Abstract
Patterns of genetic diversity between populations are often used to detect loci under selection in genome scans. Indeed, loci involved in local adaptations should show high FST values, whereas loci under balancing selection should rather show low FST values. Most tests of selection based on FST use a null distribution generated under a simple island model of population differentiation. Although this model has been shown to be robust, many species have a more complex genetic structure, with some populations sharing a recent ancestry or due to the presence of barriers to gene flow between different parts of a species range. In this paper, we propose the use of a hierarchical island model, in which demes exchange more migrants within groups than between groups, to generate the joint distribution of genetic diversity within and between populations. We show that tests not accounting for a hierarchical structure, when it exists, do generate a large excess of false positive loci, whereas the hierarchical island model is robust to uncertainties about the exact number of groups and demes per group in the system. Our approach also explicitly takes into account the mutational process, and does not just rely on allele frequencies, which is important for short tandem repeat (STR) data. An application to human and stickleback STR data sets reveals a much lower number of significant loci than previously obtained under a non-hierarchical model. The elimination of false positive loci from genome scans should allow us to better determine on which specific class of genes selection is operating.
Similar content being viewed by others
Introduction
Past episodes of selection leave distinct signatures in the genome of a population (Nielsen et al., 2007). Balancing selection leads to an excess of genetic diversity in the vicinity of the selected locus (Gillespie, 1990; Kreitman and Hudson, 1991), whereas positive selection leads to genomic regions harboring reduced levels of diversity (Kaplan et al., 1989), an altered allele frequency spectrum (Braverman et al., 1995) and a locally increased extent of linkage disequilibrium (Kim and Stephan, 2002; Przeworski, 2002; McVean, 2007). Several procedures have been proposed to detect loci under selection based on the patterns of genetic diversity found in a population (Tajima, 1989; Fu, 1997; Fay and Wu, 2000; Kim and Stephan, 2002; Sabeti et al., 2002; Nielsen et al., 2005; Voight et al., 2006; Zeng et al., 2006). However, selection can also affect genetic diversity between populations, as a locus under balancing selection should show too even allele frequencies across populations and loci under local directional selection should show large differences between populations (Cavalli-Sforza, 1966; Lewontin and Krakauer, 1973). This observation has recently led to the development of several methods comparing levels of genetic diversity and differentiation within and between populations (Beaumont and Nichols, 1996; Schlotterer, 2002; Beaumont and Balding, 2004; Tang et al., 2007; Foll and Gaggiotti, 2008; Riebler et al., 2008), which have been applied to various genome scan studies to detect specific loci under selection (see for example, Kayser et al., 2003; Storz et al., 2004; Storz, 2005; Tang et al., 2007; Thornton and Jensen, 2007; Egan et al., 2008; Mäkinen et al., 2008). Although tests of selection based on various aspects of genetic diversity within population seem very sensitive to past demographic events, such as population bottlenecks or demographic expansions (Teshima et al., 2006; Nielsen et al., 2007), the sensitivity of tests based on inter-population differences (summarized by the FST statistic) has been little investigated (but see for example, Beaumont and Nichols, 1996; Schlotterer, 2002).
Tests based on the comparison of FST across loci consists of identifying loci that present FST coefficients that are significantly more different than expected under neutrality and a given demographic model (they are called outlier loci). The main difficulty of these tests is thus to obtain the expected FST distribution (Beaumont, 2005). Lewontin and Krakauer (1973), proposed to obtain, by simulation, the expected variance of FST across loci for different underlying distributions of allele frequencies (binomial, uniform and U-shaped), and suggested a way to test whether the observed variance in FST could be obtained under neutrality, without providing a rigorous way to identify outlier loci. Beaumont and Nichols (1996) proposed to obtain the distribution of FST across loci as a function of heterozygosity between populations by carrying out simulations under an infinite island model, and to specifically identify outlier loci as being those present in the tails of the generated distribution. They have shown that this simple island model led to FST distributions that were very similar to those expected under alternative models, like scenarios of recent divergence and growth (colonization), isolation by distance (2-D stepping stone) or heterogeneous levels of gene flow between populations. Recent Bayesian approaches (Beaumont and Balding, 2004; Foll and Gaggiotti, 2008) assume that allele frequencies within population follow a multinomial Dirichlet distribution (Balding and Nichols, 1995; Rannala and Hartigan, 1996; Balding, 2003) with FST parameters that are a function of population-specific components shared among all loci and of locus-specific components shared among all populations. In these Bayesian approaches, departure from neutrality at a given locus is assumed when the locus-specific component is necessary to explain the observed pattern of diversity. A Dirichlet distribution of allele frequencies is expected to occur under a wide range of demographic models, in which sampled populations exchange genes with a unique and common migrant pool (Balding, 2003; Beaumont, 2005). The robustness of this model arises because the structure of the coalescent process in the migrant pool is similar to that of a standard coalescent (but on a different time scale), irrespective of the exact genetic structure of the underlying (meta-) population (Wakeley and Aliacar, 2001). Therefore, an infinite island model, in which the migrant pool is the set of all unsampled populations (Beaumont, 2005), should also lead to a Dirichlet distribution of allele frequencies among populations. This property explains why a simple island model can generate FST distributions similar to those expected under other models with different, but unique, migrant pools (Beaumont and Nichols, 1996). However, as noted very early on (Lewontin and Krakauer, 1975; Nei and Maruyama, 1975; Robertson, 1975a, 1975b), tests based on a single distribution of FST assume that all sampled populations are independent or contribute equally to the same migrant pool. Therefore, one would expect that current tests based on Dirichlet distributions or on the infinite or finite island model would not be appropriate if different samples are drawn from the same population; if some of the sampled populations share some recent ancestry; if some sampled populations contribute to different migrant pools; or if there is a hierarchical population structure. Thus even though recent Bayesian methods (for example, Beaumont and Balding, 2004; Foll and Gaggiotti, 2008; Riebler et al., 2008) explicitly allow sampled populations to receive unequal number of migrants from the migrant pool, they still assume that migrant genes originate from the same pool. Following Tsakas and Krimbas (1976), Vitalis et al. (2001) have proposed to over-ride the problem of shared and complex population histories by focusing on pairs of populations. They used the joint distribution of population-specific F statistics in two populations having diverged from a common ancestor, and conditioned it to a given number of observed alleles to build confidence regions for neutral loci and to identify outliers. Although this approach should be valid for pairs of populations having diverged without further migrations, its extension to more than two populations implies a correction for multiple and non-independent tests, which is far from being a trivial exercise.
To allow for heterogeneous affinities between sampled populations, we propose here to extend the FDIST approach of Beaumont and Nichols (1996) by using an explicit hierarchical island model (Slatkin and Voelm, 1991), in which populations samples are assigned to different groups (defined a priori), and allowing for different migrations rates between demes within groups and between groups. We show by simulations that the false positive rate is correct if the hierarchical analysis is used but not if the hierarchical structure is ignored. The application of our approach to human and stickleback genome scans carried out using short tandem repeats (STRs) shows a much smaller number of outlier loci than in previous studies relying on a non-hierarchical structure (Foll and Gaggiotti, 2008; Mäkinen et al., 2008).
