Key Points
-
Immunoglobulin class-switch recombination (CSR) is a B-cell specific DNA-rearrangement reaction that exchanges the expressed Ig heavy-chain constant-region gene (CH) from Cμ to another downstream CH gene. This results in the generation of antibodies of secondary isotypes (IgG, IgA or IgE), which have different effector functions.
-
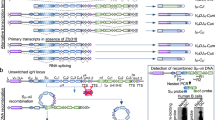
CSR occurs between repetitive DNA elements called switch (S) regions that precede each of the constant-region genes. The reaction probably proceeds through the introduction of double-strand breaks (DSBs) in the DNA of two participating S regions, followed by their fusion. During this process, the DNA between the participating S regions is lost as an extrachromosomal circle.
-
Transcription through the participating S regions is a prerequisite for CSR. The germline transcripts do not encode proteins but might have direct mechanistic roles in CSR, by forming RNA–DNA hybrids, thereby generating higher-order secondary structures. One such structure, the R loop, in which the non-template strand remains single stranded, has been demonstrated to form both in vitro and in vivo following the transcription of S regions. Other transcription-generated structures might also have roles in CSR.
-
Somatic hypermutation (SHM) is another B cell-specific reaction that introduces point mutations into the antigen-binding segment of immunoglobulin genes to allow the generation of higher-affinity antibodies.
-
Activation-induced cytidine deaminase (AID), recently identified as an enzyme expressed specifically by activated B cells, is essential for CSR and SHM. In mice and humans, inactivating mutations of AID lead to an absolute block in CSR and SHM. Ectopic expression of AID in fibroblasts can induce SHM and CSR in artificial substrates.
-
The role of AID in CSR has been intensely debated since its discovery : it has been speculated to be an mRNA-editing enzyme and also to be a DNA-cytidine deaminase. There are now strong genetic and biochemical data to support the latter possibility.
-
Recent work has provided evidence that single-stranded DNA that is generated during the transcription of S regions provides DNA substrates for AID deaminase activity. The deaminated DNA might be processed by components of the base-excision repair and mismatch-repair machineries to generate DSBs. The fusion of the two S region DSBs is probably mediated by non-homologous DNA end joining.
-
Although the discovery of AID has considerably advanced our understanding of CSR and SHM, many issues remain unresolved, including the specificity of AID activity, the mechanism by which AID recognizes its targets, and the process that converts the deaminated DNA into DSBs and ultimately resolves the DNA lesions to complete CSR. A combination of in vivo and in vitro biochemical approaches should help address these issues.
Abstract
Class-switch recombination (CSR) of immunoglobulin heavy chains is the genetic process by which a B cell switches from the production of IgM to the production of IgG, IgE or IgA. Although the general characteristics of CSR have been known for some time, the detailed molecular mechanism of this process is only now emerging. CSR is unique, in that it seems to involve transcription-generated, higher-order RNA–DNA structures, specific DNA deamination and several DNA-repair pathways. In this review, we discuss our current knowledge of the mechanism of CSR and highlight the important unanswered questions.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on SpringerLink
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
References
Bassing, C. H., Swat, W. & Alt, F. W. The mechanism and regulation of chromosomal V(D)J recombination. Cell 109, S45–S55 (2002).
Jung, D. & Alt, F. W. Unraveling V(D)J recombination: insights into gene regulation. Cell 116, 299–311 (2004).
Papavasiliou, F. N. & Schatz, D. G. Somatic hypermutation of immunoglobulin genes: merging mechanisms for genetic diversity. Cell 109, S35–S44 (2002).
Martin, A. & Scharff, M. D. AID and mismatch repair in antibody diversification. Nature Rev. Immunol. 2, 605–614 (2002).
Manis, J. P., Tian, M. & Alt, F. W. Mechanism and control of class-switch recombination. Trends Immunol. 23, 31–39 (2002).
Honjo, T., Kinoshita, K. & Muramatsu, M. Molecular mechanism of class switch recombination: linkage with somatic hypermutation. Annu. Rev. Immunol. 20, 165–196 (2002).
