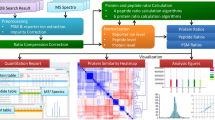
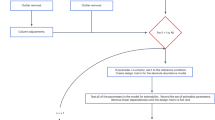
Abstract
Replicate mass spectrometry (MS) measurements and the use of multiple analytical methods can greatly expand the comprehensiveness of shotgun proteomic profiling of biological samples1,2,3,4,5. However, the inherent biases and variations in such data create computational and statistical challenges for quantitative comparative analysis6. We developed and tested a normalized, label-free quantitative method termed the normalized spectral index (SIN), which combines three MS abundance features: peptide count, spectral count and fragment-ion (tandem MS or MS/MS) intensity. SIN largely eliminated variances between replicate MS measurements, permitting quantitative reproducibility and highly significant quantification of thousands of proteins detected in replicate MS measurements of the same and distinct samples. It accurately predicts protein abundance more often than the five other methods we tested. Comparative immunoblotting and densitometry further validate our method. Comparative quantification of complex data sets from multiple shotgun proteomics measurements is relevant for systems biology and biomarker discovery.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on SpringerLink
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Durr, E. et al. Direct proteomic mapping of the lung microvascular endothelial cell surface in vivo and in cell culture. Nat. Biotechnol. 22, 985–992 (2004).
Li, Y. et al. Enhancing identifications of lipid-embedded proteins by mass spectrometry for improved mapping of endothelial plasma membranes in vivo. Mol. Cell. Proteomics 8, 1219–1235 (2009).
Oh, P. et al. Subtractive proteomic mapping of the endothelial surface in lung and solid tumours for tissue-specific therapy. Nature 429, 629–635 (2004).
Slebos, R.J. et al. Evaluation of strong cation exchange versus isoelectric focusing of peptides for multidimensional liquid chromatography-tandem mass spectrometry. J. Proteome Res. 7, 5286–5294 (2008).
Kislinger, T., Gramolini, A.O., MacLennan, D.H. & Emili, A. Multidimensional protein identification technology (MudPIT): technical overview of a profiling method optimized for the comprehensive proteomic investigation of normal and diseased heart tissue. J. Am. Soc. Mass Spectrom. 16, 1207–1220 (2005).
Wong, J.W., Sullivan, M.J. & Cagney, G. Computational methods for the comparative quantification of proteins in label-free LCn-MS experiments. Brief. Bioinform. 9, 156–165 (2008).
Oh, P. et al. Live dynamic imaging of caveolae pumping targeted antibody rapidly and specifically across endothelium in the lung. Nat. Biotechnol. 25, 327–337 (2007).
Shiio, Y. et al. Quantitative proteomic analysis of Myc oncoprotein function. EMBO J. 21, 5088–5096 (2002).
Shiio, Y. et al. Quantitative proteomic analysis of myc-induced apoptosis: a direct role for Myc induction of the mitochondrial chloride ion channel, mtCLIC/CLIC4. J. Biol. Chem. 281, 2750–2756 (2006).
Chiang, M.C. et al. Systematic uncovering of multiple pathways underlying the pathology of Huntington disease by an acid-cleavable isotope-coded affinity tag approach. Mol. Cell. Proteomics 6, 781–797 (2007).
Service, R.F. Proteomics. Proteomics ponders prime time. Science 321, 1758–1761 (2008).
Service, R.F. Proteomics. Will biomarkers take off at last? Science 321, 1760 (2008).
Kolodziej, E.P., Gray, J.L. & Sedlak, D.L. Quantification of steroid hormones with pheromonal properties in municipal wastewater effluent. Environ. Toxicol. Chem. 22, 2622–2629 (2003).
Wolf-Yadlin, A., Hautaniemi, S., Lauffenburger, D.A. & White, F.M. Multiple reaction monitoring for robust quantitative proteomic analysis of cellular signaling networks. Proc. Natl. Acad. Sci. USA 104, 5860–5865 (2007).
Kuhn, E. et al. Quantification of C-reactive protein in the serum of patients with rheumatoid arthritis using multiple reaction monitoring mass spectrometry and 13C-labeled peptide standards. Proteomics 4, 1175–1186 (2004).
Ross, P.L. et al. Multiplexed protein quantitation in Saccharomyces cerevisiae using amine-reactive isobaric tagging reagents. Mol. Cell. Proteomics 3, 1154–1169 (2004).
Koziol, J.A., Feng, A.C. & Schnitzer, J.E. Application of capture-recapture models to estimation of protein count in MudPIT experiments. Anal. Chem. 78, 3203–3207 (2006).
Ishihama, Y. et al. Exponentially modified protein abundance index (emPAI) for estimation of absolute protein amount in proteomics by the number of sequenced peptides per protein. Mol. Cell. Proteomics 4, 1265–1272 (2005).
Rappsilber, J., Ryder, U., Lamond, A.I. & Mann, M. Large-scale proteomic analysis of the human spliceosome. Genome Res. 12, 1231–1245 (2002).
Zybailov, B. et al. Statistical analysis of membrane proteome expression changes in Saccharomyces cerevisiae. J. Proteome Res. 5, 2339–2347 (2006).
Old, W.M. et al. Comparison of label-free methods for quantifying human proteins by shotgun proteomics. Mol. Cell. Proteomics 4, 1487–1502 (2005).
Silva, J.C. et al. Absolute quantification of proteins by LCMSE: a virtue of parallel MS acquisition. Mol. Cell. Proteomics 5, 144–156 (2006).
