Abstract
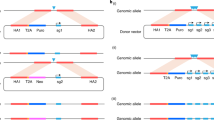
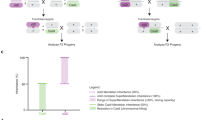
FLP/FRT-induced mitotic recombination provides a powerful method for creating genetic mosaics in Drosophila and for discerning the function of recessive genes in a heterozygous individual. Here we show that mitotic recombination can be reproducibly induced in mouse embryonic stem (ES) cells, by Cre/loxP technology, at frequencies ranging from 4.2 × 10−5 (Snrpn) to 7.0 × 10−3 (D7Mit178) for single allelic loxP sites, and to 5.0 × 10−2 (D7Mit178) for multiple allelic lox sites, after transient Cre expression. Notably, much of the recombination occurs in G2 and is followed by X segregation, where the recombinant chromatids segregate away from each other during mitosis. It is X segregation that is useful for genetic mosaic analysis because it produces clones of homozygous mutant daughter cells from heterozygous mothers. Our studies confirm the predictions made from studies in Drosophila1 that suggest that X segregation will not be limited to organisms with strong mitotic pairing, because the forces (sister-chromatid cohesion) responsible for X segregation are an elemental feature of mitosis in all eukaryotes. Our studies also show that genetic mosaic analysis in mice is feasible, at least for certain chromosomal regions.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on SpringerLink
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
References
Beumer, K.J., Pimpinelli, S. & Golic, K.G. Induced chromosomal exchange directs the segregation of recombinant chromatids in mitosis of Drosophila. Genetics 150, 173–188 (1998).
Rossant, J. & Spence, A. Chimeras and mosaics in mouse mutant analysis. Trends Genet. 14, 358–363 (1998).
Favor, J. & Neuhauser-Klaus, A. Genetic mosaicism in the house mouse. Annu. Rev. Genet. 28, 27–47 (1994).
Xu, T. & Rubin, G.M. Analysis of genetic mosaics in developing and adult Drosophila tissues. Development 117, 1223–1237 (1993).
Pimpinelli, S. & Ripoll, P. Nonrandom segregation of centromeres following mitotic recombination in Drosophila melanogaster. Proc. Natl Acad. Sci. USA 83, 3900–3903 (1986).
Stevens, N.M. A study of the germ cells of certain Diptera, with reference to the heterochromosomes and the phenomena of synapsis. J. Exp. Zool. 5, 359–383 (1908).
Metz, C.W. Chromosome studies on the Diptera II: the paired association of chromosomes in the Diptera, and its significance. J. Exp. Zool. 21, 213–279 (1916).
Golic, K.G. Site-specific recombination between homologous chromosomes in Drosophila. Science 252, 958–961 (1991).
Xu, T., Wang, W., Zhang, S., Stewart, R.A. & Yu, W. Identifying tumor suppressors in genetic mosaics: the Drosophila lats gene encodes a putative protein kinase. Development 121, 1053–1063 (1995).
Cavenee, W.K. et al. Expression of recessive alleles by chromosomal mechanisms in retinoblastoma. Nature 305, 779–784 (1983).
Mortensen, R.M., Conner, D.A., Chao, S., Geisterfer-Lowrance, A. & Seidman, J.G. Production of homozygous mutant ES cells with a single targeting construct. Mol. Cell. Biol. 12, 2391–2395 (1992).
Shao, C. et al. Mitotic recombination produces the majority of recessive fibroblast variants in heterozygous mice. Proc. Natl Acad. Sci. USA 96, 9230–9235 (1999).
Bateman, A.J. A probable case of mitotic crossing-over in the mouse. Genet. Res. 9, 375 (1967).
Panthier, J.J., Guenet, J.L., Condamine, H. & Jacob, F. Evidence for mitotic recombination in Wei/+ heterozygous mice. Genetics 125, 175–182 (1990).
Fisher, G., Stephenson, D.A. & West, J.D. Investigation of the potential for mitotic recombination in the mouse. Mutat. Res. 164, 381–388 (1986).
