Abstract
ATP-dependent allosteric regulation of the ring-shaped group II chaperonins remains ill defined, in part because their complex oligomeric topology has limited the success of structural techniques in suggesting allosteric determinants. Further, their high sequence conservation has hindered the prediction of allosteric networks using mathematical covariation approaches. Here, we develop an information theoretic strategy that is robust to residue conservation and apply it to group II chaperonins. We identify a contiguous network of covarying residues that connects all nucleotide-binding pockets within each chaperonin ring. An interfacial residue between the networks of neighboring subunits controls positive cooperativity by communicating nucleotide occupancy within each ring. Strikingly, chaperonin allostery is tunable through single mutations at this position. Naturally occurring variants at this position that double the extent of positive cooperativity are less prevalent in nature. We propose that being less cooperative than attainable allows chaperonins to support robust folding over a wider range of metabolic conditions.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$189.00 per year
only $15.75 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
Accession codes
References
Motlagh, H.N., Wrabl, J.O., Li, J. & Hilser, V.J. The ensemble nature of allostery. Nature 508, 331–339 (2014).
Hilser, V.J., Wrabl, J.O. & Motlagh, H.N. Structural and energetic basis of allostery. Annu. Rev. Biophys. 41, 585–609 (2012).
Tsai, C.-J. & Nussinov, R. A unified view of “how allostery works”. PLoS Comput. Biol. 10, e1003394 (2014).
Swain, J.F. & Gierasch, L.M. The changing landscape of protein allostery. Curr. Opin. Struct. Biol. 16, 102–108 (2006).
Monod, J., Wyman, J. & Changeux, J.P.O.N. On the nature of allosteric transitions: a plausible model. J. Mol. Biol. 12, 88–118 (1965).
Zheng, W., Brooks, B.R. & Thirumalai, D. Low-frequency normal modes that describe allosteric transitions in biological nanomachines are robust to sequence variations. Proc. Natl. Acad. Sci. USA 103, 7664–7669 (2006).
Zheng, W., Brooks, B.R., Doniach, S. & Thirumalai, D. Network of dynamically important residues in the open/closed transition in polymerases is strongly conserved. Structure 13, 565–577 (2005).
Lee, Y., Choi, S. & Hyeon, C. Mapping the intramolecular signal transduction of G-protein coupled receptors. Proteins 82, 727–743 (2014).
Lockless, S.W. & Ranganathan, R. Evolutionarily conserved pathways of energetic connectivity in protein families. Science 286, 295–299 (1999).
Smock, R.G. et al. An interdomain sector mediating allostery in Hsp70 molecular chaperones. Mol. Syst. Biol. 6, 414 (2010).
Felsenstein, J. Phylogenies and the comparative method. Am. Nat. 125, 1–15 (1985).
Morcos, F. et al. Direct-coupling analysis of residue coevolution captures native contacts across many protein families. Proc. Natl. Acad. Sci. USA 108, E1293–E1301 (2011).
Malinverni, D., Marsili, S., Barducci, A. & De Los Rios, P. Large-scale conformational transitions and dimerization are encoded in the amino-acid sequences of Hsp70 chaperones. PLoS Comput. Biol. 11, e1004262 (2015).
Skerker, J.M. et al. Rewiring the specificity of two-component signal transduction systems. Cell 133, 1043–1054 (2008).
Aakre, C.D. et al. Evolving new protein-protein interaction specificity through promiscuous intermediates. Cell 163, 594–606 (2015).
Lopez, T., Dalton, K. & Frydman, J. The mechanism and function of group II chaperonins. J. Mol. Biol. 427, 2919–2930 (2015).
Hayer-Hartl, M., Bracher, A. & Hartl, F.U. The GroEL-GroES chaperonin machine: a nano-cage for protein folding. Trends Biochem. Sci. 41, 62–76 (2016).
Yam, A.Y. et al. Defining the TRiC/CCT interactome links chaperonin function to stabilization of newly made proteins with complex topologies. Nat. Struct. Mol. Biol. 15, 1255–1262 (2008).