Materials and methods
Coalescent simulations conditioned on observed levels of diversity within and between demes
Beaumont and Nichols (1996) have proposed to detect loci under selection by identifying those loci for which the extent of differentiation between populations (summarized by the statistic FST) is incompatible with heterozygosity between populations (h1). They obtained the joint distribution of these two statistics by simulations carried out under an island model with a finite (but large) number of demes. Coalescent simulations were carried out after inferring the migration rate m between demes of size N, which would lead to the observed FST statistic using the classical relationship FST=1/(1+4Nm) obtained by Wright (1951) under the infinite island model. FST at the i-th locus was computed as described by Weir and Cockerham (1984)

where f̂0 is the average homozygosity within population and f̂1 is the probability that two genes from different demes are identical. Global FST was then computed as a weighted average among loci, where F̂STi values are weighed by the heterozygosities between populations ĥ1i=1−f̂1i (Weir and Cockerham, 1984).
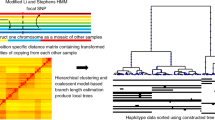
We propose here to extend this approach to the case of a hierarchical population structure, in which demes are arranged into k groups of d demes, and in which migration rates between demes are different within (m1) and between (m2) groups (see Figure 1). Slatkin and Voelm (1991) have called this particular population structure a hierarchical island model and have derived relationships between Nei's G statistics (Nei, 1973) and migration rates within and between groups. We have used their study to derive equivalent relationships between these migration rates and Wright's F statistics FST, FSC and FCT for fixed values of k and d (see Appendix). Continuous time coalescent simulations in a hierarchically structured population (Notohara, 1990; Nordborg, 1997) were implemented in a modified version of the Arlequin software package (Excoffier et al., 2005), allowing us to simulate genetic diversity in a series of samples arranged in an arbitrary hierarchical island structure, conditional on observed F-statistics globally calculated under an analysis of molecular variance framework (Excoffier et al., 1992).
Representation of the hierarchical island model used in this study. The number of groups is here k=4, and the number of demes within group is d=5. All demes are assumed to be made up of N diploid individuals. Each generation and in each deme, a proportion m1 of the 2N gene copies are immigrants from other demes belonging to the same group and a proportion m2 come from demes belonging to other groups.
Null distribution of FST under the hierarchical island model
Following Beaumont and Nichols (1996), we obtain the null distribution of FST for different levels of heterozygosity by simulation, which now takes explicitly into account a given hierarchical structure of the populations, in which populations are divided into groups given some prior knowledge (that is, geography), or according to a preliminary study of their genetic structure (using other programs, see for example, Pritchard et al., 2000; Dupanloup et al., 2002; Corander et al., 2004; Guillot et al., 2005). The number of simulated demes per group (d) and the number of groups (k) need to be specified for the simulations. The specified value of d has to be larger than the actual number of sampled demes per group and k has to be equal to or larger than the actual number of group (see Discussion for their effects on the results). Population samples belonging to the same group are then assigned to different demes of a group and sampled groups are assigned randomly to one of the k-simulated group.
We have implemented different mutation models in our coalescent simulations. For the infinite allele model (IAM) and the stepwise mutation model (SMM), we obtain for each simulation a different mutation rate by drawing a target heterozygosity at random from a uniform distribution, and use classical relationships between heterozygosity and scaled mutation rate θ=4kdNu as θ=(1−H)−1−1 under IAM (Wright, 1931), and θ=½[(1−H)−2−1] under SMM (Ohta and Kimura, 1973). For DNA data, we assumed that we would only test polymorphic loci and we have therefore implemented a single nucleotide polymorphism (SNP) mutation model, in which a mutation occurs at random along the simulated structured coalescent tree.
For each simulation, we carried out a hierarchical analysis of molecular variance and computed F-statistics, as implemented in the Arlequin software package (Excoffier et al., 2005). For SNPs, or data following the IAM model, the analysis of molecular variance was carried out using only the information on allele frequencies, whereas information on allele length was also used for data following the SMM model (Michalakis and Excoffier, 1996). Instead of computing heterozygosities between populations ĥ1 as in Beaumont and Nichols (1996), we rather computed the average heterozygosity within populations ĥ0 from which we deduced a quantity Ĥ1 equivalent to ĥ1 from eq (1), as

which shows that our test will actually compare scaled levels of diversity within (ĥ0) and between populations (F̂ST).
Locus-specific F̂ST P-values were obtained from the simulated joint distribution of Ĥ1 and F̂ST by a kernel density approach. We first estimated the density of an arbitrary value of FST (usually between 0 and 1) conditional on the observed heterozygosity Ĥ1i at the i-th locus as

where KΔF() is an Epanechnikov kernel for FST with bandwith ΔF=0.05, KΔH() is an Epanechnikov kernel for heterozygosity Ĥ1 with bandwith ΔH=0.04 and the summation is over all simulations (see for example, Beaumont et al., 2002). These bandwidths are rather arbitrary, but they allow reliable and consistent computations of P-values (see Discussion). The P-value is then obtained by (numerically) integrating the density as

where F Stmin is the minimum simulated FST value.
Test data sets
We used test data sets consisting of 50 population samples of 25 diploid individuals arranged into five groups of 10 samples each. For each data set, we simulated, under a hierarchical model (with 10 groups of 100 demes), the genetic diversity at 1000 STR loci (under a pure SMM model) or at 1000 SNP loci for two different sets of F-statistics. In the first set, the extent of differentiation between groups is larger (FCT=0.2) than that within groups (FSC=0.05), such that FST=1−(1−FSC)(1−FCT)=0.24. In the second set, groups are little differentiated (FCT=0.05), although populations within groups show more extensive differences (FSC=0.10) leading to FST=0.14. For each simulation condition, we generated and tested 100 independent data sets, such as to have a total of 100 000 loci simulated under each condition. Unless indicated otherwise, all simulated data were analysed under either a finite island model with 100 demes of size 1000 or a hierarchical island model with 10 groups of 100 demes. In the hierarchical model, the 50 populations samples are allocated as follows: 5 groups are chosen at random, and for each of these 5 selected groups, population samples are allocated to 10 randomly selected demes (among the 100 available demes).
Results
Importance of the mutation model for STR data
We first evaluated whether simulated F-statistics were correctly recovered from the data. For microsatellites, F-statistics are usually inferred under an SMM model (see for example, Slatkin, 1995), whereas current methods based on FST and aiming at inferring patterns of selection at STR loci all assume an IAM model (Beaumont and Nichols, 1996; Beaumont and Balding, 2004; Foll and Gaggiotti, 2008; Riebler et al., 2008). In Figure 2, we report FST distributions obtained under different combinations of mutation and migration models. We see that the average F-statistics are well recovered if the correct mutation and migration models are used (see Figure 2a, in which FST is computed as ρST). The use of an IAM model to estimate F-statistics under a hierarchical island model (Figure 2b) leads to an underestimation of the extent of population differentiation (F̂ST=0.2 instead of 0.24), because alleles identical in state are assumed to be identical by descent. The null distribution of FST we generated for the test in Figure 2b, is therefore shifted towards too low FST values, which should translate into an excess of loci assumed to be under directional selection and a lower power to detect loci under balancing selection. Another important difference between FST and ρST estimates is that FST computed on allele frequencies tend to decrease for very large heterozygosity levels when data are generated under a hierarchical island model. Such a constraint does not occur under an SMM model and loci with very high heterozygosities within population can show high ρST values (Figure 2a). We therefore chose to carry out our tests of selection based on the SMM model, and the FST values reported below for STR loci have been actually computed as ρST.
