgG. Color was developed using an Immun-blot Assay kit (BioRad, Richmond, CA, U.S.A.).
RESULTS
Establishment of CHO cell lines expressing OTC cDNA.
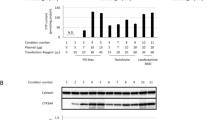
Following transfection with OTC expression constructs pHOE1 (WT), pHOE1R40H, or pHOE1R141Q, colonies of cells selected by hygromycin were studied and found to express OTC continuously. The specific activities of OTC are presented in Table 1. Untransfected CHO cells (controls) and those transformed to hygromycin resistance by pHOE1R141Q had essentially no OTC activity, whereas cells transformed by the wild-type or R40H-mutated OTC constructs each catalyzed conversion of ornithine and carbamyl phosphate to citrulline. Western analysis of the OTC proteins in these cells revealed that the threefold greater OTC activity expressed from wild-type versus R40H constructs is comparable to the relative amount of OTC cross-reactive material present (Fig. 1A, lanes 3 and 4). However, the noncatalytic R141Q enzyme from the cell line transformed by the respective vector was present in an amount approximately equal to that expressed from the wild-type construct (Fig. 1A, lanes 3 and 5). These results indicate that wild-type and R40H OTC mature enzymes have similar levels of enzymatic specific activity, and that R141Q OTC is catalytically inactive.
Western blot analysis of human OTC gene expression in CHO cell lines selected for stable transformation (A) or exposed to transient transfections (B). Each lane except for lane 1 (OTC control) was loaded with 35 μg of protein from cell lysates followed by electrophoresis separation and detection with rabbit anti-human OTC serum and alkaline phosphatase-conjugated goat anti-rabbit IgG. In both (A) and (B) lane 1 contains 1 μg purified recombinant human OTC. (A) Lane 2: nontransfected cells;lanes 3–5: cells selected for stable transformation following transfection with pHOE1 (wild type), pHOE1R40H, and pHOE1R141Q, respectively. (B) Cells transiently transfected with various vectors with the specified amounts of plasmid. Lane 2: 3 μg pIREShyg (parent vector);lanes 3–5: 3 μg of pHOE1 (wild type), pHOE1R40H, and pHOE1R141Q, respectively; lanes 6–8: 1.5 μg pHOE1 + 1.5 μg pIREShyg, 1.5 μg pHOE1 + 1.5 μg pHOE1R40H, and 1.5 μg pHOE1 + 1.5μg pHOE1R141Q, respectively. In addition to the vectors indicated above, each cell line was also co-transfected with 1 μg pGL3 for luciferase marker of transfection efficiency.
Effect of mutated OTC on wild-type OTC activity.
CHO-K1 cells were transiently transfected with mixtures of wild-type and mutated OTC expression vectors to search for any dominant negative effect of the R40H or R141Q mutations. In each transfection, total plasmid DNA was kept constant and the amount of wild-type OTC plasmid relative to mutated vector, R40H, or R141Q plasmids was varied. As seen in Table 2 and Figure 2, the amount of OTC activity expressed from mixtures of wild-type and R141Q constructs is similar to that expressed from mixtures of wild-type construct and parent vector (pIREShyg versus R141Q). However, mixtures of wild-type and R40H constructs expressed significantly more OTC activity than did mixtures of wild-type construct and parent vector (pIREShyg versus R40H). Therefore, R141Q OTC seems to have no negative effect on the activity expressed from wild-type OTC cDNA, and R40H OTC has an additive effect. To confirm that these results can be interpreted on the basis of the amount of OTC protein expressed from the various constructs, OTC in cellular lysates from these transfections was determined in Western blots. Cells transfected with 100% pHOE1, or pHOE1R141Q, or a 50/50 mixture of these constructs contains approximately equal amounts of OTC protein (Fig. 1B, lanes 3, 5, and 8). Cells transfected with 100% pHOE1R40H contain significantly less OTC protein (Fig. 1B, lane 4), and cells transfected with a 50/50 mixture of pHOE1/pHOE1R40H contain more OTC protein than cells transfected with a 50/50 mixture of pHOE1/pIREShyg (Fig. 1B, lane 7 versus lane 6).
Relative mean OTC-specific activities in CHO-K1 cells transiently transfected with mixtures of wild-type and parent vector or mutated OTC expression constructs. Enzyme activity in each experiment was averaged from four 35-mm plates and normalized to luciferase expressed from co-transfected pGL3. Across the 52 plates of transfected cells, the normalization factor ranged from 0.436 to 1.54. Error bars were eliminated for clarity and may be discerned from standard deviations presented in Table 2. The superimposition of R141Q and pIREShyg curves and the parallel R40H curve suggests no dominant negative effect of either mutant protein on the activity of wild-type OTC.