Storb, U. et al. Cis-acting sequences that affect somatic hypermutation of Ig genes. Immunol. Rev. 162, 153–160 (1998).
Lansford, R., Manis, J. P., Sonoda, E., Rajewsky, K. & Alt, F. W. Ig heavy chain class switching in Rag-deficient mice. Int. Immunol. 10, 325–332 (1998).
Dunnick, W., Hertz, G. Z., Scappino, L. & Gritzmacher, C. DNA sequences at immunoglobulin switch region recombination sites. Nucleic Acids Res. 21, 365–372 (1993).
Shinkura, R. et al. The influence of transcriptional orientation on endogenous switch region function. Nature Immunol. 4, 435–441 (2003). Provides the first evidence in vivo that the transcriptional orientation of S regions is important in CSR. In addition, the authors show that a synthetic G-rich sequence that lacks S-region motifs can replace an endogenous S region during CSR in vivo.
Chaudhuri, J. et al. Transcription-targeted DNA deamination by the AID antibody diversification enzyme. Nature 422, 726–730 (2003). Demonstrates biochemically that AID is a ssDNA deaminase. The authors also show a direct link between transcription and DNA deamination.
Neuberger, M. S., Harris, R. S., Di Noia, J. & Petersen-Mahrt, S. K. Immunity through DNA deamination. Trends Biochem. Sci. 28, 305–312 (2003).
Stavnezer, J. Immunoglobulin class switching. Curr. Opin. Immunol. 8, 199–205 (1996).
Luby, T. M., Schrader, C. E., Stavnezer, J. & Selsing, E. The μ switch region tandem repeats are important, but not required, for antibody class switch recombination. J. Exp. Med. 193, 159–168 (2001).
Stavnezer, J. Antibody class switching. Adv. Immunol. 61, 79–146 (1996).
Stavnezer, J. Molecular processes that regulate class switching. Curr. Top. Microbiol. Immunol. 245, 127–168 (2000).
Yancopoulos, G. D. et al. Secondary genomic rearrangement events in pre-B cells: VHDJH replacement by a LINE-1 sequence and directed class switching. EMBO J. 5, 3259–3266 (1986).
Stavnezer-Nordgren, J. & Sirlin, S. Specificity of immunoglobulin heavy chain switch correlates with activity of germline heavy chain genes prior to switching. EMBO J. 5, 95–102 (1986).
Reaban, M. E. & Griffin, J. A. Induction of RNA-stabilized DNA conformers by transcription of an immunoglobulin switch region. Nature 348, 342–344 (1990).
Reaban, M. E., Lebowitz, J. & Griffin, J. A. Transcription induces the formation of a stable RNA. DNA hybrid in the immunoglobulin α switch region. J. Biol. Chem. 269, 21850–21857 (1994).
Daniels, G. A. & Lieber, M. R. Strand specificity in the transcriptional targeting of recombination at immunoglobulin switch sequences. Proc. Natl Acad. Sci. USA 92, 5625–5629 (1995).
Tian, M. & Alt, F. W. Transcription-induced cleavage of immunoglobulin switch regions by nucleotide excision repair nucleases in vitro. J. Biol. Chem. 275, 24163–24172 (2000).
Daniels, G. A. & Lieber, M. R. RNA:DNA complex formation upon transcription of immunoglobulin switch regions: implications for the mechanism and regulation of class switch recombination. Nucleic Acids Res. 23, 5006–5011 (1995).
Mizuta, R. et al. Molecular visualization of immunoglobulin switch region RNA/DNA complex by atomic force microscope. J. Biol. Chem. 278, 4431–4434 (2003).
Yu, K., Chedin, F., Hsieh, C. L., Wilson, T. E. & Lieber, M. R. R-loops at immunoglobulin class switch regions in the chromosomes of stimulated B cells. Nature Immunol. 4, 442–451 (2003). Uses chemical probes to demonstrate that S regions form R loops in vivo in B cells undergoing CSR.
Tashiro, J., Kinoshita, K. & Honjo, T. Palindromic but not G-rich sequences are targets of class switch recombination. Int. Immunol. 13, 495–505 (2001).
