Abstract
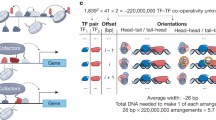
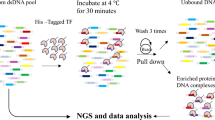
Transcription factors (TFs) interact with specific DNA regulatory sequences to control gene expression throughout myriad cellular processes. However, the DNA binding specificities of only a small fraction of TFs are sufficiently characterized to predict the sequences that they can and cannot bind. We present a maximally compact, synthetic DNA sequence design for protein binding microarray (PBM) experiments1 that represents all possible DNA sequence variants of a given length k (that is, all 'k-mers') on a single, universal microarray. We constructed such all k-mer microarrays covering all 10–base pair (bp) binding sites by converting high-density single-stranded oligonucleotide arrays to double-stranded (ds) DNA arrays. Using these microarrays we comprehensively determined the binding specificities over a full range of affinities for five TFs of different structural classes from yeast, worm, mouse and human. The unbiased coverage of all k-mers permits high-throughput interrogation of binding site preferences, including nucleotide interdependencies, at unprecedented resolution.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
References
Mukherjee, S. et al. Rapid analysis of the DNA-binding specificities of transcription factors with DNA microarrays. Nat. Genet. 36, 1331–1339 (2004).
Bulyk, M.L., Huang, X., Choo, Y. & Church, G.M. Exploring the DNA-binding specificities of zinc fingers with DNA microarrays. Proc. Natl. Acad. Sci. USA 98, 7158–7163 (2001).
Berger, M.F. & Bulyk, M.L. Protein binding microarrays (PBMs) for rapid, high-throughput characterization of the sequence specificities of DNA-binding proteins. Methods Mol. Biol. 338, 245–260 (2006).
Golomb, S. . Shift Register Sequences (Aegean Park Press, Laguna Hills, California, USA, 1967).
Kwan, A.H., Czolij, R., Mackay, J.P. & Crossley, M. Pentaprobe: a comprehensive sequence for the one-step detection of DNA-binding activities. Nucleic Acids Res. 31, e124 (2003).
Linnell, J. et al. Quantitative high-throughput analysis of transcription factor binding specificities. Nucleic Acids Res. 32, e44 (2004).
Oliphant, A.R., Brandl, C.J. & Struhl, K. Defining the sequence specificity of DNA-binding proteins by selecting binding sites from random-sequence oligonucleotides: analysis of yeast GCN4 protein. Mol. Cell. Biol. 9, 2944–2949 (1989).
Sandelin, A., Alkema, W., Engstrom, P., Wasserman, W.W. & Lenhard, B. JASPAR: an open-access database for eukaryotic transcription factor binding profiles. Nucleic Acids Res. 32, D91–D94 (2004).
Liu, J. & Stormo, G.D. Quantitative analysis of EGR proteins binding to DNA: assessing additivity in both the binding site and the protein. BMC Bioinformatics 6, 176 (2005).
Miller, J.C. & Pabo, C.O. Rearrangement of side-chains in a Zif268 mutant highlights the complexities of zinc finger–DNA recognition. J. Mol. Biol. 313, 309–315 (2001).
Bjerve, S. Error bounds for linear combinations of order statistics. Ann. Stat. 5, 357–369 (1977).
Lieb, J.D., Liu, X., Botstein, D. & Brown, P.O. Promoter-specific binding of Rap1 revealed by genome-wide maps of protein-DNA association. Nat. Genet. 28, 327–334 (2001).
Harbison, C.T. et al. Transcriptional regulatory code of a eukaryotic genome. Nature 431, 99–104 (2004).
Berg, O.G. & von Hippel, P.H. Selection of DNA binding sites by regulatory proteins. Statistical-mechanical theory and application to operators and promoters. J. Mol. Biol. 193, 723–750 (1987).
Bulyk, M.L., Johnson, P.L. & Church, G.M. Nucleotides of transcription factor binding sites exert interdependent effects on the binding affinities of transcription factors. Nucleic Acids Res. 30, 1255–1261 (2002).
Jiang, J. & Levine, M. Binding affinities and cooperative interactions with bHLH activators delimit threshold responses to the dorsal gradient morphogen. Cell 72, 741–752 (1993).
Gaudet, J. & Mango, S.E. Regulation of organogenesis by the Caenorhabditis elegans FoxA protein PHA-4. Science 295, 821–825 (2002).
Tanay, A. Extensive low-affinity transcriptional interactions in the yeast genome. Genome Res. 16, 962–972 (2006).
Warren, C.L. et al. Defining the sequence-recognition profile of DNA-binding molecules. Proc. Natl. Acad. Sci. USA 103, 867–872 (2006).
Singh-Gasson, S. et al. Maskless fabrication of light-directed oligonucleotide microarrays using a digital micromirror array. Nat. Biotechnol. 17, 974–978 (1999).
Blancafort, P., Segal, D.J. & Barbas, C.F., III. Designing transcription factor architectures for drug discovery. Mol. Pharmacol. 66, 1361–1371 (2004).
Philippakis, A.A. et al. Expression-guided in silico evaluation of candidate cis regulatory codes for Drosophila muscle founder cells. PLoS Comput. Biol. 2, e53 (2006).
