Abstract
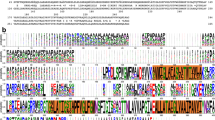
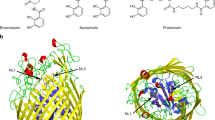
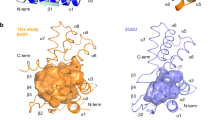
OprD proteins form a large family of substrate-specific outer-membrane channels in Gram-negative bacteria. We report here the X-ray crystal structure of OprD from Pseudomonas aeruginosa, which reveals a monomeric 18-stranded β-barrel characterized by a very narrow pore constriction, with a positively charged basic ladder on one side and an electronegative pocket on the other side. The location of highly conserved residues in OprD suggests that the structure represents the general architecture of OprD channels.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$189.00 per year
only $15.75 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout


Similar content being viewed by others
Accession codes
References
Hancock, R.E. & Brinkman, F.S. Annu. Rev. Microbiol. 56, 17–38 (2002).
Tamber, S., Ochs, M.M. & Hancock, R.E. J. Bacteriol. 188, 45–54 (2006).
Yoshimura, F. & Nikaido, H. J. Bacteriol. 152, 636–642 (1982).
Trias, J. & Nikaido, H. J. Biol. Chem. 265, 15680–15684 (1990).
Wolter, D.J., Hanson, N.D. & Lister, P.D. FEMS Microbiol. Lett. 236, 137–143 (2004).
Bagos, P.G., Liakopoulos, T.D., Spyropoulos, I.C. & Hamodrakas, S.J. Nucleic Acids Res. 32 (Web Server issue), W400–4 (2004).
Huang, H., Jeanteur, D., Pattus, F. & Hancock, R.E. Mol. Microbiol. 16, 931–941 (1995).
Yoshihara, E., Gotoh, N., Nishino, T. & Nakae, T. FEBS Lett. 394, 179–182 (1996).
Yoshihara, E., Yoneyama, H., Ono, T. & Nakae, T. Biochem. Biophys. Res. Commun. 247, 142–145 (1998).
Moraes, T.F., Bains, M., Hancock, R.E. & Strynadka, N.C. Nat. Struct. Mol. Biol. 14, 85–87 (2007).
Huang, H. & Hancock, R.E. J. Bacteriol. 178, 3085–3090 (1996).
Acknowledgements
This work was supported by a Pew Scholar award and University of Massachusetts Medical School start-up funds (B.v.d.B.), and by Syracuse University start-up funds and US National Science Foundation grant DMR-0706517 (L.M.). We thank E. Hearn for critical reading of the manuscript, K. Akpalu for purification of OprD mutants, and A.J. Wolfe, C.P. Goodrich and K.R. Howard for their help with single-channel experiments. We also thank the staff members of beamlines X6A and X25 at the National Synchrotron Light Source for beamtime and assistance.
Author information
Authors and Affiliations
Contributions
S.B. cloned, purified and crystallized OprD; M.M.M. performed single-channel electrical recordings; D.R.P. worked on the early stages of the project; L.M. supervised and designed single-channel electrical recordings; B.v.d.B. determined the OprD structure, designed research and wrote the manuscript.
Corresponding author
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–3, Supplementary Table 1, Supplementary Methods (PDF 381 kb)
Rights and permissions
About this article
Cite this article
Biswas, S., Mohammad, M., Patel, D. et al. Structural insight into OprD substrate specificity. Nat Struct Mol Biol 14, 1108–1109 (2007). https://doi.org/10.1038/nsmb1304
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nsmb1304
This article is cited by
-
Probing the binding affinities of imipenem and ertapenem for outer membrane carboxylate channel D1 (OccD1) from P. aeruginosa: simulation studies
Journal of Molecular Modeling (2017)
-
ESKAPEing the labyrinth of antibacterial discovery
Nature Reviews Drug Discovery (2015)
-
The porin and the permeating antibiotic: a selective diffusion barrier in Gram-negative bacteria
Nature Reviews Microbiology (2008)