Sen, D. & Gilbert, W. Formation of parallel four- stranded complexes by guanine-rich motifs in DNA and its implications for meiosis. Nature 334, 364–366 (1988).
Dempsey, L. A., Sun, H., Hanakahi, L. A. & Maizels, N. G4 DNA binding by LR1 and its subunits, nucleolin and hnRNP D, a role for G–G pairing in immunoglobulin switch recombination. J. Biol. Chem. 274, 1066–1071 (1999).
Ratmeyer, L., Vinayak, R., Zhong, Y. Y., Zon, G. & Wilson, W. D. Sequence specific thermodynamic and structural properties for DNA. RNA duplexes. Biochemistry 33, 5298–5304 (1994).
Kinoshita, K., Tashiro, J., Tomita, S., Lee, C. G. & Honjo, T. Target specificity of immunoglobulin class switch recombination is not determined by nucleotide sequences of S regions. Immunity 9, 849–858 (1998).
Mussmann, R., Courtet, M., Schwager, J. & Du Pasquier, L. Microsites for immunoglobulin switch recombination breakpoints from Xenopus to mammals. Eur. J. Immunol. 27, 2610–2619 (1997).
Ramiro, A. R., Stavropoulos, P., Jankovic, M. & Nussenzweig, M. C. Transcription enhances AID-mediated cytidine deamination by exposing single-stranded DNA on the nontemplate strand. Nature Immunol. 4, 452–456 (2003). Shows that AID targets the non-template strand of DNA when transcription is carried out in bacteria.
Sohail, A., Klapacz, J., Samaranayake, M., Ullah, A. & Bhagwat, A. S. Human activation-induced cytidine deaminase causes transcription-dependent, strand-biased C to U deaminations. Nucleic Acids Res. 31, 2990–2994 (2003).
Pham, P., Bransteitter, R., Petruska, J. & Goodman, M. F. Processive AID-catalysed cytosine deamination on single-stranded DNA simulates somatic hypermutation. Nature 424, 103–107 (2003). Shows that AID preferably deaminates cytidine residues within SHM motifs in an ordered fashion.
Muramatsu, M. et al. Specific expression of activation-induced cytidine deaminase (AID), a novel member of the RNA-editing deaminase family in germinal center B cells. J. Biol. Chem. 274, 18470–18476 (1999).
Muramatsu, M. et al. Class switch recombination and hypermutation require activation-induced cytidine deaminase (AID), a potential RNA editing enzyme. Cell 102, 553–563 (2000).
Revy, P. et al. Activation-induced cytidine deaminase (AID) deficiency causes the autosomal recessive form of the Hyper-IgM syndrome (HIGM2). Cell 102, 565–575 (2000).
Yoshikawa, K. et al. AID enzyme-induced hypermutation in an actively transcribed gene in fibroblasts. Science 296, 2033–2036 (2002).
Martin, A. & Scharff, M. D. Somatic hypermutation of the AID transgene in B and non-B cells. Proc. Natl Acad. Sci. USA 99, 12304–12308 (2002).
Okazaki, I. M., Kinoshita, K., Muramatsu, M., Yoshikawa, K. & Honjo, T. The AID enzyme induces class switch recombination in fibroblasts. Nature 416, 340–345 (2002). References 38 to 40 provide evidence that AID is probably the only activated B-cell factor required for SHM and CSR.
Doi, T., Kinoshita, K., Ikegawa, M., Muramatsu, M. & Honjo, T. De novo protein synthesis is required for the activation-induced cytidine deaminase function in class-switch recombination. Proc. Natl Acad. Sci. USA 100, 2634–2638 (2003).
Harris, R. S., Petersen-Mahrt, S. K. & Neuberger, M. S. RNA editing enzyme APOBEC1 and some of its homologs can act as DNA mutators. Mol. Cell 10, 1247–1253 (2002). Shows that other APOBEC1-family members have DNA-deamination activity, indicating that the RNA-editing enzyme APOBEC1 is probably an outlier in this family of proteins.
