Abstract
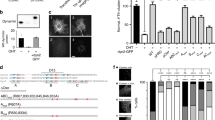
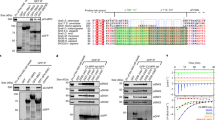
RME-1/EHD1 (receptor mediated endocytosis/Eps15 homology-domain containing 1) family proteins are key residents of the recycling endosome, which are required for endosome-to-plasma membrane transport in Caenorhabditis elegans and mammals. Recent studies suggest similarities between the RME-1/EHD proteins and the Dynamin GTPase superfamily of mechanochemical pinchases, which promote membrane fission. Here we show that endogenous C. elegans AMPH-1, the only C. elegans member of the Amphiphysin/BIN1 family of BAR (Bin1-Amphiphysin-Rvs161p/167p)-domain-containing proteins, colocalizes with RME-1 on recycling endosomes in vivo, that amph-1-deletion mutants are defective in recycling endosome morphology and function, and that binding of AMPH-1 Asn-Pro-Phe(Asp/Glu) sequences to the RME-1 EH-domain promotes the recycling of transmembrane cargo. We also show a requirement for human BIN1 (also known as Amphiphysin 2) in EHD1-regulated endocytic recycling. In vitro, we find that purified recombinant AMPH-1–RME-1 complexes produce short, coated membrane tubules that are qualitatively distinct from those produced by either protein alone. Our results indicate that AMPH-1 and RME-1 cooperatively regulate endocytic recycling, probably through functions required for the production of cargo carriers that exit the recycling endosome for the cell surface.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout








Similar content being viewed by others
References
Grant, B. et al. Evidence that RME-1, a conserved C. elegans EH-domain protein, functions in endocytic recycling. Nature Cell Biol. 3, 573–579 (2001).
Lin, S. X., Grant, B., Hirsh, D. & Maxfield, F. R. Rme-1 regulates the distribution and function of the endocytic recycling compartment in mammalian cells. Nature Cell Biol. 3, 567–572 (2001).
Caplan, S. et al. A tubular EHD1-containing compartment involved in the recycling of major histocompatibility complex class I molecules to the plasma membrane. EMBO J. 21, 2557–2567 (2002).
Lee, D. W. et al. ATP binding regulates oligomerization and endosome association of RME-1 family proteins. J. Biol. Chem. 280, 17213–17220 (2005).
Naslavsky, N., Rahajeng, J., Sharma, M., Jovic, M. & Caplan, S. Interactions between EHD proteins and Rab11-FIP2: a role for EHD3 in early endosomal transport. Mol. Biol. Cell 17, 163–177 (2006).
Sharma, M., Naslavsky, N. & Caplan, S. A role for EHD4 in the regulation of early endosomal transport. Traffic 9, 995–1018 (2008).
Shao, Y. et al. Pincher, a pinocytic chaperone for nerve growth factor/TrkA signaling endosomes. J. Cell Biol. 157, 679–691 (2002).
Guilherme, A. et al. EHD2 and the novel EH domain binding protein EHBP1 couple endocytosis to the actin cytoskeleton. J. Biol.Chem. 279, 10593–10605 (2004).
Daumke, O. et al. Architectural and mechanistic insights into an EHD ATPase involved in membrane remodelling. Nature 449, 923–927 (2007).
Sever, S. Dynamin and endocytosis. Curr. Opin. Cell Biol. 14, 463–467 (2002).
Roux, A. & Antonny, B. The long and short of membrane fission. Cell 135, 1163–1165 (2008).
Salcini, A. E. et al. Binding specificity and in vivo targets of the EH domain, a novel protein–protein interaction module. Genes Dev. 11, 2239–2249 (1997).
Grant, B. D. & Caplan, S. Mechanisms of EHD/RME-1 protein function in endocytic transport. Traffic (2008).
Kieken, F., Jovic, M., Naslavsky, N., Caplan, S. & Sorgen, P. L. EH domain of EHD1. J. Biomol. NMR 39, 323–329 (2007).
Takei, K., Slepnev, V. I., Haucke, V. & De Camilli, P. Functional partnership between amphiphysin and dynamin in clathrin-mediated endocytosis. Nature Cell Biol. 1, 33–39 (1999).
David, C., McPherson, P. S., Mundigl, O. & de Camilli, P. A role of amphiphysin in synaptic vesicle endocytosis suggested by its binding to dynamin in nerve terminals. Proc. Natl Acad. Sci. USA 93, 331–335 (1996).
Shi, A. et al. A novel requirement for C. elegans Alix/ALX-1 in RME-1-mediated membrane transport. Curr. Biol. 17, 1913–1924 (2007).
