Abstract
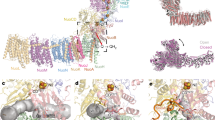
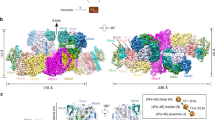
Bacterial polysulfide reductase (PsrABC) is an integral membrane protein complex responsible for quinone-coupled reduction of polysulfide, a process important in extreme environments such as deep-sea vents and hot springs. We determined the structure of polysulfide reductase from Thermus thermophilus at 2.4-Å resolution, revealing how the PsrA subunit recognizes and reduces its unique polyanionic substrate. The integral membrane subunit PsrC was characterized using the natural substrate menaquinone-7 and inhibitors, providing a comprehensive representation of a quinone binding site and revealing the presence of a water-filled cavity connecting the quinone binding site on the periplasmic side to the cytoplasm. These results suggest that polysulfide reductase could be a key energy-conserving enzyme of the T. thermophilus respiratory chain, using polysulfide as the terminal electron acceptor and pumping protons across the membrane via a previously unknown mechanism.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$189.00 per year
only $15.75 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Henne, A. et al. The genome sequence of the extreme thermophile Thermus thermophilus . Nat. Biotechnol. 22, 547–553 (2004).
Yano, T., Chu, S.S., Sled', V.D., Ohnishi, T. & Yagi, T. The proton-translocating NADH-quinone oxidoreductase (NDH-1) of thermophilic bacterium Thermus thermophilus HB-8. Complete DNA sequence of the gene cluster and thermostable properties of the expressed NQO2 subunit. J. Biol. Chem. 272, 4201–4211 (1997).
Collins, M.D., Shah, H.N. & Minnikin, D.E. A note on the separation of natural mixtures of bacterial menaquinones using reverse phase thin-layer chromatography. J. Appl. Bacteriol. 48, 277–282 (1980).
Hon-Nami, K. & Oshima, T. Purification and some properties of cytochrome c 552 from an extreme thermophile, Thermus thermophilus HB8. J. Biochem. 82, 769–776 (1977).
Mooser, D. et al. A four-subunit cytochrome bc 1 complex complements the respiratory chain of Thermus thermophilus . Biochim. Biophys. Acta 1708, 262–274 (2005).
Zimmermann, B.H., Nitsche, C.I., Fee, J.A., Rusnak, F. & Munck, E. Properties of a copper-containing cytochrome ba 3: a second terminal oxidase from the extreme thermophile Thermus thermophilus . Proc. Natl. Acad. Sci. USA 85, 5779–5783 (1988).
Fee, J.A., Yoshida, T., Surerus, K.K. & Mather, M.W. Cytochrome caa 3 from the thermophilic bacterium Thermus thermophilus: a member of the heme-copper oxidase superfamily. J. Bioenerg. Biomembr. 25, 103–114 (1993).
Soulimane, T. et al. Structure and mechanism of the aberrant ba 3-cytochrome c oxidase from Thermus thermophilus . EMBO J. 19, 1766–1776 (2000).
Keightley, J.A. et al. Molecular genetic and protein chemical characterization of the cytochrome ba 3 from Thermus thermophilus HB8. J. Biol. Chem. 270, 21016–21020 (1995).
Fee, J.A., Choc, M.G., Findling, K.L., Lorence, R. & Yoshida, T. Properties of a copper-containing cytochrome c1aa3 complex: a terminal oxidase of the extreme thermophile Thermus thermophilus HB8. Proc. Natl. Acad. Sci. USA 77, 147–151 (1980).
Brock, T.D. Thermophilic microorganisms and life at high temperatures. (Springer, New York, 1978).
Blöchl, E. et al. Pyrolobus fumarii, gen. and sp. nov., represents a novel group of archaea, extending the upper temperature limit for life to 113 °C. Extremophiles 1, 14–21 (1997).
Xu, Y., Schoonen, M.A.A., Nordstrom, D.K., Cunningham, K.M. & Ball, J.W. Sulfur geochemistry of hydrothermal waters in Yellowstone National Park, Wyoming, USA II. Formation and decomposition of thiosulfate and polythionate in Cinder Pool. J. Volcanol. Geotherm. Res. 97, 407–423 (2000).