Bigotti, M.G. & Clarke, A.R. Chaperonins: the hunt for the Group II mechanism. Arch. Biochem. Biophys. 474, 331–339 (2008).
Reissmann, S., Parnot, C., Booth, C.R., Chiu, W. & Frydman, J. Essential function of the built-in lid in the allosteric regulation of eukaryotic and archaeal chaperonins. Nat. Struct. Mol. Biol. 14, 432–440 (2007).
Yifrach, O. & Horovitz, A. Two lines of allosteric communication in the oligomeric chaperonin GroEL are revealed by the single mutation Arg196→Ala. J. Mol. Biol. 243, 397–401 (1994).
Fodor, A.A. & Aldrich, R.W. Influence of conservation on calculations of amino acid covariance in multiple sequence alignments. Proteins 56, 211–221 (2004).
Teşileanu, T., Colwell, L.J. & Leibler, S. Protein sectors: statistical coupling analysis versus conservation. PLoS Comput. Biol. 11, e1004091 (2015).
Martin, L.C., Gloor, G.B., Dunn, S.D. & Wahl, L.M. Using information theory to search for co-evolving residues in proteins. Bioinformatics 21, 4116–4124 (2005).
Dunn, S.D., Wahl, L.M. & Gloor, G.B. Mutual information without the influence of phylogeny or entropy dramatically improves residue contact prediction. Bioinformatics 24, 333–340 (2008).
Mao, W., Kaya, C., Dutta, A., Horovitz, A. & Bahar, I. Comparative study of the effectiveness and limitations of current methods for detecting sequence coevolution. Bioinformatics 31, 1929–1937 (2015).
Hill, J.E., Penny, S.L., Crowell, K.G., Goh, S.H. & Hemmingsen, S.M. cpnDB: a chaperonin sequence database. Genome Res. 14, 1669–1675 (2004).
Sievers, F. et al. Fast, scalable generation of high-quality protein multiple sequence alignments using Clustal Omega. Mol. Syst. Biol. 7, 539 (2011).
Pereira, J.H. et al. Mechanism of nucleotide sensing in group II chaperonins. EMBO J. 31, 731–740 (2012).
Shah, R. et al. Replacement of GroEL in Escherichia coli by the group II chaperonin from the Archaeon Methanococcus maripaludis. J. Bacteriol. 198, 2692–2700 (2016).
Ditzel, L. et al. Crystal structure of the thermosome, the archaeal chaperonin and homolog of CCT. Cell 93, 125–138 (1998).
Bigotti, M.G., Bellamy, S.R.W. & Clarke, A.R. The asymmetric ATPase cycle of the thermosome: elucidation of the binding, hydrolysis and product-release steps. J. Mol. Biol. 362, 835–843 (2006).
Bigotti, M.G. & Clarke, A.R. Cooperativity in the thermosome. J. Mol. Biol. 348, 13–26 (2005).
Zhang, J. et al. Cryo-EM structure of a group II chaperonin in the prehydrolysis ATP-bound state leading to lid closure. Structure 19, 633–639 (2011).
Pereira, J.H. et al. Crystal structures of a group II chaperonin reveal the open and closed states associated with the protein folding cycle. J. Biol. Chem. 285, 27958–27966 (2010).
Zhang, J. et al. Mechanism of folding chamber closure in a group II chaperonin. Nature 463, 379–383 (2010).
Douglas, N.R. et al. Dual action of ATP hydrolysis couples lid closure to substrate release into the group II chaperonin chamber. Cell 144, 240–252 (2011).
Kusmierczyk, A.R. & Martin, J. Nested cooperativity and salt dependence of the ATPase activity of the archaeal chaperonin Mm-cpn. FEBS Lett. 547, 201–204 (2003).
Kusmierczyk, A.R. & Martin, J. Nucleotide-dependent protein folding in the type II chaperonin from the mesophilic archaeon Methanococcus maripaludis. Biochem. J. 371, 669–673 (2003).