DISCUSSION
We have previously used human OTC expression in Escherichia coli to study the biochemical and physical properties of several naturally occurring OTC mutants(8). Some of the mutations occurring in the OTC gene do not affect the catalytic activity or biochemical properties when the “mature” mutant protein is purified and examined in vitro. To understand the effects of these mutations, it is essential to express the respective mutations in mammalian cells. We have now successfully used a CHO cell line for expression of wild-type and mutant human OTC genes in both stable and transient expression systems. Other eukaryotic cells (Cos1, primary hepatocytes) have been used before for transient transfection of OTC genes. The advantage of the stable expression system for OTC genes developed by us in CHO cells is that it allows studies that require ongoing synthesis and processing of the enzyme and alleviates the need for and the variability of repeated transfections. This and other expression systems for the OTC gene have the shortcoming of unregulated overexpressions of the transgene. However, even in an overexpressed system one can look for differences between wild-type and mutant proteins. The assumption we made was that the dominant negative effect would be determined by the ratio between the number of the wild-type and mutant polypeptides rather then by their absolute abundance. The effects of the mutations causing either neonatal disease (R141Q) or milder late-onset disease (R40H) were consistent in both stable and transient transfection experiments. The R141Q mutation almost completely abolishes enzyme activity as it alters an active site residue (arginine 141) responsible for binding one of the oxygen atoms of carbamyl phosphate(5, 13). However, as seen in our results, the amount of R141Q mutant protein present in both the stable and transiently transfected cells is equivalent to that seen in cells expressing the wild-type enzyme. On the other hand, the R40H mutant protein showed residual OTC activity of approximately 26 to 35% of wild type, consistent with a much milder phenotype and occasionally lack of symptoms(6). In addition, the amount of R40H mutant protein is markedly reduced to about the same degree in both stable and transient expression systems. Because the “mature” R40H mutant shows catalytic activity and biochemical and physical properties virtually identical to the wild type(8), it is likely that the mutant protein has either a shortened half-life within the cytosol or the mitochondrial matrix or has a reduced mitochondrial import rate. The leader sequence of OTC is composed of 32 neutral or basic amino acids that are cleaved upon entry of the protein into the mitochondria(1). It is possible that arginine 40 may play a role in protection of the preprotein or mature protein from protease degradation or is important for normal mitochondrial cleavage. Specific studies of R40H mutant OTC transport will need to be performed to examine these possibilities.
Gene therapy for OTC deficiency aims to deliver a normal OTC gene into hepatocytes for expression of wild-type OTC(14). Previous work suggested the occurrence of a dominant negative effect when both wild-type and mutant OTC genes are present within the cells(10). The R141Q and R40H mutations were examined in this study for the dominant negative effect they may have on wild-type OTC. We found that neither of these mutations has a negative effect on wild-type OTC activity expressed within the same cell, and that the OTC activity expressed from R40H-mutated OTC DNA is additive to that expressed from wild-type OTC. The reason for the discrepancy from a previously published study(10) is unclear to us and may be related to the different mutation examined in the previous study (R92Q), although that mutation is similar to one of the mutations examined by us (R141Q). The R141Q mutation affects the carbamyl phosphate binding site and does not affect the tertiary structure of the protein(5). Thus, it is expected to be readily incorporated into the holoenzyme without interference in the assembly of the heterotrimeric protein and, hence, lack of dominant negative effect. In such case, one would expect one or two of the three catalytic sites in the heterotrimers to remain functional, although this is difficult to study directly. The R40H has normal structure and catalytic activity(8) and would therefore not be expected to cause a dominant negative effect as documented in this work. Because CHO cells have a relatively short doubling time and are readily transfected, the method of examining mixed mutated and wild-type OTC transient expression used in this study allows rapid screening of OTC mutations for dominant negative effects. This is important given the observation that dominant negative effects of mutated OTC genes may interfere with efforts to alleviate OTCD by gene therapy.
There are now several mutations in the OTC gene recognized to cause divergent clinical phenotypes ranging from late-onset disease to lack of symptoms, even within the same family. By studying the deleterious effects of such mutations in mammalian cells and identifying the mechanisms involved, one can subsequently examine potential modifying factors that account for the various degrees of severity within the same genotype.