Rothery, R.A., Workun, G.J. & Weiner, J.H. The prokaryotic complex iron-sulfur molybdoenzyme family. Biochim. Biophys. Acta published online, doi:10.1016/j.bbamem.2007.09.002 (18 September 2007).
Rothery, R.A., Kalra, N., Turner, R.J. & Weiner, J.H. Sequence similarity as a predictor of the transmembrane topology of membrane-intrinsic subunits of bacterial respiratory chain enzymes. J. Mol. Microbiol. Biotechnol. 4, 133–150 (2002).
Jormakka, M., Törnroth, S., Byrne, B. & Iwata, S. Molecular basis of proton motive force generation: structure of formate dehydrogenase-N. Science 295, 1863–1868 (2002).
Bertero, M.G. et al. Insights into the respiratory electron transfer pathway from the structure of nitrate reductase A. Nat. Struct. Biol. 10, 681–687 (2003).
Gavira, M., Roldán, M.D., Castillo, F. & Moreno-Vivián, C. Regulation of nap gene expression and periplasmic nitrate reductase activity in the phototrophic bacterium Rhodobacter sphaeroides DSM158. J. Bacteriol. 184, 1693–1702 (2002).
Ellington, M.J. et al. Characterization of the expression and activity of the periplasmic nitrate reductase of Paracoccus pantotrophus in chemostat cultures. Microbiology 149, 1533–1540 (2003).
Ansaldi, M., Théraulaz, L., Baraquet, C., Panis, G. & Méjean, V. Aerobic TMAO respiration in Escherichia coli . Mol. Microbiol. 66, 484–494 (2007).
Raaijmakers, H.C. & Romão, M.J. Formate-reduced E. coli formate dehydrogenase H: the reinterpretation of the crystal structure suggests a new reaction mechanism. J. Biol. Inorg. Chem. 11, 849–854 (2006).
McMaster, J. & Enemark, J.H. The active sites of molybdenum- and tungsten-containing enzymes. Curr. Opin. Chem. Biol. 2, 201–207 (1998).
Hille, R. The mononuclear molybdenum enzymes. Chem. Rev. 96, 2757–2816 (1996).
Weiner, J.H., Shaw, G., Turner, R.J. & Trieber, C.A. The topology of the anchor subunit of dimethyl sulfoxide reductase of Escherichia coli . J. Biol. Chem. 268, 3238–3244 (1993).
Santini, C.L. et al. A novel Sec-independent periplasmic protein translocation pathway in Escherichia coli . EMBO J. 17, 101–112 (1998).
Schindelin, H., Kisker, C., Hilton, J., Rajagopalan, K.V. & Rees, D.C. Crystal structure of DMSO reductase: redox-linked changes in molybdopterin coordination. Science 272, 1615–1621 (1996).
Jones, T.A., Zou, J.Y., Cowan, S.W. & Kjeldgaard, M. Improved methods for building protein models in electron density maps and the location of errors in these models. Acta Crystallogr. A 47, 110–119 (1991).
Simala-Grant, J.L. & Weiner, J.H. Modulation of the substrate specificity of Escherichia coli dimethylsulfoxide reductase. Eur. J. Biochem. 251, 510–515 (1998).
Berglund, G.I. et al. The catalytic pathway of horseradish peroxidase at high resolution. Nature 417, 463–468 (2002).
Matsui, T., Ozaki, S., Liong, E., Phillips, G.N. & Watanabe, Y. Effects of the location of distal histidine in the reaction of myoglobin with hydrogen peroxide. J. Biol. Chem. 274, 2838–2844 (1999).
Ridge, J.P., Aguey-Zinsou, K.F., Bernhardt, P.V., Hanson, G.R. & McEwan, A.G. The critical role of tryptophan-116 in the catalytic cycle of dimethylsulfoxide reductase from Rhodobacter capsulatus . FEBS Lett. 563, 197–202 (2004).
Hille, R. Molybdenum enzymes. Essays Biochem. 34, 125–137 (1999).