Hua, Q. et al. A thermophilic mini-chaperonin contains a conserved polypeptide-binding surface: combined crystallographic and NMR studies of the GroEL apical domain with implications for substrate interactions. J. Mol. Biol. 306, 513–525 (2001).
Sontag, E.M. et al. Exogenous delivery of chaperonin subunit fragment ApiCCT1 modulates mutant Huntingtin cellular phenotypes. Proc. Natl. Acad. Sci. USA 110, 3077–3082 (2013).
Spiess, C., Miller, E.J., McClellan, A.J. & Frydman, J. Identification of the TRiC/CCT substrate binding sites uncovers the function of subunit diversity in eukaryotic chaperonins. Mol. Cell 24, 25–37 (2006).
Weber, F., Keppel, F., Georgopoulos, C., Hayer-Hartl, M.K. & Hartl, F.U. The oligomeric structure of GroEL/GroES is required for biologically significant chaperonin function in protein folding. Nat. Struct. Biol. 5, 977–985 (1998).
Grallert, H., Rutkat, K. & Buchner, J. Limits of protein folding inside GroE complexes. J. Biol. Chem. 275, 20424–20430 (2000).
Hyeon, C., Lorimer, G.H. & Thirumalai, D. Dynamics of allosteric transitions in GroEL. Proc. Natl. Acad. Sci. USA 103, 18939–18944 (2006).
Meyer, A.S. et al. Closing the folding chamber of the eukaryotic chaperonin requires the transition state of ATP hydrolysis. Cell 113, 369–381 (2003).
Pilak, O. et al. Chaperonins from an Antarctic archaeon are predominantly monomeric: crystal structure of an open state monomer. Environ. Microbiol. 13, 2232–2249 (2011).
Reissmann, S. et al. A gradient of ATP affinities generates an asymmetric power stroke driving the chaperonin TRIC/CCT folding cycle. Cell Rep. 2, 866–877 (2012).
Leitner, A. et al. The molecular architecture of the eukaryotic chaperonin TRiC/CCT. Structure 20, 814–825 (2012).
Kalisman, N., Adams, C.M. & Levitt, M. Subunit order of eukaryotic TRiC/CCT chaperonin by cross-linking, mass spectrometry, and combinatorial homology modeling. Proc. Natl. Acad. Sci. USA 109, 2884–2889 (2012).
Nishikori, S., Esaki, M., Yamanaka, K., Sugimoto, S. & Ogura, T. Positive cooperativity of the p97 AAA ATPase is critical for essential functions. J. Biol. Chem. 286, 15815–15820 (2011).
Sen, M. et al. The ClpXP protease unfolds substrates using a constant rate of pulling but different gears. Cell 155, 636–646 (2013).
Song, C., Wang, Q. & Li, C.-C.H. ATPase activity of p97-valosin-containing protein (VCP). D2 mediates the major enzyme activity, and D1 contributes to the heat-induced activity. J. Biol. Chem. 278, 3648–3655 (2003).
Talavera, D., Lovell, S.C. & Whelan, S. Covariation is a poor measure of molecular coevolution. Mol. Biol. Evol. 32, 2456–2468 (2015).
Lunt, B, Szurmant, H., Procaccini, A, Hoch, J.A., Hwa, T. & Weigt, M. Inference of direct residue contacts in two-component signaling. Methods Enzymol. 471, 17–41 (2010).
Dwyer, R.S., Ricci, D.P., Colwell, L.J., Silhavy, T.J. & Wingreen, N.S. Predicting functionally informative mutations in Escherichia coli BamA using evolutionary covariance analysis. Genetics 195, 443–455 (2013).
Balakrishnan, S., Kamisetty, H., Carbonell, J.G., Lee, S.-I. & Langmead, C.J. Learning generative models for protein fold families. Proteins 79, 1061–1078 (2011).