Petersen-Mahrt, S. K., Harris, R. S. & Neuberger, M. S. AID mutates E. coli suggesting a DNA deamination mechanism for antibody diversification. Nature 418, 99–103 (2002). The first paper to present the DNA-deamination model for CSR and SHM. In addition, it provides evidence that the target of AID is not RNA but DNA.
Bransteitter, R., Pham, P., Scharff, M. D. & Goodman, M. F. Activation-induced cytidine deaminase deaminates deoxycytidine on single-stranded DNA but requires the action of RNase. Proc. Natl Acad. Sci. USA 100, 4102–4107 (2003).
Dickerson, S. K., Market, E., Besmer, E. & Papavasiliou, F. N. AID mediates hypermutation by deaminating single stranded DNA. J. Exp. Med. 197, 1291–1296 (2003).
Nambu, Y. et al. Transcription-coupled events associating with immunoglobulin switch region chromatin. Science 302, 2137–2140 (2003). Shows that AID binds to chromatin that is associated with transcribed S regions, thereby providing further evidence that AID acts directly on DNA.
Wuerffel, R. A., Du, J., Thompson, R. J. & Kenter, A. L. Ig Sγ3 DNA-specific double strand breaks are induced in mitogen-activated B cells and are implicated in switch recombination. J. Immunol. 159, 4139–4144 (1997).
Catalan, N. et al. The block in immunoglobulin class switch recombination caused by activation-induced cytidine deaminase deficiency occurs prior to the generation of DNA double strand breaks in switch μ region. J. Immunol. 171, 2504–2509 (2003).
Rogakou, E. P., Pilch, D. R., Orr, A. H., Ivanova, V. S. & Bonner, W. M. DNA double-stranded breaks induce histone H2AX phosphorylation on serine 139. J. Biol. Chem. 273, 5858–5868 (1998).
Petersen, S. et al. AID is required to initiate Nbs1/γ-H2AX focus formation and mutations at sites of class switching. Nature 414, 660–665 (2001). Shows that H2AX has a role in CSR and that the role of AID in CSR is probably upstream of S-region breaks.
Bassing, C. H. et al. Histone H2AX: a dosage-dependent suppressor of oncogenic translocations and tumors. Cell 114, 359–370 (2003). Reports that haploinsufficiency of H2AX can lead to transclocations that involve S regions.
Manis, J. P. et al. 53BP1 links DNA damage response to immunoglobulin class switch recombination. Nature Immunol. 5, 481–487 (2004). Describes the effect of 53BP1 mutation in CSR and SHM.
Casellas, R. et al. Ku80 is required for immunoglobulin isotype switching. EMBO J. 17, 2404–2411 (1998).
Manis, J. P. et al. Ku70 is required for late B cell development and immunoglobulin heavy chain class switching. J. Exp. Med. 187, 2081–2089 (1998).
Manis, J. P., Dudley, D., Kaylor, L. & Alt, F. W. IgH class switch recombination to IgG1 in DNA-PKcs-deficient B Cells. Immunity 16, 607–617 (2002). Together with references 93 and 94, this paper indicates that the role of DNA-PKcs in CSR might extend beyond the role it has in NHEJ.
Bross, L., Muramatsu, M., Kinoshita, K., Honjo, T. & Jacobs, H. DNA double-strand breaks: prior to but not sufficient in targeting hypermutation. J. Exp. Med. 195, 1187–1192 (2002).
Papavasiliou, F. et al. V(D)J recombination in mature B cells: a mechanism for altering antibody responses. Science 278, 298–301 (1997).
Faili, A. et al. AID-dependent somatic hypermutation occurs as a DNA single-strand event in the BL2 cell line. Nature Immunol. 3, 815–821 (2002).
Woo, C. J., Martin, A. & Scharff, M. D. Induction of somatic hypermutation is associated with modifications in immunoglobulin variable region chromatin. Immunity 19, 479–489 (2003).
Rada, C. et al. Immunoglobulin isotype switching is inhibited and somatic hypermutation perturbed in UNG-deficient mice. Curr. Biol. 12, 1748–1755 (2002).
