Abstract
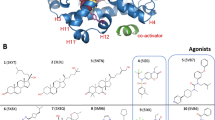
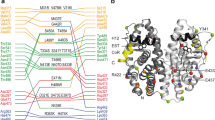
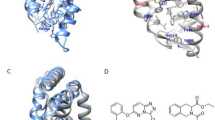
Our understanding of how steroid hormones regulate physiological functions has been significantly advanced by structural biology approaches. However, progress has been hampered by misfolding of the ligand binding domains in heterologous expression systems and by conformational flexibility that interferes with crystallization. Here, we show that protein folding problems that are common to steroid hormone receptors are circumvented by mutations that stabilize well-characterized conformations of the receptor. We use this approach to present the structure of an apo steroid receptor that reveals a ligand-accessible channel allowing soaking of preformed crystals. Furthermore, crystallization of different pharmacological classes of compounds allowed us to define the structural basis of NFκB-selective signaling through the estrogen receptor, thus revealing a unique conformation of the receptor that allows selective suppression of inflammatory gene expression. The ability to crystallize many receptor-ligand complexes with distinct pharmacophores allows one to define structural features of signaling specificity that would not be apparent in a single structure.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout







Similar content being viewed by others
Change history
01 May 2008
In the version of this article initially published, the 13th author's last name is misspelled. The author's name should read 'Andrzej Joachimiak'. Additionally, Figure 5 of this article inadvertently contains pink traces in each panel that are not attributed to any specific molecule. These errors have been corrected in the HTML and PDF versions of the article.
References
Schulman, I.G. & Heyman, R.A. The flip side: identifying small molecule regulators of nuclear receptors. Chem. Biol. 11, 639–646 (2004).
Chadwick, C.C. et al. Identification of pathway-selective estrogen receptor ligands that inhibit NF-kappaB transcriptional activity. Proc. Natl. Acad. Sci. USA 102, 2543–2548 (2005).
Steffan, R.J. et al. Synthesis and activity of substituted 4-(indazol-3-yl)phenols as pathway-selective estrogen receptor ligands useful in the treatment of rheumatoid arthritis. J. Med. Chem. 47, 6435–6438 (2004).
Darimont, B.D. et al. Structure and specificity of nuclear receptor-coactivator interactions. Genes Dev. 12, 3343–3356 (1998).
Shiau, A.K. et al. The structural basis of estrogen receptor/coactivator recognition and the antagonism of this interaction by tamoxifen. Cell 95, 927–937 (1998).
De Bosscher, K. et al. A fully dissociated compound of plant origin for inflammatory gene repression. Proc. Natl. Acad. Sci. USA 102, 15827–15832 (2005).
Weis, K.E., Ekena, K., Thomas, J.A., Lazennec, G. & Katzenellenbogen, B.S. Constitutively active human estrogen receptors containing amino acid substitutions for tyrosine 537 in the receptor protein. Mol. Endocrinol. 10, 1388–1398 (1996).
Zhong, L. & Skafar, D.F. Mutations of tyrosine 537 in the human estrogen receptor-alpha selectively alter the receptor's affinity for estradiol and the kinetics of the interaction. Biochemistry 41, 4209–4217 (2002).
Carlson, K.E., Choi, I., Gee, A., Katzenellenbogen, B.S. & Katzenellenbogen, J.A. Altered ligand binding properties and enhanced stability of a constitutively active estrogen receptor: evidence that an open pocket conformation is required for ligand interaction. Biochemistry 36, 14897–14905 (1997).
Tremblay, G.B., Tremblay, A., Labrie, F. & Giguere, V. Ligand-independent activation of the estrogen receptors alpha and beta by mutations of a conserved tyrosine can be abolished by antiestrogens. Cancer Res. 58, 877–881 (1998).
Yudt, M.R. et al. Function of estrogen receptor tyrosine 537 in hormone binding, DNA binding, and transactivation. Biochemistry 38, 14146–14156 (1999).