Example of FST distributions obtained under different combinations of mutation and migration models. The diversity of 1000 short term repeat (STR) loci (open circles) was simulated under a hierarchical island model with 10 groups of 100 demes. The migration rates within and between groups were adjusted such as to have FSC=0.05 and FCT=0.2, implying an FST of 0.240. The joint null distribution of FST and heterozygosity (20 000 grey dots) was then obtained under either a hierarchical island (a, b) or a finite island model (c, d), based on F-statistics computed either assuming an infinite allele (b, d) or a stepwise mutation model (a, c). \(\overline{F}\)ST or \(\overline{Ro}\)ST are the weighed average F-statistics computed over 1000 loci for a given genetic structure under the infinite allele model (IAM) or the stepwise mutation model (SMM), respectively.
The loci simulated under a hierarchical island model were also analysed under a finite island model (Figures 1d and 2c). In that case, estimated average FST values are also biased downwards, but this is because FST here is close to an average computed between all pairs of populations, irrespective of the group to which they belong, and the comparison of populations from the same group will largely contribute to a reduced global FST. In contrast, the global FST estimated under a hierarchical island model is close to the average FST computed between populations of different groups (Figure 2a). Another major drawback of the use of the finite island model is that it generates overly narrow FST distributions and leads to a large excess of false positive results (see Figures 2c and d), as all populations are assumed independent. We further quantify this effect in the next section.
False positive rates under finite and hierarchical island models
In Table 1, we report the analysis of STR and SNP loci simulated under a hierarchical island model and analysed under either a finite island or a hierarchical island model. We generally see that data analysed without taking the hierarchical genetic structure into account show a large excess of false positives for all significance levels because of narrower simulated null distributions under the finite island than under the hierarchical island model (compare Figure 2a with Figure 2c, and Figures 3a and b with Figures 3c and d).
Example of FST distributions obtained under different migration models for single nucleotide polymorphism (SNP) data. The diversity of 1000 SNP loci (open circles) was simulated under a hierarchical island model with 10 groups of 100 demes for two combinations of migration rates within and between groups. The joint null distribution of FST and heterozygosity (30 000 grey dots) was then obtained under either a hierarchical island (a, b) or a finite island model (c, d) based on observed average F-statistics. \(\overline{F}\)ST is the weighed average F-statistics computed over 1000 loci.
For STR data, when groups of populations are well differentiated (FCT=0.2), about 24% of the loci seem to be under balancing selection (too low FST) at the 1% significance level and about 12% of the loci show significantly too high FST. When groups of populations are less differentiated (FCT=0.05), there is still an overall excess of false positives, but this excess is less pronounced than for FCT=0.2. For example, at the 1% level of significance 3.5% of the loci appear under directional selection and 4% under balancing selection. This lower rate of false positives is expected as small values of FCT imply more migrations between groups of demes, which would then tend to behave more like a single migrant pool. However, analyses carried out under the hierarchical island model show false significant rates very close to their expectations for directional selection and slightly overestimated for balancing selection when groups are little differentiated (FCT=0.05).
For SNP data and for the different migration models, we report two analyses in Table 1. The first one was done on all SNPs irrespective of their heterozygosity, and the second one was done only on SNPs with a global scaled heterozygosity larger than 0.2, which was typically about 30% of all 100 000 tested SNPs. FST distributions generated under the finite island model (Figures 3c and d) are generally too narrow and shifted towards lower FST values compared with distributions generated under the hierarchical island model (Figures 3a and b). Under the hierarchical island model, false positive rates are globally slightly underestimated for SNPs under directional selection and quite largely overestimated for SNPs under balancing selection. Note, however, that these levels become correctly estimated when considering common SNPs with Ĥ1>0.2. As expected, we find a large excess of false positives under the finite island model, when groups are well differentiated (FCT=0.2), with about 5% and 15 % of significant SNPs for directional and balancing selection at a 1% level, respectively. When groups are less differentiated (FCT=0.05), false positive rates under the finite island model are relatively well estimated for directional selection, and still considerably overestimated for balancing selection. Note that SNPs with Ĥ1>0.2 show even larger false positive rates when FCT=0.2, which is clearly visible in Figure 3c. When FCT=0.05 and Ĥ1>0.2, the false positive rate for SNPs under balancing selection is correctly estimated, whereas the false positive rate for directional selection is overestimated under the finite island model.
Analysis of human STR data
We first analysed a human data set consisting of 783 autosomal STR loci genotyped in 1048 individuals from 53 Human genome diversity project (HGDP) populations worldwide (Ramachandran et al., 2005), available on http://rosenberglab.bioinformatics.med.umich.edu/diversity.html#data2. The alleles originally coded as fragment lengths were transformed into number of repeats after identification of their motif length, allowing us to estimate unbiased ρ-statistics (Excoffier and Hamilton, 2003). Alleles not fitting a pure SMM were coded as missing data. Five loci for which at least one population contained only missing data after allele recoding were excluded from the analysis (D8S503 in Colombian; ATT015 in Tuscans; AGAT114 in Colombian; AGAT110P in Papuan; and AAAT007 in Colombian), leaving us with 778 loci. Mutation rates were chosen by drawing heterozygosities randomly between 0.2 and 1 and using classical relationships between heterozygosities and the scaled mutation parameter θ=4kdNu (see above).
In line with a previous analysis of 560 of these 783 loci under a Bayesian framework (Foll and Gaggiotti, 2008), which showed that 23% of the loci could be under selection (probability of locus-specific effect >99%), we found a large fraction of STR loci showing signs of selection when data are analyzed under a finite island model (see outlier loci in Figure 4a). Indeed at a 1% level of significance, 91 loci show sign of balancing selection (11.7%) and 74 other loci show sign of directional selection (9.5%). However, when the data are analysed under a hierarchical island model with populations arranged into five continental groups defined by Rosenberg et al. (2002), we found only 8 out of 778 loci (1%) under directional selection and 6 out of 778 loci (0.8%) under balancing selection at the 1% level of significance. These proportions are close to the expected false significance rate of 1%. It suggests that the hierarchical island model can, quite well, reproduce the observed data (Figure 4b) when one takes into account the fact that some sets of populations are quite similar to one another, but also that HGDP STR loci show very little sign of selection, at odds with previous analyses (Foll and Gaggiotti, 2008). Finally, note that a few loci in Figure 4b show very large FST values (that is, FST>0.3) and Ĥ1>1, but are not found significant. We carefully checked the P-value of these loci, and found that they are correctly estimated on the basis of the simulated data points. Thus even though they may visually appear at outliers because the density of simulated points is low for such large Ĥ1 values, there are always more than 1% of the simulated data points that have larger FST values for such high levels of heterozygosity.