Imai, K. et al. Human uracil-DNA glycosylase deficiency associated with profoundly impaired immunoglobulin class-switch recombination. Nature Immunol. 4, 1023–1028 (2003). References 60 and 61 show that UNG has a crucial role in CSR and thereby provide evidence for a DNA-deamination step in CSR.
Vora, K. A. et al. Severe attenuation of the B cell immune response in Msh2-deficient mice. J. Exp. Med. 189, 471–482 (1999).
Ehrenstein, M. R. & Neuberger, M. S. Deficiency in Msh2 affects the efficiency and local sequence specificity of immunoglobulin class-switch recombination: parallels with somatic hypermutation. EMBO J. 18, 3484–3490 (1999).
Schrader, C. E., Edelmann, W., Kucherlapati, R. & Stavnezer, J. Reduced isotype switching in splenic B cells from mice deficient in mismatch repair enzymes. J. Exp. Med. 190, 323–330 (1999).
Yu, K. & Lieber, M. R. Nucleic acid structures and enzymes in the immunoglobulin class switch recombination mechanism. DNA Repair (Amst.) 2, 1163–1174 (2003).
Rush, J. S., Fugmann, S. D. & Schatz, D. G. Staggered AID-dependent DNA double strand breaks are the predominant DNA lesions targeted to Sμ in immunoglobulin class switch recombination. Int. Immunol. 16, 549–557 (2004).
Faili, A. et al. DNA Polymerase η is involved in hypermutation occurring during immunoglobulin class switch recombination. J. Exp. Med. 199, 265–270 (2004).
Schrader, C. E., Vardo, J. & Stavnezer, J. Role for mismatch repair proteins Msh2, Mlh1, and Pms2 in immunoglobulin class switching shown by sequence analysis of recombination junctions. J. Exp. Med. 195, 367–373 (2002).
Min, I. M. et al. The Sμ tandem repeat region is critical for Ig isotype switching in the absence of Msh2. Immunity 19, 515–524 (2003).
Paques, F. & Haber, J. E. Multiple pathways of recombination induced by double-strand breaks in Saccharomyces cerevisiae. Microbiol. Mol. Biol. Rev. 63, 349–404 (1999).
Tian, M., Jones, A. M., Smith, M., Shinkura, R. & Alt, F. W. Deficiency in nuclease activity of xeroderma pigmentosum G in mice leads to hypersensitivity to UV radiation. Mol. Cell. Biol. 24, 2237–2242 (2004).
Tian, M., Shinkura, R., Shinkura, N. & Alt, F. W. Growth retardation, early death, and DNA repair defects in mice deficient for the nucleotide excision repair enzyme XPF. Mol. Cell. Biol. 24, 1200–1205 (2004).
Hodgkin, P. D., Lee, J. H. & Lyons, A. B. B cell differentiation and isotype switching is related to division cycle number. J. Exp. Med. 184, 277–281 (1996).
Kuzminov, A. Single-strand interruptions in replicating chromosomes cause double-strand breaks. Proc. Natl Acad. Sci. USA 98, 8241–8246 (2001).
Dudley, D. D. et al. Internal IgH class switch region deletions are position-independent and enhanced by AID expression. Proc. Natl Acad. Sci. USA 99, 9984–9989 (2002). Shows that S-region internal deletions are AID dependent and provides evidence that synapsis between two S regions is not required for targeting AID.
Alt, F. W., Rosenberg, N., Casanova, R. J., Thomas, E. & Baltimore, D. Immunoglobulin heavy-chain expression and class switching in a murine leukaemia cell line. Nature 296, 325–331 (1982).
Gu, H., Zou, Y. R. & Rajewsky, K. Independent control of immunoglobulin switch recombination at individual switch regions evidenced through Cre–loxP-mediated gene targeting. Cell 73, 1155–1164 (1993).
Schrader, C. E. et al. Mutations occur in the Ig Sμ region but rarely in Sγ regions prior to class switch recombination. EMBO J. 22, 5893–5903 (2003).
Reina-San-Martin, B. et al. H2AX is required for recombination between immunoglobulin switch regions but not for intra-switch region recombination or somatic hypermutation. J. Exp. Med. 197, 1767–1778 (2003). Indicates that H2AX might have a role in the synapsis of two S regions during CSR.