Lazennec, G., Ediger, T.R., Petz, L.N., Nardulli, A.M. & Katzenellenbogen, B.S. Mechanistic aspects of estrogen receptor activation probed with constitutively active estrogen receptors: correlations with DNA and coregulator interactions and receptor conformational changes. Mol. Endocrinol. 11, 1375–1386 (1997).
Hsieh, R.W. et al. Identification of ligands with bicyclic scaffolds provides insights into mechanisms of estrogen receptor subtype selectivity. J. Biol. Chem. 281, 17909–17919 (2006).
Nettles, K.W. et al. Structural plasticity in the oestrogen receptor ligand-binding domain. EMBO Rep. 8, 563–568 (2007).
Compton, D.R. et al. Pyrazolo[1,5-a]pyrimidines: estrogen receptor ligands possessing estrogen receptor beta antagonist activity. J. Med. Chem. 47, 5872–5893 (2004).
De Angelis, M., Stossi, F., Carlson, K.A., Katzenellenbogen, B.S. & Katzenellenbogen, J.A. Indazole estrogens: highly selective ligands for the estrogen receptor beta. J. Med. Chem. 48, 1132–1144 (2005).
De Angelis, M., Stossi, F., Waibel, M., Katzenellenbogen, B.S. & Katzenellenbogen, J.A. Isocoumarins as estrogen receptor beta selective ligands: Isomers of isoflavone phytoestrogens and their metabolites. Bioorg. Med. Chem. 13, 6529–6542 (2005).
Kim, S.H. & Katzenellenbogen, J.A. Hormone-PAMAM dendrimer conjugates: polymer dynamics and tether structure affect ligand access to receptors. Angew. Chem. Int. Edn Engl. 45, 7243–7248 (2006).
Muthyala, R.S., Sheng, S., Carlson, K.E., Katzenellenbogen, B.S. & Katzenellenbogen, J.A. Bridged bicyclic cores containing a 1,1-diarylethylene motif are high-affinity subtype-selective ligands for the estrogen receptor. J. Med. Chem. 46, 1589–1602 (2003).
Zhou, H.B., Comninos, J.S., Stossi, F., Katzenellenbogen, B.S. & Katzenellenbogen, J.A. Synthesis and evaluation of estrogen receptor ligands with bridged oxabicyclic cores containing a diarylethylene motif: estrogen antagonists of unusual structure. J. Med. Chem. 48, 7261–7274 (2005).
Zhou, H.B. et al. Elemental isomerism: a boron-nitrogen surrogate for a carbon-carbon double bond increases the chemical diversity of estrogen receptor ligands. Chem. Biol. 14, 659–669 (2007).
Zhou, H.B. et al. Structure-guided optimization of estrogen receptor binding affinity and antagonist potency of pyrazolopyrimidines with basic side chains. J. Med. Chem. 50, 399–403 (2007).
Zhang, J.X., Labaree, D.C. & Hochberg, R.B. Nonpolar and short side chain groups at C-11beta of estradiol result in antiestrogens. J. Med. Chem. 48, 1428–1447 (2005).
Bennion, B.J. et al. PhIP carcinogenicity in breast cancer: computational and experimental evidence for competitive interactions with human estrogen receptor. Chem. Res. Toxicol. 18, 1528–1536 (2005).
Nissen, R.M. & Yamamoto, K.R. The glucocorticoid receptor inhibits NFkappaB by interfering with serine-2 phosphorylation of the RNA polymerase II carboxy-terminal domain. Genes Dev. 14, 2314–2329 (2000).
Nettles, K.W. et al. CBP is a dosage dependent regulator of NFκB suppression by the estrogen receptor. Mol. Endocrinol. 22, 263–272 (2008).
Pike, A.C. et al. Structural insights into the mode of action of a pure antiestrogen. Structure 9, 145–153 (2001).