Sato, K. et al. Differential requirements for clathrin in receptor-mediated endocytosis and maintenance of synaptic vesicle pools. Proc. Natl Acad. Sci. USA 106, 1139–1144 (2009).
Chen, C. C. et al. RAB-10 is required for endocytic recycling in the Caenorhabditis elegans intestine. Mol. Biol. Cell 17, 1286–1297 (2006).
Zhang, B. & Zelhof, A. C. Amphiphysins: raising the BAR for synaptic vesicle recycling and membrane dynamics. Bin-Amphiphysin-Rvsp. Traffic 3, 452–460 (2002).
Razzaq, A. et al. Amphiphysin is necessary for organization of the excitation-contraction coupling machinery of muscles, but not for synaptic vesicle endocytosis in Drosophila. Genes Dev. 15, 2967–2979 (2001).
Zelhof, A. C. et al. Drosophila Amphiphysin is implicated in protein localization and membrane morphogenesis but not in synaptic vesicle endocytosis. Development 128, 5005–5015 (2001).
Leventis, P. A. et al. Drosophila Amphiphysin is a post-synaptic protein required for normal locomotion but not endocytosis. Traffic 2, 839–850 (2001).
Grabs, D. et al. The SH3 domain of amphiphysin binds the proline-rich domain of dynamin at a single site that defines a new SH3 binding consensus sequence. J. Biol Chem. 272, 13419–13425 (1997).
Burack, M. A., Silverman, M. A. & Banker, G. The role of selective transport in neuronal protein sorting. Neuron 26, 465–472 (2000).
Naslavsky, N., Weigert, R. & Donaldson, J. G. Characterization of a nonclathrin endocytic pathway: membrane cargo and lipid requirements. Mol. Biol. Cell 5, 3542–3552 (2004).
Naslavsky, N., Weigert, R. & Donaldson, J. G. Convergence of non-clathrin- and clathrin-derived endosomes involves Arf6 inactivation and changes in phosphoinositides. Mol. Biol. Cell 14, 417–431 (2003).
Farsad, K. et al. A putative role for intramolecular regulatory mechanisms in the adaptor function of amphiphysin in endocytosis. Neuropharmacology 45, 787–796 (2003).
McMahon, H. T., Wigge, P. & Smith, C. Clathrin interacts specifically with amphiphysin and is displaced by dynamin. FEBS Lett. 413, 319–322 (1997).
Olesen, L. E. et al. Solitary and repetitive binding motifs for the AP2 complex alpha-appendage in amphiphysin and other accessory proteins. J. Biol. Chem. 283, 5099–5109 (2008).
Owen, D. J. et al. Crystal structure of the amphiphysin-2 SH3 domain and its role in the prevention of dynamin ring formation. EMBO J. 17, 5273–5285 (1998).
Slepnev, V. I. & De Camilli, P. Accessory factors in clathrin-dependent synaptic vesicle endocytosis. Nature Rev. Neurosci. 1, 161–172 (2000).
Yoshida, Y. et al. The stimulatory action of amphiphysin on dynamin function is dependent on lipid bilayer curvature. EMBO J. 23, 3483–3491 (2004).
Lee, E. et al. Amphiphysin 2 (Bin1) and T-tubule biogenesis in muscle. Science 297, 1193–1196 (2002).
Muller, A. J. et al. Targeted disruption of the murine Bin1/Amphiphysin II gene does not disable endocytosis but results in embryonic cardiomyopathy with aberrant myofibril formation. Mol. Cell. Biol. 23, 4295–4306 (2003).
Jovic, M., Kieken, F., Naslavsky, N., Sorgen, P. L. & Caplan, S. Eps15 homology domain 1-associated tubules contain phosphatidylinositol-4-phosphate and phosphatidylinositol-(4, 5)-bisphosphate and are required for efficient recycling. Mol. Biol. Cell 20, 2731–2743 (2009).
Yamashiro, D. J., Tycko, B., Fluss, S. R. & Maxfield, F. R. Segregation of transferrin to a mildly acidic (pH 6.5) para-Golgi compartment in the recycling pathway. Cell 37, 789–800 (1984).
Takei, K., Slepnev, V. I. & De Camilli, P. Interactions of dynamin and amphiphysin with liposomes. Methods Enzymol. 329, 478–486 (2001).
Brown, F. D., Rozelle, A. L., Yin, H. L., Balla, T. & Donaldson, J. G. Phosphatidylinositol 4, 5-bisphosphate and Arf6-regulated membrane traffic. J. Cell Biol. 154, 1007–1017 (2001).
Pucadyil, T. J. & Schmid, S. L. Real-time visualization of dynamin-catalyzed membrane fission and vesicle release. Cell 135, 1263–1275 (2008).
Bashkirov, P. V. et al. GTPase cycle of Dynamin is coupled to membrane squeeze and release, leading to spontaneous fission. Cell (2008).