Bassing, C. H. & Alt, F. W. H2AX may function as an anchor to hold broken chromosomal DNA ends in close proximity. Cell Cycle 3, 149–153 (2004).
Shiloh, Y. Ataxia-telangiectasia and the Nijmegen breakage syndrome: related disorders but genes apart. Annu. Rev. Genet. 31, 635–662 (1997).
Pan, Q. et al. Alternative end joining during switch recombination in patients with ataxia-telangiectasia. Eur. J. Immunol. 32, 1300–1308 (2002).
Pan-Hammarstrom, Q. et al. ATM is not required in somatic hypermutation of VH, but is involved in the introduction of mutations in the switch μ region. J. Immunol. 170, 3707–3716 (2003).
Burma, S., Chen, B. P., Murphy, M., Kurimasa, A. & Chen, D. J. ATM phosphorylates histone H2AX in response to DNA double-strand breaks. J. Biol. Chem. 276, 42462–42467 (2001).
Stiff, T. et al. ATM and DNA-PK function redundantly to phosphorylate H2AX after exposure to ionizing radiation. Cancer Res. 64, 2390–2396 (2004).
Ward, I. M. & Chen, J. Histone H2AX is phosphorylated in an ATR-dependent manner in response to replicational stress. J. Biol. Chem. 276, 47759–47762 (2001).
Xu, Y. et al. Targeted disruption of ATM leads to growth retardation, chromosomal fragmentation during meiosis, immune defects, and thymic lymphoma. Genes Dev. 10, 2411–2422 (1996).
DeFazio, L. G., Stansel, R. M., Griffith, J. D. & Chu, G. Synapsis of DNA ends by DNA-dependent protein kinase. EMBO J. 21, 3192–3200 (2002).
Ehrenstein, M. R., Rada, C., Jones, A. M., Milstein, C. & Neuberger, M. S. Switch junction sequences in PMS2-deficient mice reveal a microhomology-mediated mechanism of Ig class switch recombination. Proc. Natl Acad. Sci. USA 98, 14553–14558 (2001).
Hall, M. C., Wang, H., Erie, D. A. & Kunkel, T. A. High affinity cooperative DNA binding by the yeast Mlh1–Pms1 heterodimer. J. Mol. Biol. 312, 637–647 (2001).
Schrader, C. E., Vardo, J. & Stavnezer, J. Mlh1 can function in antibody class switch recombination independently of Msh2. J. Exp. Med. 197, 1377–1383 (2003).
Brown, K. D. et al. The mismatch repair system is required for S-phase checkpoint activation. Nature Genet. 33, 80–84 (2003).
Bosma, G. C. et al. DNA-dependent protein kinase activity is not required for immunoglobulin class switching. J. Exp. Med. 196, 1483–1495 (2002).
Cook, A. J. et al. Reduced switching in SCID B cells is associated with altered somatic mutation of recombined S regions. J. Immunol. 171, 6556–6564 (2003).
Ma, Y., Pannicke, U., Schwarz, K. & Lieber, M. R. Hairpin opening and overhang processing by an Artemis/DNA-dependent protein kinase complex in nonhomologous end joining and V(D)J recombination. Cell 108, 781–794 (2002).
Sekiguchi, J. et al. Genetic interactions between ATM and the nonhomologous end-joining factors in genomic stability and development. Proc. Natl Acad. Sci. USA 98, 3243–3248 (2001).
Ta, V. T. et al. AID mutant analyses indicate requirement for class-switch-specific cofactors. Nature Immunol. 4, 843–848 (2003).
Barreto, V., Reina-San-Martin, B., Ramiro, A. R., McBride, K. M. & Nussenzweig, M. C. C-terminal deletion of AID uncouples class switch recombination from somatic hypermutation and gene conversion. Mol. Cell 12, 501–508 (2003). References 97 and 98 provide evidence that the carboxyl terminus of AID could have specific roles in CSR that might be downstream of DNA deamination.
Bemark, M. et al. Somatic Hypermutation in the absence of DNA-dependent protein kinase catalytic subunit (DNA-PKcs) or recombination-activating gene (RAG) 1 activity. J. Exp. Med. 192, 1509–1514 (2000).