Ekena, K., Weis, K.E., Katzenellenbogen, J.A. & Katzenellenbogen, B.S. Identification of amino acids in the hormone binding domain of the human estrogen receptor important in estrogen binding. J. Biol. Chem. 271, 20053–20059 (1996).
Bruning, J.B. et al. Partial agonists activate PPARgamma using a helix 12 independent mechanism. Structure 15, 1258–1271 (2007).
Johnson, B.A. et al. Ligand-induced stabilization of PPARgamma monitored by NMR spectroscopy: implications for nuclear receptor activation. J. Mol. Biol. 298, 187–194 (2000).
Stols, L. et al. A new vector for high-throughput, ligation-independent cloning encoding a tobacco etch virus protease cleavage site. Protein Expr. Purif. 25, 8–15 (2002).
Otwinowski, Z. & Minor, W. Processing of X-ray diffraction data collected in oscillation mode. Methods Enzymol. 276, 307–326 (1997).
Collaborative Computational Project, Number 4. The CCP4 suite: programs for protein crystallography. Acta Crystallogr. D Biol. Crystallogr. 50, 760–763 (1994).
Acknowledgements
We are very grateful to S. Rajan for her work on refinement and model building of the structures. We thank T. Tellinghiusen and J. Cleveland for comments on the manuscript. The authors also thank J. Chrzas, G. Sahle and J. Habel for data collection at APS SER-CAT, and C. Smith, G. Card and J. Habel for data collection at SSRL beamlines. Portions of data were collected at Southeast Regional Collaborative Access Team (SER-CAT) 22-ID (or 22-BM) beamline at the Advanced Photon Source, Argonne National Laboratory. Portions of this research were carried out at the Stanford Synchrotron Radiation Laboratory, a national user facility operated by Stanford University on behalf of the US Department of Energy, Office of Basic Energy Sciences. This work was supported by the US National Institutes of Health (1R21 NS056998-01 (K.W.N.); 5R01 CA89489 (G.L.G.); 5R37 DK15556 (J.A.K.); 5R01 CA18119 (B.S.K.); R01 HL61432 and R01 CA37799 (R.B.H.)), the Ludwig Fund for Cancer Research (G.L.G.) and US Department of Defense grant W81XWH-04-1-0791 (G.L.G.).
Author information
Authors and Affiliations
Contributions
K.W.N., J.B.B., J.N., S.K.S. and J.B.H. worked on crystallization. K.W.N., J.B.B. and Y.K. worked on X-ray data collection and data analysis. K.K., R.B.H., H.Z., J.A.K. and B.S.K. worked on generating reagents and chemical synthesis. G.G. performed the mammalian cell–based experiments. K.W.N., J.B.B., J.A.K., A.J. and G.L.G. designed and supervised experiments and wrote the paper.
Corresponding authors
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–5, Supplementary Tables 1 and 2, and Supplementary Methods (PDF 3269 kb)
Rights and permissions
About this article
Cite this article
Nettles, K., Bruning, J., Gil, G. et al. NFκB selectivity of estrogen receptor ligands revealed by comparative crystallographic analyses. Nat Chem Biol 4, 241–247 (2008). https://doi.org/10.1038/nchembio.76
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nchembio.76
This article is cited by
-
A PROSS-designed extensively mutated estrogen receptor α variant displays enhanced thermal stability while retaining native allosteric regulation and structure
Scientific Reports (2021)
-
A molecular switch regulating transcriptional repression and activation of PPARγ
Nature Communications (2020)
-
Endocrine therapy-resistant breast cancer model cells are inhibited by soybean glyceollin I through Eleanor non-coding RNA
Scientific Reports (2018)
-
Unraveling the clinicopathological features driving the emergence of ESR1 mutations in metastatic breast cancer
npj Breast Cancer (2018)
-
Dynamic changes in binding interaction networks of sex steroids establish their non-classical effects
Scientific Reports (2017)