Abstract
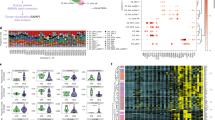
Autoimmune diseases are common and debilitating, but their severe manifestations could be reduced if biomarkers were available to allow individual tailoring of potentially toxic immunosuppressive therapy. Gene expression–based biomarkers facilitating such tailoring of chemotherapy in cancer, but not autoimmunity, have been identified and translated into clinical practice1,2. We show that transcriptional profiling of purified CD8+ T cells, which avoids the confounding influences of unseparated cells3,4, identifies two distinct subject subgroups predicting long-term prognosis in two autoimmune diseases, antineutrophil cytoplasmic antibody (ANCA)-associated vasculitis (AAV), a chronic, severe disease characterized by inflammation of medium-sized and small blood vessels5, and systemic lupus erythematosus (SLE), characterized by autoantibodies, immune complex deposition and diverse clinical manifestations ranging from glomerulonephritis to neurological dysfunction6. We show that the subset of genes defining the poor prognostic group is enriched for genes involved in the interleukin-7 receptor (IL-7R) pathway and T cell receptor (TCR) signaling and those expressed by memory T cells. Furthermore, the poor prognostic group is associated with an expanded CD8+ T cell memory population. These subgroups, which are also found in the normal population and can be identified by measuring expression of only three genes, raise the prospect of individualized therapy and suggest new potential therapeutic targets in autoimmunity.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Accession codes
References
Golub, T.R. et al. Molecular classification of cancer: class discovery and class prediction by gene expression monitoring. Science 286, 531–537 (1999).
van't Veer, L.J. & Bernards, R. Enabling personalized cancer medicine through analysis of gene-expression patterns. Nature 452, 564–570 (2008).
Bryant, P.A., Smyth, G.K., Robins-Browne, R. & Curtis, N. Detection of gene expression in an individual cell type within a cell mixture using microarray analysis. PLoS One 4, e4427 (2009).
Lyons, P.A. et al. Novel expression signatures identified by transcriptional analysis of separated leukocyte subsets in SLE and vasculitis. Ann. Rheum. Dis. published online, doi:10.1136/ard.2009.108043 (2009).
Lane, S.E., Watts, R.A., Shepstone, L. & Scott, D.G. Primary systemic vasculitis: clinical features and mortality. QJM 98, 97–111 (2005).
Tan, E.M. et al. The 1982 revised criteria for the classification of systemic lupus erythematosus. Arthritis Rheum. 25, 1271–1277 (1982).
Booth, A.D. et al. Outcome of ANCA-associated renal vasculitis: a 5-year retrospective study. Am. J. Kidney Dis. 41, 776–784 (2003).
Jennette, J.C., Xiao, H. & Falk, R.J. Pathogenesis of vascular inflammation by anti-neutrophil cytoplasmic antibodies. J. Am. Soc. Nephrol. 17, 1235–1242 (2006).
Langford, C.A. Antineutrophil cytoplasmic antibodies should not be used to guide treatment in Wegener′s granulomatosis. Clin. Exp. Rheumatol. 22, S3–S6 (2004).
Lyons, P.A. et al. Microarray analysis of human leucocyte subsets: the advantages of positive selection and rapid purification. BMC Genomics 8, 64 (2007).
Le Brigand, K. et al. An open-access long oligonucleotide microarray resource for analysis of the human and mouse transcriptomes. Nucleic Acids Res. 34, e87 (2006).
Abdulahad, W.H., Stegeman, C.A., Limburg, P.C. & Kallenberg, C.G. CD4-positive effector memory T cells participate in disease expression in ANCA-associated vasculitis. Ann. NY Acad. Sci. 1107, 22–31 (2007).
Lamprecht, P. Off balance: T-cells in antineutrophil cytoplasmic antibody (ANCA)-associated vasculitides. Clin. Exp. Immunol. 141, 201–210 (2005).
Torrente, A., Kapushesky, M. & Brazma, A. A new algorithm for comparing and visualizing relationships between hierarchical and flat gene expression data clusterings. Bioinformatics 21, 3993–3999 (2005).
Monti, S. Consensus Clustering: A resampling-based method for class discovery and visualisation of gene expression microarray data. Mach. Learn. 52, 91–118 (2003).
Blanco, P. et al. Increase in activated CD8+ T lymphocytes expressing perforin and granzyme B correlates with disease activity in patients with systemic lupus erythematosus. Arthritis Rheum. 52, 201–211 (2005).
Ippolito, A. & Petri, M. An update on mortality in systemic lupus erythematosus. Clin. Exp. Rheumatol. 26, S72–S79 (2008).
Illei, G.G. & Lipsky, P.E. Biomarkers in systemic lupus erythematosus. Curr. Rheumatol. Rep. 6, 382–390 (2004).
Subramanian, A. et al. Gene set enrichment analysis: a knowledge-based approach for interpreting genome-wide expression profiles. Proc. Natl. Acad. Sci. USA 102, 15545–15550 (2005).
Buentke, E.M.A., Tolaini, M., Di Santo, J., Zamoyska, R. & Seddon, B. Do CD8 effector cells need IL7-R expression to become resting memory cells? Blood 108, 1949–1956 (2006).
Pellegrini, M., Belz, G., Bouillet, P. & Strasser, A. Shutdown of an acute T cell immune response to viral infection is mediated by the proapoptotic Bcl-2 homology 3-only protein Bim. Proc. Natl. Acad. Sci. USA 100, 14175–14180 (2003).