Morcos, F., Hwa, T., Onuchic, J.N. & Weigt, M. in Protein Structure Prediction (ed. Kihara, D.) 55–70 (Springer, New York, 2014).
Jones, D.T., Buchan, D.W.A., Cozzetto, D. & Pontil, M. PSICOV: precise structural contact prediction using sparse inverse covariance estimation on large multiple sequence alignments. Bioinformatics 28, 184–190 (2012).
Ovchinnikov, S. et al. Large-scale determination of previously unsolved protein structures using evolutionary information. eLife 4, e09248 (2015).
Cong, Y. et al. Symmetry-free cryo-EM structures of the chaperonin TRiC along its ATPase-driven conformational cycle. EMBO J. 31, 720–730 (2012).
Weber, F. & Hayer-Hartl, M. in Chaperonin Protocols (ed. Schneider, C.) 117–126 (Springer, New York, 2000).
Acknowledgements
We thank N. Douglas for advice and manuscript comments, R. Pfuetzner and A. Brunger for help with the SEC-MALS experiments, A. Colavin and K. Huang for discussions of the coupling algorithms and members of the Frydman lab for helpful discussions. Research was supported by the NIH awards GM074074 (JF), GM062868 (VP) and by award DE-SC0008504 from the Department of Energy. K.D. was a recipient of a Stanford Graduate Fellowship, and T.L. was supported by T32GM007276.
Author information
Authors and Affiliations
Contributions
T.L. did all experiments, K.D. developed the algorithm and did analyses, A.T. assisted with light scattering measurements, V.P. directed algorithm development and J.F. directed all aspects of work. All authors contributed to the final manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Integrated supplementary information
Supplementary Figure 1 Comparison of covariation measures
Using synthetic alignments, we investigate the entropy dependence of various covariation measures including normalized mutual information (NMI), average product corrected normalized mutual information (APC-NMI), and statistical coupling analysis version 5 (SCA5).
(A) The entropy dependence for a range of covariation scenarios. Covariation for the 500 sequence alignment with two residue types shown on the left was computed with four common measures. The entropy dependence of the upper right quadrant for the measure is plotted in the bottom row. Restricting analysis to this quadrant avoids including coupling between identical columns. 2-D histograms report the average value of the measure for a combination of marginal entropies. Z-scores (number of standard deviations from the mean value for each distribution) are computed from the raw measure and thus lower scores imply less covariation and higher values of APCNMI indicate more covariation between residue pairs. Negative z-scores correspond to values which are below the mean of the distribution encompassing all residue pairs for the alignment. The Results show that normalized and average product corrected normalized mutual information have a comparatively flat dependence on the entropy of the input residues and favor residue pairs with similar entropies.
(B) Noise sensitivity of measures was evaluated by calculating the measures on an alignment of two residue types over a range of entropies generated by randomized insertion of the second residue type. The values were converted into Z-scores before binning to aid in comparison. The results show that APC-NMI has a very flat dependence on marginal entropy and tends to downweight couples with high entropy.
Supplementary Figure 2 Covariation of the archaeal chaperonin alignment
(A) Alignment of 573 archaeal chaperonin peptide sequences from the CpnDB.
(B) Approximate maximum likelihood tree of the archaeal chaperonin sequences. Generated using FasTree on codon aligned nucleotide sequences of the same 573 taxa illustrated in the peptide sequence (A). Render from FigTree.
(C) Comparison of average product corrected normalized mutual information (APC-NMI, bottom) with normalized mutual information (NMI, top) computed for the alignment in panel A. Measures are presented as z-scores to ease comparison. Higher scores are more significant.
(D) Comparison of average product corrected normalized mutual information (APC-NMI, bottom) with normalized average product corrected mutual information (MIp, top) computed for the alignment in panel A. Measures are presented as z-scores to ease comparison. Higher scores are more significant.
Supplementary Figure 3 Spectral Decomposition of APC-NMI
(A)The eigenvalues of the APC-NMI matrix computed from the alignment of archaeal chaperonins.