Hensel, M., Hinsley, A.P., Nikolaus, T., Sawers, G. & Berks, B.C. The genetic basis of tetrathionate respiration in Salmonella typhimurium . Mol. Microbiol. 32, 275–287 (1999).
Nagarajan, K. et al. Structural and functional analogue of the active site of polysulfide reductase from Wolinella succinogenes . Inorg. Chem. 43, 4532–4533 (2004).
Berks, B.C. et al. Sequence analysis of subunits of the membrane-bound nitrate reductase from a denitrifying bacterium: the integral membrane subunit provides a prototype for the dihaem electron-carrying arm of a redox loop. Mol. Microbiol. 15, 319–331 (1995).
Page, C.C., Moser, C.C., Chen, X. & Dutton, P.L. Natural engineering principles of electron tunneling in biological oxidation-reduction. Nature 402, 47–52 (1999).
Cheng, V.W., Rothery, R.A., Bertero, M.G., Strynadka, N.C. & Weiner, J.H. Investigation of the environment surrounding iron-sulfur cluster 4 of Escherichia coli dimethylsulfoxide reductase. Biochemistry 44, 8068–8077 (2005).
Dietrich, W. & Klimmek, O. The function of methyl-menaquinone-6 and polysulfide reductase membrane anchor (PsrC) in polysulfide respiration of Wolinella succinogenes . Eur. J. Biochem. 269, 1086–1095 (2002).
Szilágyi, A. & Závodszky, P. Structural differences between mesophilic, moderately thermophilic and extremely thermophilic protein subunits: results of a comprehensive survey. Structure 8, 493–504 (2000).
Yankovskaya, V. et al. Architecture of succinate dehydrogenase and reactive oxygen species generation. Science 299, 700–704 (2003).
Lancaster, C.R., Kröger, A., Auer, M. & Michel, H. Structure of fumarate reductase from Wolinella succinogenes at 2.2 Å resolution. Nature 402, 377–385 (1999).
Iverson, T.M., Luna-Chavez, C., Cecchini, G. & Rees, D.C. Structure of the Escherichia coli fumarate reductase respiratory complex. Science 284, 1961–1966 (1999).
Sun, F. et al. Crystal structure of mitochondrial respiratory membrane protein complex II. Cell 121, 1043–1057 (2005).
Huang, L.S. et al. 3-nitropropionic acid is a suicide inhibitor of mitochondrial respiration that, upon oxidation by complex II, forms a covalent adduct with a catalytic base arginine in the active site of the enzyme. J. Biol. Chem. 281, 5965–5972 (2006).
Horsefield, R. et al. Structural and computational analysis of the quinone-binding site of complex II (succinate-ubiquinone oxidoreductase): a mechanism of electron transfer and proton conduction during ubiquinone reduction. J. Biol. Chem. 281, 7309–7316 (2006).
Hedderich, R. et al. Anaerobic respiration with elemental sulfur and with disulfides. FEMS Microbiol. Rev. 22, 353–381 (1998).
Iwata, S., Ostermeier, C., Ludwig, B. & Michel, H. Structure at 2.8 resolution of cytochrome C oxidase from Paracoccus denitrificans . Nature 376, 660–669 (1995).
Tsukihara, T. et al. The whole structure of the 13-subunit oxidized cytochrome C oxidase at 2.8. Science 272, 1136–1144 (1996).
Holt, P.J., Morgan, D.J. & Sazanov, L.A. The location of NuoL and NuoM subunits in the membrane domain of the Escherichia coli complex I: implications for the mechanism of proton pumping. J. Biol. Chem. 278, 43114–43120 (2003).
Ohnishi, T. & Salerno, J.C. Conformation-driven and semiquinone-gated proton-pump mechanism in the NADH-ubiquinone oxidoreductase (complex I). FEBS Lett. 579, 4555–4561 (2005).
Bogachev, A.V., Murtazina, R.A. & Skulachev, V.P.H. +/e− stoichiometry for NADH dehydrogenase I and dimethyl solfoxide reductase in anaerobically grown Escherichia coli cells. J. Bacteriol. 178, 6233–6237 (1996).
Yokoyama, K., Akabane, Y., Ishii, N. & Yoshida, M. Isolation of prokaryotic V0V1-ATPase from a thermophilic eubacterium Thermus thermophilus . J. Biol. Chem. 269, 12248–12253 (1994).
