Abstract
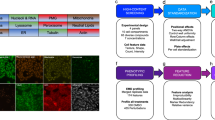
Quantitative analytical approaches for discovering new compound mechanisms are required for summarizing high-throughput, image-based drug screening data. Here we present a multivariate method for classifying untreated and treated human cancer cells based on ∼300 single-cell phenotypic measurements. This classification provides a score, measuring the magnitude of the drug effect, and a vector, indicating the simultaneous phenotypic changes induced by the drug. These two quantities were used to characterize compound activities and identify dose-dependent multiphasic responses. A systematic survey of profiles extracted from a 100-compound compendium of image data revealed that only 10–15% of the original features were required to detect a compound effect. We report the most informative image features for each compound and fluorescence marker set using a method that will be useful for determining minimal collections of readouts for drug screens. Our approach provides human-interpretable profiles and automatic determination of on- and off-target effects.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
References
Boland, M.V. & Murphy, R.F. After sequencing: quantitative analysis of protein localization. IEEE Eng. Med. Biol. Mag. 18, 115–119 (1999).
Lang, P., Yeow, K., Nichols, A. & Scheer, A. Cellular imaging in drug discovery. Nat. Rev. Drug Discov. 5, 343–356 (2006).
Price, J.H. et al. Advances in molecular labeling, high throughput imaging and machine intelligence portend powerful functional cellular biochemistry tools. J. Cell. Biochem. Suppl. 39, 194–210 (2002).
Zhou, X. & Wong, S.T.C. Informatics challenges of high-throughput microscopy. IEEE Signal Process. Mag. 23, 63–72 (2006).
Perlman, Z.E. et al. Multidimensional drug profiling by automated microscopy. Science 306, 1194–1198 (2004).
Tanaka, M. et al. An unbiased cell morphology-based screen for new, biologically active small molecules. PLoS Biol. 3, e128 (2005).
Boland, M.V. & Murphy, R.F. A neural network classifier capable of recognizing the patterns of all major subcellular structures in fluorescence microscope images of HeLa cells. Bioinformatics 17, 1213–1223 (2001).
Conrad, C. et al. Automatic identification of subcellular phenotypes on human cell arrays. Genome Res. 14, 1130–1136 (2004).
Neumann, B. et al. High-throughput RNAi screening by time-lapse imaging of live human cells. Nat. Methods 3, 385–390 (2006).
Ohya, Y. et al. High-dimensional and large-scale phenotyping of yeast mutants. Proc. Natl. Acad. Sci. USA 102, 19015–19020 (2005).
Gasparri, F., Mariani, M., Sola, F. & Galvani, A. Quantification of the proliferation index of human dermal fibroblast cultures with the ArrayScan high-content screening reader. J. Biomol. Screen. 9, 232–243 (2004).
Giuliano, K.A. et al. Systems cell biology knowledge created from high content screening. Assay Drug Dev. Technol. 3, 501–514 (2005).
Vapnik, V.N. Statistical Learning Theory. (John Wiley & Sons, New York, 1998).
Duda, R.O., Hart, P.E. & Stork, D.G. Pattern Classification 2nd edn. (John Wiley & Sons, New York, 2001).
Guyon, I., Weston, J., Barnhill, S. & Vapnik, V. Gene selection for cancer classification using support vector machines. Mach. Learn. 46, 389–422 (2002).
Haralick, R.M. Statistical and structural approaches to texture. Proc. IEEE 67, 786–804 (1979).
Teh, C.H. & Chin, R.T. On image analysis by the methods of moments. IEEE Trans. Pattern Anal. Mach. Intell. 10, 496–513 (1988).
Lundholt, B.K., Scudder, K.M. & Pagliaro, L. A simple technique for reducing edge effect in cell-based Assays. J. Biomol. Screen. 8, 566–570 (2003).
Malo, N., Hanley, J.A., Cerquozzi, S., Pelletier, J. & Nadon, R. Statistical practice in high-throughput screening data analysis. Nat. Biotechnol. 24, 167–175 (2006).
Hastie, T., Tibshirani, R. & Friedman, J. The Elements of Statistical Learning: Data Mining, Inference, and Prediction. (Springer-Verlag, New York, 2001).
Yeung, T.K., Germond, C., Chen, X. & Wang, Z. The mode of action of taxol: apoptosis at low concentration and necrosis at high concentration. Biochem. Biophys. Res. Commun. 263, 398–404 (1999).
Yoo, C.B. & Jones, P.A. Epigenetic therapy of cancer: past, present and future. Nat. Rev. Drug Discov. 5, 37–50 (2006).
Perlman, Z.E., Mitchison, T.J. & Mayer, T.U. High-content screening and profiling of drug activity in an automated centrosome-duplication assay. ChemBioChem 6, 145–151 (2005).
Bollag, D.M. et al. Epothilones, a new class of microtubule-stabilizing agents with a taxol-like mechanism of action. Cancer Res. 55, 2325–2333 (1995).
Panda, D., Rathinasamy, K., Santra, M.K. & Wilson, L. Kinetic suppression of microtubule dynamic instability by griseofulvin: implications for its possible use in the treatment of cancer. Proc. Natl. Acad. Sci. USA 102, 9878–9883 (2005).
Clemons, P.A. Complex phenotypic assays in high-throughput screening. Curr. Opin. Chem. Biol. 8, 334–338 (2004).
Dove, A. Drug screening-beyond the bottleneck. Nat. Biotechnol. 17, 859–863 (1999).
Maciag, K. et al. Systems-level analyses identify extensive coupling among gene expression machines. Mol. Syst. Biol. [online] 2, 0003 (2006) (doi:10.1038/msb4100045).
Hanley, J.A. & McNeil, B.J. The meaning and use of the area under a receiver operating characteristic (ROC) curve. Radiology 143, 29–36 (1982).
Acknowledgements
We thank S. Johnson from the Texas Advanced Computing Center for high-performance computing technical assistance, and all members of the Altschuler and Wu lab at the UT Southwestern Medical Center for stimulating discussions. This research was supported by the Endowed Scholars program at UT Southwestern Medical Center and the Welch Foundation (I-1619, I-1644).
Author information
Authors and Affiliations
Contributions
L.-H.L. designed, implemented and performed the profiling methods. All authors contributed to the conception of the overall approach, statistical analysis of the methods and writing of the manuscript.
Note: Supplementary information is available on the Nature Methods website.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Fig. 1
Distributions of the number of selected features. (PDF 67 kb)
Supplementary Fig. 2
Common phenotypic changes for the DNA-SC35-anillin, DNA-p53-cFos, and DNA-MT-actin marker sets. (PDF 1442 kb)
Supplementary Fig. 3
Some of the common phenotypic changes are cell-cycle independent. (PDF 161 kb)
Rights and permissions
About this article
Cite this article
Loo, LH., Wu, L. & Altschuler, S. Image-based multivariate profiling of drug responses from single cells. Nat Methods 4, 445–453 (2007). https://doi.org/10.1038/nmeth1032
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nmeth1032
This article is cited by
-
Single object profiles regression analysis (SOPRA): a novel method for analyzing high-content cell-based screens
BMC Bioinformatics (2022)
-
Self-supervised deep learning encodes high-resolution features of protein subcellular localization
Nature Methods (2022)
-
A statistical framework for high-content phenotypic profiling using cellular feature distributions
Communications Biology (2022)
-
Image-based profiling for drug discovery: due for a machine-learning upgrade?
Nature Reviews Drug Discovery (2021)
-
Organoids in image-based phenotypic chemical screens
Experimental & Molecular Medicine (2021)