(B) Comparison of two most significant eigenvectors of APC-NMI. Each dot is colored by the Shannon entropy of the corresponding residue from the protein MSA (Figure S2A).
(C) Comparison of the second and third most significant eigenvectors of APC-NMI. Each dot is colored by the Shannon entropy of the corresponding residue from the protein MSA (Figure S2A).
Supplementary Figure 4 Summary of subnetworks
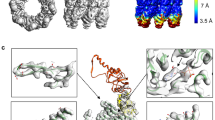
(A) (Left) Surface rendering all residues participating in couples below a bootstrap p-value threshold of 0.005 shown on the structure of the closed MmCpn (PDBID: 3RUW). Residues are colored by the first component of the APC-NMI matrix. (Right) Residues viewed on a single ring looking at the base of the equatorial domain towards the chamber lid along the eight-fold symmetry axis.
(B) All subnetworks from Figure 1F have been given identifiers for observation in the pymol session included in supplemental data set 1 and the renderings in panel C.
(C) Small subnetworks that correlate strongly with the first component of the APC-NMI matrix shown on single Cpn subunits (Upper) and as enhanced views (Lower).
Supplementary Figure 5 Binding site architecture for Met-47
(A) Surface rendering of binding site filled by Met47 (left) and alone (right).
(B) Hypothetical structures of TRiC-like mutants, M47I (left) and M47L (right). Rotamers were limited to only those that were attainable at the observed backbone angles.
(C) Mutants predicted to interfere with the steric architecture of the binding site. (left) Mutation to alanine would empty the binding site. (right) Deleting the residue responsible for creating the constricted binding site, Ile-512, would similarly create a largely empty pocket.
Supplementary Figure 6 HPLC-SEC-MALLS of chaperonin mutants
(A) Size exclusion chromatography of Cpn. Mutants were run on 500Å gel filtration column, as in Figure 2B.
(B) Calculated molecular masses and observed polydispersities by SEC-MALLS/QELS shown with the calculated uncertainty based on Debye plot fit.
Supplementary Figure 7 Nucleotide cycling of WT and M47L
(A) Nucleotide hydrolysis measured by an enzyme coupled assay at 1 mM ATP, from Figure 3C. ADP generated calculated by monitoring NADH oxidation at 340 nm. Single representative shown.
(B) Hill equation parameters for the first allosteric transition of nucleotide cycling with the standard deviation of three trials.
Supplementary Figure 8 Nucleotide hydrolyzed by the chaperonin
Amount of hydrolyzed nucleotide calculated as the difference between α- and γ-labeled 32P-ATP recoveries from Figure 4B i-iii.
Supplementary Figure 9 Suppression of Substrate Aggregation
Substrate aggregation monitored after dilution from denaturant seen in Figure 5B.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–9 and Supplementary Note 1 (PDF 1824 kb)
Supplementary Data set 1
Identified covarying networks from APC-NMI analysis of archaeal group II chaperonins. All identified subnetworks from Figure 1F and S4B shown on Cpn from M. maripaludis. PDBID: 3RUW. (ZIP 6390 kb)
Rights and permissions
About this article
Cite this article
Lopez, T., Dalton, K., Tomlinson, A. et al. An information theoretic framework reveals a tunable allosteric network in group II chaperonins. Nat Struct Mol Biol 24, 726–733 (2017). https://doi.org/10.1038/nsmb.3440
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nsmb.3440
This article is cited by
-
Snapshots of actin and tubulin folding inside the TRiC chaperonin
Nature Structural & Molecular Biology (2022)
-
CryoEM reveals the stochastic nature of individual ATP binding events in a group II chaperonin
Nature Communications (2021)
-
REP-X: An Evolution-guided Strategy for the Rational Design of Cysteine-less Protein Variants
Scientific Reports (2020)
-
Identification of an allosteric network that influences assembly and function of group II chaperonins
Nature Structural & Molecular Biology (2017)