Braun, P. et al. Proteome-scale purification of human proteins from bacteria. Proc. Natl. Acad. Sci. USA 99, 2654–2659 (2002).
Li, M.Z. & Elledge, S.J. MAGIC, an in vivo genetic method for the rapid construction of recombinant DNA molecules. Nat. Genet. 37, 311–319 (2005).
Dudley, A.M., Aach, J., Steffen, M.A. & Church, G.M. Measuring absolute expression with microarrays with a calibrated reference sample and an extended signal intensity range. Proc. Natl. Acad. Sci. USA 99, 7554–7559 (2002).
Morton, T.A. & Myszka, D.G. Kinetic analysis of macromolecular interactions using surface plasmon resonance biosensors. Methods Enzymol. 295, 268–294 (1998).
Wilmen, A., Pick, H., Niedenthal, R.K., Sen-Gupta, M. & Hegemann, J.H. The yeast centromere CDEI/Cpf1 complex: differences between in vitro binding and in vivo function. Nucleic Acids Res. 22, 2791–2800 (1994).
Christy, B. & Nathans, D. DNA binding site of the growth factor–inducible protein Zif268. Proc. Natl. Acad. Sci. USA 86, 8737–8741 (1989).
Okkema, P.G. & Fire, A. The Caenorhabditis elegans NK-2 class homeoprotein CEH-22 is involved in combinatorial activation of gene expression in pharyngeal muscle. Development 120, 2175–2186 (1994).
Klemm, J.D., Rould, M.A., Aurora, R., Herr, W. & Pabo, C.O. Crystal structure of the Oct-1 POU domain bound to an octamer site: DNA recognition with tethered DNA-binding modules. Cell 77, 21–32 (1994).
Acknowledgements
We thank T.V.S. Murthy, Leo Brizuela and Josh LaBaer for providing the Cbf1 and Rap1 clones, Gwenael Badis-Breard and Tim Hughes for providing the Zif268 DNA-binding domain clone and Shufen Meng for assistance with the Biacore technology. We also thank Stephen Gisselbrecht, Amy Donner and Rachel McCord for critical reading of the manuscript. This work was funded in part by grants R01 HG003985 and R01 HG003420 from National Institutes of Health/National Human Genome Research Institute to M.L.B. M.F.B. was supported in part by a National Science Foundation Graduate Research Fellowship. A.A.P. was supported in part by a National Defense Science and Engineering Graduate Fellowship, a National Science Foundation Graduate Research Fellowship and an Athinoula Martinos Fellowship.
Author information
Authors and Affiliations
Contributions
M.F.B. participated in the array design, experimental design, analysis of results and drafting of the manuscript, and performed the experiments; A.A.P. conceived the idea of using de Bruijn sequences and participated in the binding site survey, array design, experimental design, analysis of results and drafting of the manuscript; A.M.Q. provided linear feedback shift register expertise and assisted in the array design; F.S.H. participated in the binding site survey; P.W.E. III conceived the concept of the compact universal array; M.L.B. conceived the concept of the compact universal array and participated in the array design, experimental design, analysis of the results and drafting of the manuscript.
Corresponding author
Ethics declarations
Competing interests
A subset of the authors—M.L.B., A.A.P and P.W.E. 3rd—have filed a patent application, through Brigham and Women's Hospital, covering the sequence design described in this paper.
Supplementary information
Supplementary Fig. 1
Survey of binding sites in the JASPAR database. (PDF 570 kb)
Supplementary Fig. 2
Reproducibility of Cy3 dUTP signal intensities. (PDF 512 kb)
Supplementary Fig. 3
Correlation of PBM signal intensities with affinities. (PDF 679 kb)
Supplementary Fig. 4
Effects of binding site position and orientation on PBM signal. (PDF 629 kb)
Supplementary Fig. 5
Comparison of median signal intensities for 28 Zif268 variants for fixed versus variable position, orientation, and flanking sequence. (PDF 654 kb)
Supplementary Fig. 6
Correspondence between median signal intensities for 7-mers on distinct de Bruijn sequences. (PDF 296 kb)
Supplementary Fig. 7
Biacore measurements supporting interdependence between the first two positions of the Cbf1 binding site. (PDF 920 kb)
Supplementary Fig. 8
Minimum number of unique features on an array for different values of k. (PDF 652 kb)
Supplementary Fig. 9
Dependence of Cy3 dUTP incorporation upon sequence context. (PDF 609 kb)
Rights and permissions
About this article
Cite this article
Berger, M., Philippakis, A., Qureshi, A. et al. Compact, universal DNA microarrays to comprehensively determine transcription-factor binding site specificities. Nat Biotechnol 24, 1429–1435 (2006). https://doi.org/10.1038/nbt1246
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nbt1246
This article is cited by
-
KaScape: a sequencing-based method for global characterization of protein‒DNA binding affinity
Scientific Reports (2023)
-
Widespread perturbation of ETS factor binding sites in cancer
Nature Communications (2023)
-
Sequence determinants of human gene regulatory elements
Nature Genetics (2022)
-
The transcriptional regulator HDP1 controls expansion of the inner membrane complex during early sexual differentiation of malaria parasites
Nature Microbiology (2022)
-
Simple synthesis of massively parallel RNA microarrays via enzymatic conversion from DNA microarrays
Nature Communications (2022)