Schluns, K.S. & Lefrancois, L. Cytokine control of memory T-cell development and survival. Nat. Rev. Immunol. 3, 269–279 (2003).
Joshi, N.S. & Kaech, S.M. Effector CD8 T cell development: a balancing act between memory cell potential and terminal differentiation. J. Immunol. 180, 1309–1315 (2008).
Willinger, T., Freeman, T., Hasegawa, H., McMichael, A.J. & Callan, M.F. Molecular signatures distinguish human central memory from effector memory CD8 T cell subsets. J. Immunol. 175, 5895–5903 (2005).
Mao, M. et al. T lymphocyte activation gene identification by coregulated expression on DNA microarrays. Genomics 83, 989–999 (2004).
Galon, J. et al. Gene profiling reveals unknown enhancing and suppressive actions of glucocorticoids on immune cells. FASEB J. 16, 61–71 (2002).
Sallusto, F., Lenig, D., Forster, R., Lipp, M. & Lanzavecchia, A. Two subsets of memory T lymphocytes with distinct homing potentials and effector functions. Nature 401, 708–712 (1999).
Morley, M. et al. Genetic analysis of genome-wide variation in human gene expression. Nature 430, 743–747 (2004).
Vezys, V. et al. Memory CD8 T-cell compartment grows in size with immunological experience. Nature 457, 196–199 (2009).
Bevan, M.J. Helping the CD8+ T-cell response. Nat. Rev. Immunol. 4, 595–602 (2004).
Williams, M.A. & Bevan, M.J. Effector and memory CTL differentiation. Annu. Rev. Immunol. 25, 171–192 (2007).
Hou, S., Hyland, L., Ryan, K.W., Portner, A. & Doherty, P.C. Virus-specific CD8+ T-cell memory determined by clonal burst size. Nature 369, 652–654 (1994).
Alizadeh, A.A. et al. Distinct types of diffuse large B-cell lymphoma identified by gene expression profiling. Nature 403, 503–511 (2000).
van 't Veer, L.J. et al. Gene expression profiling predicts clinical outcome of breast cancer. Nature 415, 530–536 (2002).
van de Vijver, M.J. et al. A gene-expression signature as a predictor of survival in breast cancer. N. Engl. J. Med. 347, 1999–2009 (2002).
Stone, J.H. et al. A disease-specific activity index for Wegener's granulomatosis: modification of the Birmingham Vasculitis Activity Score. International Network for the Study of the Systemic Vasculitides (INSSYS). Arthritis Rheum. 44, 912–920 (2001).
Willcocks, L.C. et al. Copy number of FCGR3B, which is associated with systemic lupus erythematosus, correlates with protein expression and immune complex uptake. J. Exp. Med. 205, 1573–1582 (2008).
Acknowledgements
We thank D. Fearon and J. Todd for critical review of the manuscript, T. Rayner, M. Kapushesky and M. Clatworthy for helpful discussions, K. Townsend for help with subject recruitment, H. Woffendin and T. Freeman for help with optimizing the custom array platform and A. Hatton and H. Ratlamwala for technical support, along with all subjects and clinicians involved in enrollment and clinical follow up. We are grateful to E. Clutterbuck, R. Lazarus and the volunteers at the Oxford Vaccine Group for their help with recruitment of healthy controls. This work was supported by the UK National Institute of Health Research, the Cambridge Biomedical Research Centre, the Wellcome Trust (Programme Grant number 083650/Z/07/Z), the UK Medical Research Council, the Evelyn Trust, Kidney Research UK and the National Medical Research Council of Singapore (grant reference IRG07nov089). E.F.M. holds a Wellcome Fellowship, and L.C.W. a Medical Research Council, Clinical Training Fellowship. K.G.C.S. is a Lister Prize Fellow and Khoo Oon Teik Professor of Nephrology at the National University of Singapore. The Cambridge Institute for Medical Research is in receipt of a Wellcome Trust Strategic Award (079895).
Author information
Authors and Affiliations
Contributions
K.G.C.S. and P.A.L. designed the study and wrote the paper along with E.F.M. E.F.M. analyzed the data with help from A.B., carried out experiments and collected and analyzed clinical data along with J.L.H., M.K., D.R.W.J., L.C.W. and A.N.C. E.J.C. contributed to experiments in healthy control subjects along with A.J.P. and performed validation of microarray results. Singaporean data was collected and analyzed by V.J., D.M.K. and P.A.M. along with E.F.M., P.A.L. and K.G.C.S.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–13, Supplementary Tables 1–7, Supplementary Methods and Supplementary Discussion (PDF 9203 kb)
Rights and permissions
About this article
Cite this article
McKinney, E., Lyons, P., Carr, E. et al. A CD8+ T cell transcription signature predicts prognosis in autoimmune disease. Nat Med 16, 586–591 (2010). https://doi.org/10.1038/nm.2130
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nm.2130
This article is cited by
-
Single-cell multi-omics analysis identifies two distinct phenotypes of newly-onset microscopic polyangiitis
Nature Communications (2023)
-
The promise of precision medicine in rheumatology
Nature Medicine (2022)
-
Transcriptional analysis of peripheral memory T cells reveals Parkinson’s disease-specific gene signatures
npj Parkinson's Disease (2022)
-
Regulation of activated T cell survival in rheumatic autoimmune diseases
Nature Reviews Rheumatology (2022)
-
Characterization of interstitial infiltrates in MPO and PR3 anti-neutrophil cytoplasmic antibody glomerulonephritis
Journal of Nephrology (2022)