German, J. Cytological evidence for crossing-over in vitro in human lymphoid cells. Science 144, 298–301 (1964).
Ellis, N.A. et al. The Bloom's syndrome gene product is homologous to RecQ helicases. Cell 83, 655–666 (1995).
Johnson, R.D. & Jasin, M. Double-strand-break-induced homologous recombination in mammalian cells. Biochem. Soc. Trans. 29, 196–201 (2001).
Herault, Y., Rassoulzadegan, M., Cuzin, F. & Duboule, D. Engineering chromosomes in mice through targeted meiotic recombination (TAMERE). Nature Genet. 20, 381–384 (1998).
Zheng, B., Sage, M., Sheppeard, E.A., Jurecic, V. & Bradley, A. Engineering mouse chromosomes with Cre-loxP: range, efficiency, and somatic applications. Mol. Cell. Biol. 20, 648–655 (2000).
Ramirez-Solis, R., Liu, P. & Bradley, A. Chromosome engineering in mice. Nature 378, 720–724 (1995).
Matzuk, M.M., Finegold, M.J., Su, J.G., Hsueh, A.J. & Bradley, A. α-inhibin is a tumour-suppressor gene with gonadal specificity in mice. Nature 360, 313–319 (1992).
Yang, T. et al. A mouse model for Prader-Willi syndrome imprinting-centre mutations. Nature Genet. 19, 25–31 (1998).
Gabriel, J.M. et al. A transgene insertion creating a heritable chromosome deletion mouse model of Prader-Willi and angelman syndromes. Proc. Natl Acad. Sci. USA 96, 9258–9263 (1999).
Baubonis, W. & Sauer, B. Genomic targeting with purified Cre recombinase. Nucleic Acids Res. 21, 2025–2029 (1993).
Riesselmann, L. & Haaf, T. Preferential S-phase pairing of the imprinted region on distal mouse chromosome 7. Cytogenet. Cell Genet. 86, 39–42 (1999).
LaSalle, J. M. & Lalande, M. Homologous association of oppositely imprinted chromosomal domains. Science 272, 725–728 (1996).
Araki, K., Araki, M. & Yamamura, K. Targeted integration of DNA using mutant lox sites in embryonic stem cells. Nucleic Acids Res. 25, 868–872 (1997).
Luo, G. et al. Cancer predisposition caused by elevated mitotic recombination in bloom mice. Nature Genet. 26, 424–429 (2000).
Cattanach, B.M. & Jones, J. Genetic imprinting in the mouse: implications for gene regulation. J. Inherit. Metab. Dis. 17, 403–420 (1994).
Lee, G. & Saito, I. Role of nucleotide sequences of loxP spacer region in Cre-mediated recombination. Gene 216, 55–65 (1998).
Acknowledgements
We thank A. Bradley for providing AB2.2 ES cells, STO feeder cells, an HPRT1 minigene, a D11Mit71 genomic fragment and PolII-neor and puro selection cassettes, and C. Brannan for the Snrpn promoter probe. We also thank W.F. Dove, who provided the inspiration for these experiments. This research was supported by the National Cancer Institute, Department of Health and Human Services.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Liu, P., Jenkins, N. & Copeland, N. Efficient Cre-loxP–induced mitotic recombination in mouse embryonic stem cells. Nat Genet 30, 66–72 (2002). https://doi.org/10.1038/ng788
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ng788
This article is cited by
-
Genome engineering of Agrobacterium tumefaciens using the lambda Red recombination system
Applied Microbiology and Biotechnology (2014)
-
Dissection of gene function at clonal level using mosaic analysis with double markers
Frontiers in Biology (2013)
-
A CO-FISH assay to assess sister chromatid segregation patterns in mitosis of mouse embryonic stem cells
Chromosome Research (2013)
-
Biased segregation of DNA and centrosomes — moving together or drifting apart?
Nature Reviews Molecular Cell Biology (2009)
-
A Novel Genotyping Strategy Based on Allele-specific Inverse PCR for Rapid and Reliable Identification of Conditional FADD Knockout Mice
Molecular Biotechnology (2008)