Human genome diversity project (HGDP) short term repeat (STR) data. Joint distribution of FST (computed as ρST) against heterozygosity. Black open circles correspond to observed STR loci, whereas grey dots are simulated loci under either (a) finite island model or (b) hierarchical island model. Significant loci (P<0.01) are shown as large black dots in (b).
Analysis of stickleback STR data
The second data set we analysed consisted of 103 STR markers genotyped in four freshwater and three marine populations of sticklebacks (Gasterosteus aculeatus) in northern Europe and in the Balkan (Mäkinen et al., 2008). Original allele definition was recoded to be proportional to the number of repeats in the STR sequence, and one locus was eliminated during this process, due to the presence of imperfect repeats in one population, leaving us with 102 loci to analyse. A Bayesian analysis implemented in the program BayesFST (Beaumont and Balding, 2004) had shown that three STR loci (2.9%) could be under directional selection, whereas about 15 STR loci (14.6%) could be under balancing selection (Mäkinen et al., 2008; see Figure 5a). In line with these results, the re-analysis of the data under a finite island model with FST computed from allele frequencies showed that seven loci could be under directional selection and 17 other loci under balancing selection at 1% level of significance (Figure 5a). This large fraction of loci found under balancing selection was indeed found puzzling. It had been attributed to the high mutation rate at STR loci (Beaumont, 2008), which is not correctly dealt with current methods that only compare STR allele frequencies between populations. Indeed, one can see in Figure 5a that the loci appearing under balancing selection show low FST values associated with an extremely high heterozygosity, which is the same artefact as that described in Figure 2d. To see whether these outliers are removed by taking the SMM model into account, we carried out the same analysis as in Figure 5a, but we computed ρST (Michalakis and Excoffier, 1996) instead of FST. We see in Figure 5b that loci with high heterozygosity can also have high ρST values, many of them being still outlier loci (35 out of 102), without being under a form of balancing selection, having led to low levels of differentiation between populations. We finally carried out an analysis under the hierarchical island model by pooling marine populations into a single group and leaving the remaining four freshwater populations in four different groups. This assumed genetic structured was based on the observation by Mäkinen et al., 2008 that marine sticklebacks showed much lower levels of divergence than freshwater sticklebacks. The result of this hierarchical analysis carried out by assuming the presence of 10 groups of 100 demes is shown on Figure 5c. At 1% level of significance, we find evidence for selection at only seven loci: two loci seem to be under directional selection (STN365 and STN90) in agreement with previous results (Mäkinen et al., 2008); three loci seem under balancing selection (GAest3, GAest61 and the locus GAest60 that was also identified as such in Mäkinen et al., 2008); two remaining outliers (GAest30 and GAest63), which have too low FST for the observed heterozygosity level, are difficult to explain using conventional selection models. Indeed, these last two loci may be outliers because the hierarchical island model has problems in simulating data, which have both high heterozygosity and high FST. Note that the use of a hierarchical island model not only leads to a larger variance of FST for a given heterozygosity level, but also to a right shift of the whole joint distribution. It is, therefore, likely that taking into account additional levels of genetic structure would probably shift the distribution even further to the right and these two loci would not be considered as outliers anymore.
Sticklebacks short term repeat (STR) data. Joint distribution of FST (computed either as FST or ρST) against heterozygosity. Black open circles correspond to observed STR loci, whereas grey dots are simulated loci under either (a and b) the finite island model, or (c) the hierarchical island model. Significant loci at 1% level of significance are shown as large black dots in (c).
Discussion
We show here that testing procedures that do not take into account the hierarchical structure of the populations lead to overly narrow null distributions of FST, and thus to a clear excess of false significant loci. This underestimation of FST variability when populations have a complex history has been realized earlier (Nei and Maruyama, 1975; Robertson, 1975a, 1975b). As clearly pointed out by Robertson (1975a), the variance of FST will indeed not decrease with increasing number of sampled populations if there is a hierarchical structure of the population. This variance will rather depend on the (unknown) correlation structure among all populations. In a hierarchical island model, in which migrations between groups is much smaller than migration within groups, the variance of FST will actually be inversely proportional to the number of sampled groups, rather than to the number of sample demes as in a simple island model. It therefore suggests that increasing the number of samples within groups will not reduce this variability by much. As expected, the false positive rate, incurred by ignoring the hierarchical structure of the populations, decreases with the extent of differentiation between groups of related populations (Table 1). Indeed, in absence of any differentiation between groups, one would get back to a simple island model in which the variance of FST would be proportional to the number of sampled demes. A hierarchical structure of the population may thus explain the large number of loci found to be under selection in some genome scans (for example, Emelianov et al., 2004; Egan et al., 2008; Foll and Gaggiotti, 2008; Namroud et al., 2008). However, our approach based on a simple hierarchical island model considerably reduces the rate of false positives, and this for both directional and balancing selection. This is because the hierarchical island model explicitly allows for the existence of correlations between samples, leading to a much larger variance of the FST distribution (see Figures 2 and 3), and therefore to lower false positive rates. Although recent Bayesian approaches to detect loci under selection (for example, Beaumont and Balding, 2004; Foll and Gaggiotti, 2008) allow for unequal levels of differentiation among populations, they seem unable to cope with the presence of hierarchical structure. Indeed, both the HGDP and the stickleback STR data sets have been analysed under Bayesian approaches (Beaumont and Balding, 2004; Foll and Gaggiotti, 2008; Mäkinen et al., 2008), but found to have a proportion of sites under balancing or directional selection that is very similar to what is inferred under the finite island IAM model (see Results section). This is in keeping with an earlier observation (Beaumont and Balding, 2004) that BayesFST approach gave results very similar to the FDIST procedure (Beaumont and Nichols, 1996), which is actually completely analogous to the finite island IAM approach described here.
Importance of mutation model for STR data
Whichever the migration model, it also seems important to correctly account for the right mutation model to get correct P-values and to properly interpret them. For instance, in a pure island model without recurrent bottlenecks, loci with high mutation rates and recurrent mutations (like STRs) cannot show large levels of population differentiation based on the comparison of allele frequencies (Hedrick, 2005), and the relationship between heterozygosity (taken as a proxy for mutation rate) and FST is non-monotonous (Figures 2b and d). Therefore, in this case STR loci with very large mutation rates will often show low FST, and if they are found significant they could be incorrectly interpreted as being subject to balancing selection (Beaumont, 2008). Note that this interpretation problem does not occur when one computes ρST instead of FST because highly mutable loci can show both large heterozygosity and large ρST (see Figure 2a). Thus, although it has been advocated that the use of FST should be preferred over that of ρST for estimating migration rates (for example, Gaggiotti et al., 1999), we find here that the use of ρST seems more appropriate than FST when detecting STR loci under selection because ρST is essentially unbiased (Slatkin, 1995, and Figure 2), and because the testing procedure based on coalescent simulations adequately handles ρST variability across loci.
