Abstract
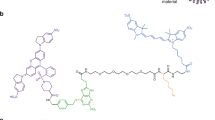
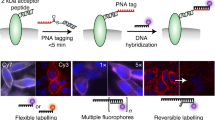
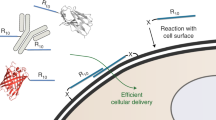
Live cell imaging is a powerful method to study protein dynamics at the cell surface, but conventional imaging probes are bulky, or interfere with protein function1,2, or dissociate from proteins after internalization3,4. Here, we report technology for covalent, specific tagging of cellular proteins with chemical probes. Through rational design, we redirected a microbial lipoic acid ligase (LplA)5 to specifically attach an alkyl azide onto an engineered LplA acceptor peptide (LAP). The alkyl azide was then selectively derivatized with cyclo-octyne6 conjugates to various probes. We labeled LAP fusion proteins expressed in living mammalian cells with Cy3, Alexa Fluor 568 and biotin. We also combined LplA labeling with our previous biotin ligase labeling7,8, to simultaneously image the dynamics of two different receptors, coexpressed in the same cell. Our methodology should provide general access to biochemical and imaging studies of cell surface proteins, using small fluorophores introduced via a short peptide tag.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on SpringerLink
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Debant, A., Ponzio, G., Clauser, E., Contreres, J.O. & Rossi, B. Receptor cross-linking restores an insulin metabolic effect altered by mutation on tyrosine 1162 and tyrosine 1163. Biochemistry 28, 14–17 (1989).
Weiss, A. & Littman, D.R. Signal transduction by lymphocyte antigen receptors. Cell 76, 263–274 (1994).
Anderson, R.G., Brown, M.S., Beisiegel, U. & Goldstein, J.L. Surface distribution and recycling of the low density lipoprotein receptor as visualized with antireceptor antibodies. J. Cell Biol. 93, 523–531 (1982).
Barak, L.S. & Webb, W.W. Fluorescent low density lipoprotein for observation of dynamics of individual receptor complexes on cultured human fibroblasts. J. Cell Biol. 90, 595–604 (1981).
Green, D.E., Morris, T.W., Green, J., Cronan, J.E., Jr. & Guest, J.R. Purification and properties of the lipoate protein ligase of Escherichia coli. Biochem. J. 309, 853–862 (1995).
Agard, N.J., Baskin, J.M., Prescher, J.A., Lo, A. & Bertozzi, C.R. A comparative study of bioorthogonal reactions with azides. ACS Chem. Biol. 1, 644–648 (2006).
Howarth, M., Takao, K., Hayashi, Y. & Ting, A.Y. Targeting quantum dots to surface proteins in living cells with biotin ligase. Proc. Natl. Acad. Sci. USA 102, 7583–7588 (2005).
Howarth, M. et al. A monovalent streptavidin with a single femtomolar biotin binding site. Nat. Methods 3, 267–273 (2006).
Marks, K.M. & Nolan, G.P. Chemical labeling strategies for cell biology. Nat. Methods 3, 591–596 (2006).
Prescher, J.A. & Bertozzi, C.R. Chemistry in living systems. Nat. Chem. Biol. 1, 13–21 (2005).
Ali, S.T. & Guest, J.R. Isolation and characterization of lipoylated and unlipoylated domains of the E2p subunit of the pyruvate dehydrogenase complex of Escherichia coli. Biochem. J. 271, 139–145 (1990).
Fujiwara, K. et al. Crystal structure of lipoate-protein ligase A from Escherichia coli. Determination of the lipoic acid-binding site. J. Biol. Chem. 280, 33645–33651 (2005).
Green, J.D., Laue, E.D., Perham, R.N., Ali, S.T. & Guest, J.R. Three-dimensional structure of a lipoyl domain from the dihydrolipoyl acetyltransferase component of the pyruvate dehydrogenase multienzyme complex of Escherichia coli. J. Mol. Biol. 248, 328–343 (1995).
Kiick, K.L., Saxon, E., Tirrell, D.A. & Bertozzi, C.R. Incorporation of azides into recombinant proteins for chemoselective modification by the Staudinger ligation. Proc. Natl. Acad. Sci. USA 99, 19–24 (2002).
Griffin, R.J. The medicinal chemistry of the azido group. Prog. Med. Chem. 31, 121–232 (1994).
Chen, I., Howarth, M., Lin, W. & Ting, A.Y. Site-specific labeling of cell surface proteins with biophysical probes using biotin ligase. Nat. Methods 2, 99–104 (2005).