Imai, K. et al. Hyper-IgM syndrome type 4 with a B lymphocyte-intrinsic selective deficiency in Ig class- switch recombination. J. Clin. Invest. 112, 136–142 (2003). Reports on a newly characterized human immunodeficiency, in which the defect in CSR is probably downstream of AID-mediated DNA deamination.
Shanmugam, A., Shi, M. J., Yauch, L., Stavnezer, J. & Kenter, A. L. Evidence for class-specific factors in immunoglobulin isotype switching. J. Exp. Med. 191, 1365–1380 (2000).
Ma, L., Wortis, H. H. & Kenter, A. L. Two new isotype-specific switching activities detected for Ig class switching. J. Immunol. 168, 2835–2846 (2002).
Ito, S. et al. Activation-induced cytidine deaminase shuttles between nucleus and cytoplasm like apolipoprotein B mRNA editing catalytic polypeptide 1. Proc. Natl Acad. Sci. USA 101, 1975–1980 (2004).
Martin, A. et al. Msh2 ATPase activity is essential for somatic hypermutation at A–T basepairs and for efficient class switch recombination. J. Exp. Med. 198, 1171–1178 (2003).
Bardwell, P. D. et al. Altered somatic hypermutation and reduced class-switch recombination in exonuclease 1-mutant mice. Nature Immunol. 5, 224–229 (2004).
Acknowledgements
We thank John Manis, Ming Tian, Ali Zarrin, Sheila Ranganath, Craig Bassing and Chan Khuong and other members of the laboratory for careful evaluation of this manuscript. Work on CSR in the Alt laboratory is supported in part by the National Institutes of Health. F.W.A. is an Investigator of the Howard Hughes Medical Institute.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Glossary
- NON-HOMOLOGOUS END JOINING
-
(NHEJ). The process that joins broken DNA ends without depending on extended homology. Components of this pathway include the DNA-end-binding proteins Ku70 and Ku80, the endonuclease ARTEMIS, X-ray repair cross-complementing protein 4 (XRCC4), DNA ligase IV and the catalytic subunit of DNA-dependent protein kinase (DNA-PKcs).
- CYTOKINES
-
Biologically active molecules that are released by cells and affect the function of other cells.
- RGYW MOTIFS
-
The nucleotide sequence RGYW (where R denotes A or G, Y denotes C or T, and W denotes A or T) is considered a 'hot spot' for somatic hypermutation, as it is considerably more mutable than other random sequences.
- LIGATION-MEDIATED PCR
-
(lm-pcr). This method, which involves the ligation of unphosphorylated adaptors to broken DNA followed by PCR, can detect DNA double-stranded breaks in defined regions of the genome. It has been used extensively to analyse recombination-activating gene (RAG)-dependent breaks during V(D)J recombination, and more recently, in variable (V) gene segments and switch (S) regions during somatic hypermutation and class-switch recombination, respectively.
- SYNAPSIS
-
Non-covalent juxtaposition of two non-adjacent stretches of DNA.
Rights and permissions
About this article
Cite this article
Chaudhuri, J., Alt, F. Class-switch recombination: interplay of transcription, DNA deamination and DNA repair. Nat Rev Immunol 4, 541–552 (2004). https://doi.org/10.1038/nri1395
Issue Date:
DOI: https://doi.org/10.1038/nri1395
This article is cited by
-
Risk of carcinogenesis in the biliary epithelium of children with congenital biliary dilatation through epigenetic and genetic regulation
Surgery Today (2022)
-
Spt5-mediated enhancer transcription directly couples enhancer activation with physical promoter interaction
Nature Genetics (2020)
-
More than one antibody of individual B cells revealed by single-cell immune profiling
Cell Discovery (2019)
-
Uracil-DNA glycosylase is not implicated in the choice of the DNA repair pathway during B-cell class switch recombination
Cellular & Molecular Immunology (2019)
-
Essential role of the initial activation signal in isotype selection upon deletion of a transcriptionally committed promoter
Scientific Reports (2019)