Uncertainty about population structure
We assume here that the genetic structure of the population is known in advance, but when this is not the case several approaches could be used to define this genetic structure from the data (for example, Pritchard et al., 2000; Dupanloup et al., 2002; Corander et al., 2004; Guillot et al., 2005). Once the genetic structure has been identified, one also needs to define how many groups of population (k) and how many demes per group (d) to simulate. To see the effect of this choice on estimated P-values, we report in Table 2 the determination coefficient (R2) between P-values obtained from simulations under the correct hierarchical island model (k=10, d=100) and those obtained by simulating data under different models (k={1, 5, 10, 20} and d={20, 50, 100, 200}). As expected, R2 values are much lower for a non-hierarchical (k=1, R2=0.15–0.21) than for a hierarchical analysis (k=5, 10 or 20, R2=0.87–0.99). We see that the P-values obtained under the correct model are highly correlated among runs (R2=0.99), but that this correlation decreases quite sharply when the number of demes within group is small (d=20, R2=0.87–0.93). When the number of demes per group becomes larger (that is, d⩾50), the P-values become very close to those obtained under the correct settings (for example, d=50, R2=0.95–0.98), and R2 values are also very high when the number of simulated demes is much larger than that used to generate the data (when d=200, R2=0.96–0.99). It suggests that the use of a hierarchical island model with a relatively large number of groups (that is, at least twice as many groups than reported in the genetic structure) and a large number of demes per group (d⩾100) should give consistent and reliable results. In any case, as the analysis of thousands of loci with 50 000 simulations usually takes only a few hours, replicated analyses using different values of k and d could be done to assess their influence on the results of specific data sets.
To see the effect of an alternative population structure and evolutionary process on the testing procedure, we also simulated 10 data sets of 1000 STR loci under a spatially explicit model using the SPLATCHE software (Currat et al., 2004). We used the human example as a template and we simulated a population expansion on a world map under a two dimensional stepping-stone model (see Supplementary Figure S1). The spatial expansion was assumed to have occurred some 40 000 generations ago from Ethiopia (Prugnolle et al., 2005; Ray et al., 2005). The parameters of the model, such as mutation rate (μ=5 × 10−4), deme carrying capacity (K=500), level of gene flow between neighbouring demes (m=0.1) and level of local population growth (r=0.8) were chosen to approximately lead to the observed global level of differentiation between human population as ρST ≅0.11 (Excoffier and Hamilton, 2003). Using a coalescent-based approach implemented in SPLATCHE, we simulated genetic data at the same geographic coordinates (see Supplementary Figure S1) and using the same sample size as those defined for the 53 populations of HGDP (Cann et al., 2002). On the basis of the analysis of 10 data sets of 1000 STR loci thus generated, we found that the use of a hierarchical island model partitioning samples into five continental groups (as in Figure 4) markedly reduces the rate of false positives as compared with the use of a finite island model (Table 3), but we still observe about a two times excess of significant loci under the hierarchical island model. Note that results under the hierarchical island model do not seem to depend much on the number of assumed groups in this case, as we obtain extremely similar false positive rates when simulating 5 or 10 groups of demes (Table 3). Although it is difficult to generalize this analysis to all possible scenarios of range expansions, it suggests that the distribution of FST under a spatial expansion in a stepping-stone model cannot be perfectly replicated in a hierarchical island model.
Effect of using a wrong genetic structure
Although we have assumed that population samples were correctly assigned to different groups, the genetic structure could sometimes be mis-estimated by users. To see the effect of incorrect population assignments to groups, we simulated genetic diversity in 10 populations arranged in various numbers of groups (k={1, 2, 5, or 8}) and analyzed the data set by allocating populations to similar or to different number of groups (in this case g={1, 2, 5, or 8}). In all cases, FCT between groups was set to 0.2 and FST between demes to 0.24. We report the results in Table 4, in which we see that the proportion of false positives is clearly overestimated when k≠g. When g>k, which corresponds to the splitting of some real groups of population, the false positive rate generally decreases when g increases. When g<k, which corresponds to the lumping of some distinct groups of population in the same group, the false positive rate also decreases with increasing g, as g approaches closer to k. Even though we have studied a very limited number of cases, it seems that the splitting of existing groups is less detrimental than the lumping of those groups, which produces false positives at larger rates. It is also interesting to note that the false positive rate is largest when one assumes no hierarchical genetic structure when there is one. In other words, it seems always better to assume a hierarchical structure than ignoring this hierarchical structure, even if the true hierarchical genetic structure is not correctly inferred. We also note that the largest false positive rates were obtained when data resulting from the simulation of two groups of demes were analysed with an incorrect genetic structure. This result makes sense in light of the prediction of Robertson (1975a), which suggests that the variance in FST in a hierarchically structured populations increases with the correlation in gene frequencies between demes, which it is thus largest in our hierarchical island model when k=2.
Genes under selection in humans
The application of our hierarchical analysis on human and stickleback data sets leads to a considerably lower number of significant loci than previously published (Foll and Gaggiotti, 2008; Mäkinen et al., 2008), and this number only slightly exceeds that expected by chance. We attribute this result to our use of more realistic mutation and demographic models. In the case of humans, it is interesting to note that the distribution of observed FST seems to fit very well to the hierarchical island model, even though it has been proposed that human genetic diversity could be adequately explained by a simple range expansion out of Africa (Prugnolle et al., 2005; Ramachandran et al., 2005). We would have indeed expected to find a larger proportion of significant loci if observed diversity at STR loci would have been generated under a pure range expansion model (see Table 3). It suggests that human history has been more complex than a simple range expansion out of Africa, with periods of isolation during ice ages and possible long-distance migrations between continents (Foley and Lahr, 2001). However, even if selection had been affecting genetic diversity at many loci it is likely that genome scan with ‘only’ a few hundred markers evenly distributed over the genome would not be able to detect them all, as the effect of selection in subdivided populations is usually restricted to relatively small segments around the selected loci (where r/s≪1, see for example, Charlesworth et al., 1997; Slatkin and Wiehe, 1998). There are actually several examples of loci having recently responded to selection, where allele frequency differences between populations are restricted to a few kb around the selected site (see for example, Turner and Hahn, 2007; Seehausen et al., 2008; Wood et al., 2008). Among the 783 human STR loci analysed here, 488 STRs could be mapped on the genome. Among these, a relatively large proportion (162 out of 488) are found within gene transcripts and 210 are within 100 kb of a nearby known gene (Hofer et al., 2008). Interestingly, among the eight loci potentially, found here, under directional selection, four are found within transcripts and one other is less than 1 kb away from any known gene. Among the six loci potentially under balancing selection, two are within transcripts and one is only 3 kb away from a gene (see Table 5). STRs within or close to gene transcripts are thus good candidates for having been influenced by selection. Note that only two of the genes reported in Table 5 have been reported to be under selection in previous studies: PHACTR1 (Williamson et al., 2007; Foll and Gaggiotti, 2008) and E2F6 (Kayser et al., 2003). Using a non-hierarchical Bayesian approach, Foll and Gaggiotti (2008) also found that six of the 14 outliers loci were under selection, but they did not find evidence for selection at three other loci (Table 5), even though one of these is found within a gene transcript (MAGI2). This discrepancy is in line with the low correlation in P-values found between the finite island and the hierarchical island model (Table 3), and suggests that our approach can also detect new genes under selection. Among the biological processes in which our 14 outlier genes are involved, we only find an enrichment for neuronal activities (P=0.05 after Bonferroni correction), a category that was already shown to be enriched among selected genes in previous studies (see for example, Wang et al., 2006; Haygood et al., 2007).