Lin, C.W. & Ting, A.Y. Transglutaminase-catalyzed site-specific conjugation of small-molecule probes to proteins in vitro and on the surface of living cells. J. Am. Chem. Soc. 128, 4542–4543 (2006).
Adams, S.R. et al. New biarsenical ligands and tetracysteine motifs for protein labeling in vitro and in vivo: synthesis and biological applications. J. Am. Chem. Soc. 124, 6063–6076 (2002).
Willnow, T.E. The low-density lipoprotein receptor gene family: multiple roles in lipid metabolism. J. Mol. Med. 77, 306–315 (1999).
Reche, P. & Perham, R.N. Structure and selectivity in post-translational modification: attaching the biotinyl-lysine and lipoyl-lysine swinging arms in multifunctional enzymes. EMBO J. 18, 2673–2682 (1999).
Pasquale, E.B. Eph receptor signalling casts a wide net on cell behaviour. Nat. Rev. Mol. Cell Biol. 6, 462–475 (2005).
Singh, A.B. & Harris, R.C. Autocrine, paracrine and juxtacrine signaling by EGFR ligands. Cell. Signal. 17, 1183–1193 (2005).
Tuli, S.S. et al. Immunohistochemical localization of EGF, TGF-alpha, TGF-beta, and their receptors in rat corneas during healing of excimer laser ablation. Curr. Eye Res. 31, 709–719 (2006).
Flanagan, J.G. & Vanderhaeghen, P. The ephrins and Eph receptors in neural development. Annu. Rev. Neurosci. 21, 309–345 (1998).
Wimmer-Kleikamp, S.H. & Lackmann, M. Eph-modulated cell morphology, adhesion and motility in carcinogenesis. IUBMB Life 57, 421–431 (2005).
George, N., Pick, H., Vogel, H., Johnsson, N. & Johnsson, K. Specific labeling of cell surface proteins with chemically diverse compounds. J. Am. Chem. Soc. 126, 8896–8897 (2004).
Griffin, B.A., Adams, S.R. & Tsien, R.Y. Specific covalent labeling of recombinant protein molecules inside live cells. Science 281, 269–272 (1998).
Brock, R., Hamelers, I.H. & Jovin, T.M. Comparison of fixation protocols for adherent cultured cells applied to a GFP fusion protein of the epidermal growth factor receptor. Cytometry 35, 353–362 (1999).
McLean, A.J. & Milligan, G. Ligand regulation of green fluorescent protein-tagged forms of the human beta(1)- and beta(2)-adrenoceptors; comparisons with the unmodified receptors. Br. J. Pharmacol. 130, 1825–1832 (2000).
Zhou, Z. et al. Genetically encoded short peptide tags for orthogonal protein labeling by Sfp and AcpS phosphopantetheinyl transferases. ACS Chem. Biol. 2, 337–346 (2007).
Acknowledgements
The authors thank Mark Howarth, John Cronan, Irwin Chen, Chi-Wang Lin, Robin Prince and Martin Lackmann for their assistance and advice. This work was supported by the National Institutes of Health (R01 GM072670-01 to A.Y.T. and GM58867 to C.R.B.), the Sloan Foundation, the Dreyfus Foundation, a La Caixa Foundation predoctoral fellowship (to M.F.-S.), and National Science Foundation and National Science Defense and Engineering predoctoral fellowships (to J.M.B.).
Author information
Authors and Affiliations
Contributions
M.F.-S., H.B., L.M.-H. and A.Y.T. designed the research; M.F.-S., H.B., L.M.-H. and K.T.X. performed the research; J.M.B. and C.R.B. provided cyclo-octyne starting material; M.F.-S., H.B. and A.Y.T. analyzed data; M.F.-S. and A.Y.T. wrote the paper.
Corresponding author
Ethics declarations
Competing interests
Massachusetts Institute of Technology is seeking to file a patent application covering part of the information contained in this article.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–6; Supplementary Methods (PDF 405 kb)
Rights and permissions
About this article
Cite this article
Fernández-Suárez, M., Baruah, H., Martínez-Hernández, L. et al. Redirecting lipoic acid ligase for cell surface protein labeling with small-molecule probes. Nat Biotechnol 25, 1483–1487 (2007). https://doi.org/10.1038/nbt1355
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nbt1355
This article is cited by
-
Location-agnostic site-specific protein bioconjugation via Baylis Hillman adducts
Nature Communications (2024)
-
Further assessments of ligase LplA-mediated modifications of proteins in vitro and in cellulo
Molecular Biology Reports (2022)
-
Quantifying residue-specific conformational dynamics of a highly reactive 29-mer peptide
Scientific Reports (2020)