Abstract
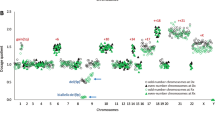
Chromosomal imbalances such as deletions and amplifications are common rearrangements in most tumors1. Specific rearrangements are consistently associated with specific tumor types or stages, implicating the role of the genes in a region of chromosomal imbalance in tumor initiation and progression2. The development of comparative genomic hybridization (CGH)3 has obviated the need to obtain metaphase spreads from tumors, so that the chromosomal imbalances in many solid tumors may be revealed using an extracted genomic DNA sample. However, the resolution of the cytogenetic method remains and the extreme technical difficulty of CGH has restricted its use. Conceptually, DNA microarray4–based CGH is an obvious solution to all of the limitations of conventional CGH. Although arrays have been used for CGH studies5,6,7,8, their success has been limited by poor specific signal-to-noise ratios. Here we demonstrate a microarray-based CGH method that allows reliable detection of chromosomal deletions and amplifications with high resolution. Our microarray system is fundamentally different from most current microarray technologies in that activated DNA is printed on natural glass surfaces while other systems almost exclusively focus on activating the surfaces, a strategy that invariably introduces hybridization backgrounds. The concept of using pre-modification may be generally applied for making arrays of other biological materials, as modifying the substrates will be more controllable in solution than on surfaces.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
References
Mitelman, F., Mertens, F. & Johansson, B. A breakpoint map of recurrent chromosomal rearrangements in human neoplasia. Nat. Genet. 15 (Spec No.), 417–474 (1997).
Lengauer, C., Kinzler, K.W. & Vogelstein, B. Genetic instabilities in human cancers. Nature 396, 643–649 (1998).
Kallioniemi, A. et al. Comparative genomic hybridization for molecular cytogenetic analysis of solid tumors. Science 258, 818–821 (1992).
Schena, M., Shalon, D., Davis, R.W. & Brown, P.O. Quantitative monitoring of gene-expression patterns with a complementary-DNA microarray. Science 270, 467–470 (1995).
Solinas-Toldo, S. et al. Matrix-based comparative genomic hybridization: biochips to screen for genomic imbalances. Genes Chromosomes Cancer 20, 399–407 (1997).
Pinkel, D. et al. High-resolution analysis of DNA copy number variation using comparative genomic hybridization to microarrays. Nat. Genet. 20, 207–211 (1998).
Pollack, J.R. et al. Genome-wide analysis of DNA copy-number changes using cDNA microarrays. Nat. Genet. 23, 41–46 (1999).
Hodgson, G. et al. Genome scanning with array CGH delineates regional alterations in mouse islet carcinomas. Nat. Genet. 29, 459–464 (2001).
Cai, W.W., Chow, C.W., Damani, S., Simon, G.G. & Bradley, A. An SSLP marker anchored BAC framework map of the mouse genome. Nat. Genet. 29, 133–134 (2001).
Okano, H. et al. Homozygous deletions and point mutations of the Ikaros gene in γ-ray-induced mouse thymic lymphomas. Oncogene 18, 6677–6683 (1999).
Matsumoto, Y. et al. Allelic loss analysis of γ-ray-induced mouse thymic lymphomas: two candidate tumor suppressor gene loci on chromosomes 12 and 16. Oncogene 16, 2747–2754 (1998).
Liu, P., Zhang, H., McLellan, A., Vogel, H. & Bradley, A. Embryonic lethality and tumorigenesis caused by segmental aneuploidy on mouse chromosome 11. Genetics 150, 1155–1168 (1998).
Dietrich, W.F. et al. Genome-wide search for loss of heterozygosity in transgenic mouse tumors reveals candidate tumor suppressor genes on chromosomes 9 and 16. Proc. Natl. Acad. Sci. USA 91, 9451–9455 (1994).
Parangi, S., Dietrich, W., Christofori, G., Lander, E.S. & Hanahan, D. Tumor suppressor loci on mouse chromosome 9 and 16 are lost at distinct stages of tumorigenesis in a transgenic model of islet cell carcinoma. Cancer Res. 55, 6071–6076 (1995).
The BAC Resource Consortium. Integration of cytogenetic landmarks into the draft sequence of the human genome. Nature 409, 953–958 (2001).
The International Human Genome Mapping Consortium. A physical map of the human genome. Nature 409, 934–941 (2001).
International Human Sequencing Consortium. Initial sequencing and analysis of the human genome. Nature 409, 860–921 (2001).
Venter, J.C. et al. The sequence of the human genome. Science 291, 1304–1351 (2001).
Cai, W.W., Reneker, J., Chow, C.W., Vaishnav, M. & Bradley, A. An anchored framework BAC map of mouse chromosome 11 assembled using multiplex oligonucleotide hybridization. Genomics 54, 387–397 (1998).
Donehower, L.A. et al. Mice deficient for p53 are developmentally normal but susceptible to spontaneous tumours. Nature 356, 215–221 (1992).
Acknowledgements
W.W.C. was a Jane Coffin Childs research fellow. This work was supported by NCI and the Baylor prostate SPORE program. We thank Alec Wang for the cDNA clones and other valuable reagents.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Cai, WW., Mao, JH., Chow, CW. et al. Genome-wide detection of chromosomal imbalances in tumors using BAC microarrays. Nat Biotechnol 20, 393–396 (2002). https://doi.org/10.1038/nbt0402-393
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1038/nbt0402-393
This article is cited by
-
FISH molecular testing in cytological preparations from solid tumors
Molecular Cytogenetics (2014)
-
Hipk2 cooperates with p53 to suppress γ-ray radiation-induced mouse thymic lymphoma
Oncogene (2012)
-
An HDAC1-binding domain within FATS bridges p21 turnover to radiation-induced tumorigenesis
Oncogene (2010)
-
Atm heterozygosity does not increase tumor susceptibility to ionizing radiation alone or in a p53 heterozygous background
Oncogene (2008)
-
Array-based comparative genomic hybridization of mapped BAC DNA clones to screen for chromosome 14 copy number abnormalities in meningiomas
European Journal of Human Genetics (2008)