Possible applications and limitations of hierarchical analyses
Although adaptations in a given portion of a species range should globally lead to increased levels of differentiation (and hence global FST), the observation of a locus with an elevated FST value does not necessarily imply that an adaptive event occurred. Indeed, allele surfing during range expansions (Klopfstein et al., 2006) can also lead to high levels of differentiation between populations in the periphery of the range at a random locus (Hallatschek et al., 2007; Excoffier and Ray, 2008). A mutation linked to dispersal and affecting migration rates or a mutation influencing mate choice could also alter FST levels through their effects on local inbreeding and effective population size. Markers located on chromosomes with different ploidy levels (for example, cpDNA, mtDNA, X and Y chromosome) should also have a distinct effective size (see for example, Pool and Nielsen, 2007) potentially leading to more extreme FST, and they would thus need to be analysed separately.
It seems therefore important to be able to distinguish among these different situations, and new methods aiming at specifically finding loci showing geographically restricted differentiation indicative of local adaptive selection would be desirable. A hierarchical approach seems well suited for this purpose. For instance, one may be interested in finding loci that are particularly differentiated between two geographic regions, two species or sub-species, or between two groups of population showing distinct phenotypes or ecotypes (Campbell and Bernatchez, 2004; Kane and Rieseberg, 2007; Egan et al., 2008; Nosil et al., 2008). A nested analysis of variance under a hierarchical island model would allow one to use replicated population samples within each of the group to be contrasted, which should be more powerful than an approach based on population pairwise comparisons, and solve the problem of non-independence in multiple pairwise tests occurring in many studies (for example, Schlotterer, 2002; Bonin et al., 2006; Kane and Rieseberg, 2007; Nosil et al., 2008; Oetjen and Reusch, 2007).
Although the hierarchical island model can certainly reduce the number of false positives arising in studies involving related populations samples, it can still lead to an excess of false positives if the underlying genetic structure is more complex than what we have modelled, for instance in case of complex demographic histories, involving population splits, range expansions (Table 3), bottlenecks or admixture events. However, the present approach seems quite conservative and always leads to lesser false positives than when assuming that populations are independent (Table 4). As current genome scans aiming at detecting loci under selection are now typically carried out on hundreds of thousands of markers (see for example, Pickrell et al., 2009), it still seems worthwhile trying to reduce the number of candidate loci to a few dozen or a few hundreds that could then be further studied, perhaps by examining their genomic distribution and identifying genes in their neighbourhood. Further research on better ways to take into account the true underlying genetic structure of the populations is needed and should lead to more powerful procedure to detect loci under selection.
References
Balding DJ (2003). Likelihood-based inference for genetic correlation coefficients. Theor Popul Biol 63: 221–230.
Balding DJ, Nichols RA (1995). A method for quantifying differentiation between populations at multi-allelic loci and its implications for investigating identity and paternity. Genetica 96: 3–12.
Beaumont MA (2005). Adaptation and speciation: What can F-st tell us? Trends Ecol Evol 20: 435–440.
Beaumont MA (2008). Selection and sticklebacks. Mol Ecol 17: 3425–3427.
Beaumont MA, Balding DJ (2004). Identifying adaptive genetic divergence among populations from genome scans. Mol Ecol 13: 969–980.
Beaumont MA, Nichols RA (1996). Evaluating loci for use in the genetic analysis of population structure. Proc R Soc Lond B 263: 1619–1626.
Beaumont MA, Zhang W, Balding DJ (2002). Approximate Bayesian computation in population genetics. Genetics 162: 2025–2035.
Bonin A, Taberlet P, Miaud C, Pompanon F (2006). Explorative genome scan to detect candidate loci for adaptation along a gradient of altitude in the common frog (Rana temporaria). Mol Biol Evol 23: 773–783.
Braverman JM, Hudson RR, Kaplan NL, Langley CH, Stephan W (1995). The hitchhiking effect on the site frequency spectrum of DNA polymorphisms. Genetics 140: 783–796.
Campbell D, Bernatchez L (2004). Generic scan using AFLP markers as a means to assess the role of directional selection in the divergence of sympatric whitefish ecotypes. Mol Biol Evol 21: 945–956.
Cann HM, de Toma C, Cazes L, Legrand M-F, Morel V, Piouffre L et al. (2002). A human genome diversity cell line panel. Science 296: 261b–2262.
Cavalli-Sforza LL (1966). Population structure and human evolution. Proc R Soc Lond B Biol Sci 164: 362–379.
Charlesworth B, Nordborg M, Charlesworth D (1997). The effects of local selection, balanced polymorphism and background selection on equilibrium patterns of genetic diversity in subdivided populations. Genet Res 70: 155–174.
Corander J, Waldmann P, Marttinen P, Sillanpaa MJ (2004). BAPS 2: Enhanced possibilities for the analysis of genetic population structure. Bioinformatics 20: 2363–2369.
Currat M, Ray N, Excoffier L (2004). SPLATCHE: A program to simulate genetic diversity taking into account environmental heterogeneity. Mol Ecol Notes 4: 139–142.
Dupanloup I, Schneider S, Excoffier L (2002). A simulated annealing approach to define the genetic structure of populations. Mol Ecol 11: 2571–2581.
Egan SP, Nosil P, Funk DJ (2008). Selection and genomic differentiation during ecological speciation: Isolating the contributions of host association via a comparative genome scan of Neochlamisus bebbianae leaf beetles. Evolution 62: 1162–1181.
Emelianov I, Marec F, Mallet J (2004). Genomic evidence for divergence with gene flow in host races of the larch budmoth. Proc Biol Sci 271: 97–105.
Excoffier L, Hamilton G (2003). Comment on ‘Genetic Structure of Human Populations’. Science 300: 1877b–1877.
Excoffier L, Laval G, Schneider S (2005). Arlequin (version 3.0): An integrated software package for population genetics data analysis. Evol Bioinform Online 1: 47–50.
Excoffier L, Ray N (2008). Surfing during population expansions promotes genetic revolutions and structuration. Trends Ecol Evol 23: 347–351.
Excoffier L, Smouse P, Quattro J (1992). Analysis of molecular variance inferred from metric distances among DNA haplotypes: Application to human mitochondrial DNA restriction data. Genetics 131: 479–491.