Klimek, J. et al. The standard protein mix database: a diverse data set to assist in the production of improved Peptide and protein identification software tools. J. Proteome Res. 7, 96–103 (2008).
Bland, J.M. & Altman, D.G. Statistical methods for assessing agreement between two methods of clinical measurement. Lancet 1, 307–310 (1986).
Callister, S.J. et al. Normalization approaches for removing systematic biases associated with mass spectrometry and label-free proteomics. J. Proteome Res. 5, 277–286 (2006).
Wang, W. et al. Quantification of proteins and metabolites by mass spectrometry without isotopic labeling or spiked standards. Anal. Chem. 75, 4818–4826 (2003).
Lukas, T.J. et al. Informatics-assisted protein profiling in a transgenic mouse model of amyotrophic lateral sclerosis. Mol. Cell. Proteomics 5, 1233–1244 (2006).
Forner, F. et al. Quantitative proteomic comparison of rat mitochondria from muscle, heart, and liver. Mol. Cell. Proteomics 5, 608–619 (2006).
Choi, H., Fermin, D. & Nesvizhskii, A.I. Significance analysis of spectral count data in label-free shotgun proteomics. Mol. Cell. Proteomics 7, 2373–2385 (2008).
Baggerly, K.A. et al. A comprehensive approach to the analysis of matrix-assisted laser desorption/ionization-time of flight proteomics spectra from serum samples. Proteomics 3, 1667–1672 (2003).
Wagner, M., Naik, D. & Pothen, A. Protocols for disease classification from mass spectrometry data. Proteomics 3, 1692–1698 (2003).
Anderle, M. et al. Quantifying reproducibility for differential proteomics: noise analysis for protein liquid chromatography-mass spectrometry of human serum. Bioinformatics 20, 3575–3582 (2004).
Kramer, C.Y. Extension of multiple range tests to group means with unequal numbers of replications. Biometrics 12, 309–310 (1956).
Tukey, J.W. Some selected quick and easy methods of statistical analysis. Trans. N.Y. Acad. Sci. 16, 88–97 (1953).
Oh, P. & Schnitzer, J.E. Isolation and subfractionation of plasma membranes to purify caveolae separately from glycosyl-phosphatidylinositol-anchored protein microdomain. in Cell Biology: A Laboratory Handbook (ed. C.J.) 34–36 (Academic Press, Orlando, FL, USA, 1998).
Schnitzer, J.E. et al. Separation of caveolae from associated microdomains of GPI-anchored proteins. Science 269, 1435–1439 (1995).
Beissbarth, T. et al. Statistical modeling of sequencing errors in SAGE libraries. Bioinformatics 20 Suppl 1, i31–i39 (2004).
Kendall, M. Multivariate Analysis, edn. 2 (Macmillan, New York, 1980).
Mirkin, B. Mathematical Classification and Clustering (Kluwer Academic Publishers, Dordrecht, The Netherlands; 1996).
Cheng, Y. & Church, G.M. in Proceedings of the 8th International Conference on Intelligent Systems for Molecular Biology 93-1032000, August 19–23, 2000 (AAAI Press, Menlo Park, CA, 2000).
Hartigan, J. Direct clustering of a data matrix. J. Amer. Stat. Assoc. 67, 123–129 (1972).
Acknowledgements
This work was supported by National Institute of Health grants (to J.E.S): RO1HL074063, R33CA118602 and P01CA104898.
Author information
Authors and Affiliations
Contributions
N.M.G. designed, developed and analyzed the methods, provided some of the mass spectrometry data, performed the spiking experiments and analysis and wrote the manuscript; J.Y. initiated the project, designed, tested and implemented the methods; F.L. developed the scripts for data extraction; P.O. performed western blot analysis and densitometry; S.S. performed western blot analysis; Y.L. provided key mass spectrometry data; J.A.K. provided direction for statistical analysis; J.E.S supervised the project, designed specific tests and helped to write the manuscript. All authors have read and agreed to all the content in this manuscript.
Corresponding author
Supplementary information
Supplementary Text and Figures
Supplementary Figs. 1–7, Supplementary Table 1, Supplementary Notes, Supplementary Methods, Supplementary Data and Supplementary Discussion (PDF 827 kb)
Rights and permissions
About this article
Cite this article
Griffin, N., Yu, J., Long, F. et al. Label-free, normalized quantification of complex mass spectrometry data for proteomic analysis. Nat Biotechnol 28, 83–89 (2010). https://doi.org/10.1038/nbt.1592
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nbt.1592
This article is cited by
-
Identification of the Beta Subunit Fas1p of Fatty Acid Synthetase as an Interacting Partner of Yeast Calcium/Calmodulin-Dependent Protein Kinase Cmk2p Through Mass Spectrometry Analysis
Applied Biochemistry and Biotechnology (2024)
-
metaSpectraST: an unsupervised and database-independent analysis workflow for metaproteomic MS/MS data using spectrum clustering
Microbiome (2023)
-
Spectrin-beta 2 facilitates the selective accumulation of GABAA receptors at somatodendritic synapses
Communications Biology (2023)
-
High-quality and robust protein quantification in large clinical/pharmaceutical cohorts with IonStar proteomics investigation
Nature Protocols (2023)
-
MetaAnalyst: a user-friendly tool for metagenomic biomarker detection and phenotype classification
BMC Medical Research Methodology (2022)