Braun, A. et al. EHD proteins associate with syndapin I and II and such interactions play a crucial role in endosomal recycling. Mol. Biol. Cell 16, 3642–3658 (2005).
Roux, A., Uyhazi, K., Frost, A. & De Camilli, P. GTP-dependent twisting of dynamin implicates constriction and tension in membrane fission. Nature 441, 528–531 (2006).
Kamath, R. S. & Ahringer, J. Genome-wide RNAi screening in Caenorhabditis elegans. Methods 30, 313–321 (2003).
Balklava, Z., Pant, S., Fares, H. & Grant, B. D. Genome-wide analysis identifies a general requirement for polarity proteins in endocytic traffic. Nature Cell Biol. 9, 1066–1073 (2007).
Choi, J. Y., Stukey, J., Hwang, S. Y. & Martin, C. E. Regulatory elements that control transcription activation and unsaturated fatty acid-mediated repression of the Saccharomyces cerevisiae OLE1 gene. J. Biol. Chem. 271, 3581–3589 (1996).
Grant, B. & Greenwald, I. Structure, function, and expression of SEL-1, a negative regulator of LIN-12 and GLP-1 in C. elegans. Development 124, 637–644 (1997).
Brenner, S. The genetics of Caenorhabditis elegans. Genetics 77, 71–94 (1974).
Praitis, V., Casey, E., Collar, D. & Austin, J. Creation of low-copy integrated transgenic lines in Caenorhabditis elegans. Genetics 157, 1217–1226 (2001).
Edelhoch, H. Spectroscopic determination of tryptophan and tyrosine in proteins. Biochemistry 6, 1948–1954 (1967).
Acknowledgements
We thank P. McPherson, Z. Zhou, B. Kay, G. Prandergast and S. Mitani for important reagents. The monoclonal against RME-1 was made at Washington University School of Medicine and funded by Grant R24RR22234 (PI M. L. Nonet). We also thank C. Martin, P. Yurchenco, D. Winkelman, F. Matsumura, S. Yamashiro and S. H. deGregori for sharing their instruments, reagents and protocols; A. Shi for the construction and gift of the GFP–SDPN-1 strain; R. Patel and V. Starovoytov for expert assistance with electron microscopy protocols and Z. Pan for usage of the confocal microscopy facility; O. Daumke and T. Pucadyil for discussions and advice on liposome tubulation experiments; P. Schweinsberg for technical assistance. This work was supported by NIH Grants GM67237 to B.D.G and GM074876 to S.C., a NJCSCR Graduate Fellowship 06-2915-SCR-E-0, an Anne B. and James B. Leathem Fellowship, a McCallum Fund Fellowship Grant to S.P., an American Heart Association student fellowship to M.S. and a Rutgers Summer Undergraduate Research Fellowship (SURF) to K.P.
Author information
Authors and Affiliations
Contributions
S.P. participated in experimental design, performed the C. elegans-related experiments and the biochemical and liposome experiments (Figs 1, 2, 3, 4, 6, 7, 8 and Supplementary Information, Figs S1–5 and S7) and wrote the manuscript. M.S. performed the mammalian cell experiments (Fig. 5 and Supplementary Information, Fig. S6) and helped to write those sections of the manuscript. K.P. contributed to developing some strains used in the study. S.C. designed the mammalian cell experiments. C.M.C. designed the experiments related to liposome biochemistry and trained S.P. to do those experiments. B.D.G. designed the experiments, trained S.P. in all C. elegans experiments and wrote the paper.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Information
Supplementary Information (PDF 1871 kb)
Supplementary Information
Supplementary Table 1 (XLS 21 kb)
Supplementary Information
Supplementary Table 2 (XLS 28 kb)
Rights and permissions
About this article
Cite this article
Pant, S., Sharma, M., Patel, K. et al. AMPH-1/Amphiphysin/Bin1 functions with RME-1/Ehd1 in endocytic recycling. Nat Cell Biol 11, 1399–1410 (2009). https://doi.org/10.1038/ncb1986
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ncb1986
This article is cited by
-
BIN1 knockdown rescues systolic dysfunction in aging male mouse hearts
Nature Communications (2024)
-
BIN1 in the Pursuit of Ousting the Alzheimer’s Reign: Impact on Amyloid and Tau Neuropathology
Neurotoxicity Research (2023)
-
BIN1 is a key regulator of proinflammatory and neurodegeneration-related activation in microglia
Molecular Neurodegeneration (2022)
-
Metabolic disorder in Alzheimer’s disease
Metabolic Brain Disease (2021)
-
BIN1 protein isoforms are differentially expressed in astrocytes, neurons, and microglia: neuronal and astrocyte BIN1 are implicated in tau pathology
Molecular Neurodegeneration (2020)