Abbreviations
- OTC:
-
ornithine transcarbamylase
- OTCD:
-
ornithine transcarbamylase deficiency
- CHO:
-
Chinese hamster ovary
References
Horwich AL, Kalousek F, Fenton WA, Pollock RA, Rosenberg LE 1986 Targeting of pre-ornithine transcarbamylase to mitochondria: definition of critical regions and residues in the leader peptide. Cell 44: 451–459.
Tuchman M, Morizono H, Rajagopal BS, Plante RJ, Allewell NM 1998 The biochemical and molecular spectrum of ornithine transcarbamylase deficiency. J Inher Metab Dis 21: 40–58.
Horwich AL, Fenton WA, Williams KR, Kalousek F, Kraus JP, Doolittle RF, Konigsberg W, Rosenberg LE 1984 Structure and expression of a complementary DNA for the nuclear coded precursor of human mitochondrial ornithine transcarbamylase. Science 224: 1068–1074.
Lee JT, Nussbaum RL 1989 An arginine to glutamine mutation in residue 109 of human ornithine transcarbamylase completely abolishes enzymatic activity in Cos1 cells. J Clin Invest 84: 1762–1766.
Shi D, Morizono H, Ha Y, Aoyagi M, Tuchman M, Allewell NM 1998 1. J Biol Chem 273: 34247–34254.
Matsuda I, Matsuura T, Nishiyori A, Komaki S, Hoshide R, Matsumoto T, Funakoshi M, Kiwaki K, Endo F, Hata A, Shimadzu M, Yoshino M 1996 Phenotypic variability in male patients carrying the mutant ornithine transcarbamylase (OTC) allele, Arg40His, ranging from a child with an unfavourable prognosis to an asymptomatic older adult. J Med Genet 33: 645–648.
Nishiyori A, Yoshino M, Kato H, Matsuura T, Hoshide R, Matsuda I, Kuno T, Miyazaki S, Hirose S, Kuromaru R, Mori M 1997 The R40H mutation in a late onset type of human ornithine transcarbamylase deficiency in male patients. Hum Genet 99: 171–176.
Morizono H, Listrom CD, Rajagopal BS, Aoyagi M, McCann MT, Allewell NM, Tuchman M 1997 Late onset ornithine transcarbamylase deficiency: function of three purified recombinant mutant enzymes. Hum Mol Genet 6: 963–968.
Zhang W, Holzknecht RA, Tuchman M 1990 Immunochemical analysis of carbamyl phosphate synthetase I and ornithine transcarbamylase deficient livers: elevated N-acetylglutamate level in CRM negative carbamyl phosphate synthetase I deficiency. Clin Invest Med 13: 183–188.
Morsy MA, Zhao JZ, Ngo TT, Warman AW, O'Brien WE, Graham FL, Caskey CT 1996 Patient selection may affect gene therapy success. J Clin Invest 97: 826–832.
Seraphin B, Kandels-Lewis S 1996 An efficient PCR mutagenesis strategy without gel purification step that is amenable to automation. Nucleic Acids Res 24: 3276–3277.
Ye X, Robinson MB, Batshaw ML, Furth EE, Smith I, Wilson JM 1996 Prolonged metabolic correction in adult ornithine transcarbamylase-deficient mice with adenoviral vectors. J Biol Chem 271: 3639–3646.
Allewell NM, Shi D, Morizono H, Tuchman M 1999 Molecular recognition by ornithine and aspartate transcarbamylases. Acc Chem Res 32: 885–894.
Raper SE, Wilson JM, Judkoff M, Robinson MB, Ye X, Batshaw ML 1998 Developing adenoviral-mediated in vivo gene therapy for ornithine transcrabamylase deficiency. J Inher Metab Dis 21: 119
Author information
Authors and Affiliations
Additional information
This work was supported by public health service grant DK47870 from the National Institute of Diabetes Digestive and Kidney Diseases.
Rights and permissions
About this article
Cite this article
Augustin, L., Mavinakere, M., Morizono, H. et al. Expression of Wild-Type and Mutant Human Ornithine Transcarbamylase Genes in Chinese Hamster Ovary Cells and Lack of Dominant Negative Effect of R141Q and R40H Mutants. Pediatr Res 48, 842–846 (2000). https://doi.org/10.1203/00006450-200012000-00023
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1203/00006450-200012000-00023
This article is cited by
-
A dual AAV system enables the Cas9-mediated correction of a metabolic liver disease in newborn mice
Nature Biotechnology (2016)