Otwinowski, Z. & Minor, W. Processing of x-ray diffraction data collected in oscillation mode. Methods Enzymol. 276, 307–326 (1997).
Leslie, A.G.W. Recent changes to the MOSFLM package for processing film and image plate data. Joint CCP4 and ESF-EAMCB Newsletter on Protein Crystallography, No. 26 (Daresbury Laboratory, Warrington, UK, 1992).
Collaborative Computational Project. Number 4. The CCP4 Suite: Programs for Protein Crystallography. Acta Crystallogr. D50, 760–763 (1994).
Weeks, C.M. & Miller, R. The design and implementation of SnB v2.0. J. Appl. Crystallogr. 32, 120–124 (1999).
Perrakis, A., Morris, R.M. & Lamzin, V.S. Automated protein model building combined with iterative structure refinement. Nat. Struct. Biol. 6, 458–463 (1999).
Brünger, A.T. et al. Crystallography & NMR system: a new software suite for macromolecular structure determination. Acta Crystallogr. D Biol. Crystallogr. 54, 905–921 (1998).
Laskowski, R.A., MacArthur, M.W., Moss, D.S. & Thornton, J.M. PROCHECK—a program to check the stereochemical quality of protein structures. J. Appl. Crystallogr. 26, 283–291 (1993).
Acknowledgements
This study was supported by the Australian Research Council (ARC; DP0666970 to M.J.), the European Membrane Protein Consortium (EU-PF6 E-MEP to S.I.) and Grants-in-Aid from the Ministry of Education, Science, Sports and Culture of Japan (No. 18370055 and No. 19042008 to K.Y.). M.J. was a recipient of the European Molecular Biology Organization (EMBO) Long-term Fellowship. Data collection was done at the European Synchrotron Radiation Facility (ESRF) and GM/CA-CAT at Advanced Photon Source (APS). We acknowledge the support of S. Corcoran, N. Venugopolan and M. Becker at GM/CA-CAT, and beamline scientists at ID23-1, ESRF, for scientific support and assistance with data collection. GM/CA-CAT beamline (ID23) is supported by the US National Cancer Institute and the US National Institute of General Medical Science. Visits to APS were supported by the Australian Nuclear Science Technology Organization (ANSTO). Part of the work was performed at the Membrane Protein Laboratory at Diamond Light Source, UK, which is funded by Wellcome Trust grant 079209/Z/06/Z. We thank U. Kappler, M. Maher, M. Rapp, B. Byrne and L. Carpenter for critical assessment of the manuscript.
Author information
Authors and Affiliations
Contributions
K.Y. and M.J. designed the research; K.Y. and T.Y. oversaw experimental design of biochemical studies, largely performed by S.A.; M.T. performed protein expression and optimization; T.S. generated antibodies for protein detection; P.C. assisted with data collection and interpretation; M.J. performed Psr purification, crystallization, data collection and structure determination; M.J. and S.I. performed initial phasing and structure interpretation. M.J. and S.I. prepared the manuscript, and all authors discussed the results and commented on the manuscript.
Corresponding authors
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–4 (PDF 1959 kb)
Rights and permissions
About this article
Cite this article
Jormakka, M., Yokoyama, K., Yano, T. et al. Molecular mechanism of energy conservation in polysulfide respiration. Nat Struct Mol Biol 15, 730–737 (2008). https://doi.org/10.1038/nsmb.1434
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nsmb.1434
This article is cited by
-
Active anaerobic methane oxidation and sulfur disproportionation in the deep terrestrial subsurface
The ISME Journal (2022)
-
An unprecedented function for a tungsten-containing oxidoreductase
JBIC Journal of Biological Inorganic Chemistry (2022)
-
Active sulfur cycling in the terrestrial deep subsurface
The ISME Journal (2020)
-
Single cell electron collectors for highly efficient wiring-up electronic abiotic/biotic interfaces
Nature Communications (2020)
-
Structure of the respiratory MBS complex reveals iron-sulfur cluster catalyzed sulfane sulfur reduction in ancient life
Nature Communications (2020)