Fay JC, Wu CI (2000). Hitchhiking under positive Darwinian selection. Genetics 155: 1405–1413.
Foley RA, Lahr MM (2001). The anthropological, demographic and ecological context of human evolutionary genetics. In: Donnelly P and Foley RA (eds). Genes, Fossils and Behavior. IOS Press: Amsterdam. Vol. 310, pp 223–245.
Foll M, Gaggiotti OE (2008). A genome scan method to identify selected loci appropriate for both dominant and codominant markers: A Bayesian perspective. Genetics 180: 977–993.
Fu Y-X (1997). Statistical tests of neutrality of mutations against population growth, hitchhiking and background selection. Genetics 147: 915–925.
Gaggiotti OE, Lange O, Rassmann K, Gliddon C (1999). A comparison of two indirect methods for estimating average levels of gene flow using microsatellite data. Mol Ecol 8: 1513–1520.
Gillespie JH (1990). The molecular nature of allelic diversity for two models of balancing selection. Theor Popul Biol 37: 91–109.
Guillot G, Estoup A, Mortier F, Cosson JF (2005). A spatial statistical model for landscape genetics. Genetics 170: 1261–1280.
Hallatschek O, Hersen P, Ramanathan S, Nelson DR (2007). Genetic drift at expanding frontiers promotes gene segregation. Proc Natl Acad Sci USA 104: 19926–19930.
Haygood R, Fedrigo O, Hanson B, Yokoyama KD, Wray GA (2007). Promoter regions of many neural- and nutrition-related genes have experienced positive selection during human evolution. Nat Genet 39: 1140–1144.
Hedrick PW (2005). A standardized genetic differentiation measure. Evolution 59: 1633–1638.
Hofer T, Ray N, Wegmann D, Excoffier L (2008). Large allele frequency differences between human continental groups are more likely to have occurred by drift during range expansions than by selection. Ann Hum Genet 73: 95–108.
Kane NC, Rieseberg LH (2007). Selective sweeps reveal candidate genes for adaptation to drought and salt tolerance in common sunflower, Helianthus annuus. Genetics 175: 1823–1834.
Kaplan NL, Hudson RR, Langley CH (1989). The ‘hitchhiking effect’ revisited. Genetics 123: 887–899.
Kayser M, Brauer S, Stoneking M (2003). A genome scan to detect candidate regions influenced by local natural selection in human populations. Mol Biol Evol 20: 893–900.
Kim Y, Stephan W (2002). Detecting a local signature of genetic hitchhiking along a recombining chromosome. Genetics 160: 765–777.
Klopfstein S, Currat M, Excoffier L (2006). The fate of mutations surfing on the wave of a range expansion. Mol Biol Evol 23: 482–490.
Kreitman M, Hudson RR (1991). Inferring the evolutionary histories of the Adh and Adh-dup loci in Drosophila melanogaster from patterns of polymorphism and divergence. Genetics 127: 565–582.
Lewontin RC, Krakauer J (1973). Distribution of gene frequency as a test of the theory of the selective neutrality of polymorphisms. Genetics 74: 175–195.
Lewontin RC, Krakauer J (1975). Letters to the editors: testing the heterogeneity of F values. Genetics 80: 397–398.
Mäkinen HS, Cano JM, Merila J (2008). Identifying footprints of directional and balancing selection in marine and freshwater three-spined stickleback (Gasterosteus aculeatus) populations. Mol Ecol 17: 3565–3582.
McVean G (2007). The structure of linkage disequilibrium around a selective sweep. Genetics 175: 1395–1406.
Michalakis Y, Excoffier L (1996). A generic estimation of population subdivision using distances between alleles with special reference for microsatellite loci. Genetics 142: 1061–1064.
Namroud MC, Beaulieu J, Juge N, Laroche J, Bousquet J (2008). Scanning the genome for gene single nucleotide polymorphisms involved in adaptive population differentiation in white spruce. Mol Ecol 17: 3599–3613.
Nei M (1973). Analysis of gene diversity in subdivided populations. Proc Natl Acad Sci USA 70: 3321–3323.
Nei M, Maruyama T (1975). Letters to the editors: Lewontin–Krakauer test for neutral genes. Genetics 80: 395.
Nielsen R, Hellmann I, Hubisz M, Bustamante C, Clark AG (2007). Recent and ongoing selection in the human genome. Nat Rev Genet 8: 857–868.
Nielsen R, Williamson S, Kim Y, Hubisz MJ, Clark AG, Bustamante C (2005). Genomic scans for selective sweeps using SNP data. Genome Res 15: 1566–1575.
Nordborg M (1997). Structured coalescent processes on different time scales. Genetics 146: 1501–1514.
Nosil P, Egan SP, Funk DJ (2008). Heterogeneous genomic differentiation between walking-stick ecotypes: ‘Isolation by adaptation’ and multiple roles for divergent selection. Evolution 62: 316–336.
Notohara M (1990). The coalescent and the genealogical process in geographically structured population. J Math Biol 29: 59–75.
Oetjen K, Reusch TB (2007). Genome scans detect consistent divergent selection among subtidal vs. intertidal populations of the marine angiosperm Zostera marina. Mol Ecol 16: 5156–5167.
Ohta T, Kimura M (1973). A model of mutation appropriate to estimate the number of electrophoretically detectable alleles in a finite population. Genet Res 22: 201–204.
Pickrell JK, Coop G, Novembre J, Kudaravalli S, Li JZ, Absher D et al. (2009). Signals of recent positive selection in a worldwide sample of human populations. Genome Res 19: 826–837.
Pool JE, Nielsen R (2007). Population size changes reshape genomic patterns of diversity. Evolution 61: 3001–3006.
Pritchard JK, Stephens M, Donnelly P (2000). Inference of population structure using multilocus genotype data. Genetics 155: 945–959.
Prugnolle F, Manica A, Balloux F (2005). Geography predicts neutral genetic diversity of human populations. Curr Biol 15: R159–R160.
Przeworski M (2002). The signature of positive selection at randomly chosen loci. Genetics 160: 1179–1189.
Ramachandran S, Deshpande O, Roseman CC, Rosenberg NA, Feldman MW, Cavalli-Sforza LL (2005). Support from the relationship of genetic and geographic distance in human populations for a serial founder effect originating in Africa. Proc Natl Acad Sci USA 102: 15942–15947.
Rannala B, Hartigan JA (1996). Estimating gene flow in island populations. Genet Res 67: 147–158.
Ray N, Currat M, Berthier P, Excoffier L (2005). Recovering the geographic origin of early modern humans by realistic and spatially explicit simulations. Genome Res 15: 1161–1167.
Riebler A, Held L, Stephan W (2008). Bayesian variable selection for detecting adaptive genomic differences among populations. Genetics 178: 1817–1829.
Robertson A (1975a). Gene frequency distributions as a test of selective neutrality. Genetics 81: 775–785.
Robertson A (1975b). Letters to the editors: remarks on the Lewontin–Krakauer test. Genetics 80: 396.
Rosenberg NA, Pritchard JK, Weber JL, Cann HM, Kidd KK, Zhivotovsky LA et al. (2002). Genetic structure of human populations. Science 298: 2381–2385.
Sabeti PC, Reich DE, Higgins JM, Levine HZ, Richter DJ, Schaffner SF et al. (2002). Detecting recent positive selection in the human genome from haplotype structure. Nature 419: 832–837.
Schlotterer C (2002). A microsatellite-based multilocus screen for the identification of local selective sweeps. Genetics 160: 753–763.
Seehausen O, Terai Y, Magalhaes IS, Carleton KL, Mrosso HD, Miyagi R et al. (2008). Speciation through sensory drive in cichlid fish. Nature 455: 620–626.
Slatkin M (1991). Inbreeding coefficients and coalescence times. Genet Res 58: 167–175.
Slatkin M (1995). A measure of population subdivision based on microsatellite allele frequencies. Genetics 139: 457–462.
Slatkin M, Voelm L (1991). FST in a hierarchical island model. Genetics 127: 627–629.
Slatkin M, Wiehe T (1998). Genetic hitch-hiking in a subdivided population. Genet Res 71: 155–160.
Storz JF (2005). Using genome scans of DNA polymorphism to infer adaptive population divergence. Mol Ecol 14: 671–688.
Storz JF, Payseur BA, Nachman MW (2004). Genome scans of DNA variability in humans reveal evidence for selective sweeps outside of Africa. Mol Biol Evol 21: 1800–1811.
Strobeck K (1987). Average number of nucleotide differences in a sample from a single subpopulation: A test for population subdivision. Genetics 117: 149–153.
Tajima F (1989). Statistical method for testing the neutral mutation hypothesis by DNA polymorphism. Genetics 123: 585–595.
Tang K, Thornton KR, Stoneking M (2007). A new approach for using genome scans to detect recent positive selection in the human genome. PLoS Biol 5: e171.
Teshima KM, Coop G, Przeworski M (2006). How reliable are empirical genomic scans for selective sweeps? Genome Res 16: 702–712.
Thomas PD, Campbell MJ, Kejariwal A, Mi H, Karlak B, Daverman R et al. (2003). PANTHER: A library of protein families and subfamilies indexed by function. Genome Res 13: 2129–2141.
Thornton KR, Jensen JD (2007). Controlling the false-positive rate in multilocus genome scans for selection. Genetics 175: 737–750.
Tsakas S, Krimbas CB (1976). Testing the heterogeneity of F values: A suggestion and a correction. Genetics 84: 399–401.
Turner TL, Hahn MW (2007). Locus- and population-specific selection and differentiation between incipient species of Anopheles gambiae. Mol Biol Evol 24: 2132–2138.
Vitalis R, Dawson K, Boursot P (2001). Interpretation of variation across marker loci as evidence of selection. Genetics 158: 1811–1823.
Voight BF, Kudaravalli S, Wen X, Pritchard JK (2006). A map of recent positive selection in the human genome. PLoS Biol 4: e72.
Wakeley J, Aliacar N (2001). Gene genealogies in a metapopulation. Genetics 159: 893–905.
Wang ET, Kodama G, Baldi P, Moyzis RK (2006). Global landscape of recent inferred Darwinian selection for Homo sapiens. Proc Natl Acad Sci USA 103: 135–140.
Weir BS, Cockerham CC (1984). Estimating F-statistics for the analysis of population structure. Evolution 38: 1358–1370.
Williamson SH, Hubisz MJ, Clark AG, Payseur BA, Bustamante CD, Nielsen R (2007). Localizing recent adaptive evolution in the human genome. PLoS Genet 3: e90.
Wood HM, Grahame JW, Humphray S, Rogers J, Butlin RK (2008). Sequence differentiation in regions identified by a genome scan for local adaptation. Mol Ecol 17: 3123–3135.
Wright S (1931). Evolution in Mendelian populations. Genetics 16: 97–159.
Wright S (1951). The genetical structure of populations. Ann Eugen 15: 323–354.
Zeng K, Fu YX, Shi S, Wu CI (2006). Statistical tests for detecting positive selection by utilizing high-frequency variants. Genetics 174: 1431–1439.
Acknowledgements
We are grateful to Hannu Mäkinen for his comments on a previous version of the paper and making his stickleback data set available to us, as well as to two anonymous reviewers for their useful comments. We are also indebted to Mark Beaumont for detailed comments and useful suggestions. The methodology presented in this paper has been incorporated in the Arlequin package and will be made available in a future release. This work was supported by a Swiss NSF Grant No. 3100A0–112072 to LE.
Author information
Authors and Affiliations
Corresponding author
Additional information
Supplementary Information accompanies the paper on Heredity website (http://www.nature.com/hdy)
Supplementary information
Appendix
Appendix
As shown previously (Slatkin, 1991; Slatkin and Voelm, 1991), Wright's F-statistics can be expressed as a function of mean coalescence times of two genes drawn in the same deme (t̄0), in two different demes from the same group (t̄1), and in two demes from different groups (t̄2) as,

The average coalescence time of two genes drawn from the same deme in a subdivided population is the total number of genes in the population (Strobeck, 1987). If we assume that there are k groups of demes, each one being made up of d demes, and that deme size is N, then

The average coalescence time of genes drawn from different demes can be divided into two phases. To coalesce, two genes need to be in the same group, which will take a time t̄1′ for two genes in different demes but in the same group, and a time t̄2′ for two genes found in different groups. Once they are in the same deme, it will take them t̄0 more generations to coalesce, such that t̄i=t̄i′+t̄0 (i=1, 2) (Slatkin and Voelm, 1991). Assuming that m1≫m2, Slatkin and Voelm (1991) have derived the t̄i′s as

Combining equations (A5), (A6) and (A7), Wright's F-statistics can therefore be written as



showing that in a hierarchical context, FST obtained in (A6) is approximately equal to a non-hierarchical FST that is computed among groups, ignoring subdivisions within group.
Rights and permissions
About this article
Cite this article
Excoffier, L., Hofer, T. & Foll, M. Detecting loci under selection in a hierarchically structured population. Heredity 103, 285–298 (2009). https://doi.org/10.1038/hdy.2009.74
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/hdy.2009.74
Keywords
This article is cited by
-
Conservation implications for the world’s most widely distributed cavefish species complex based on population genomics (Typhlichthys, Percopsiformes)
Conservation Genetics (2024)
-
Genome Scan of Rice Landrace Populations Collected Across Time Revealed Climate Changes’ Selective Footprints in the Genes Network Regulating Flowering Time
Rice (2023)
-
Key triggers of adaptive genetic variability of sessile oak [Q. petraea (Matt.) Liebl.] from the Balkan refugia: outlier detection and association of SNP loci from ddRAD-seq data
Heredity (2023)
-
Global invasion history of the emerging plant pathogen Phytophthora multivora
BMC Genomics (2022)
-
Physical geography, isolation by distance and environmental variables shape genomic variation of wild barley (Hordeum vulgare L. ssp. spontaneum) in the Southern Levant
Heredity (2022)