Abstract
Background:
Glutathione S-transferase 1 (GSTP1) inactivation is associated with CpG island promoter hypermethylation in the majority of prostate cancers (PCs). This study assessed whether the level of circulating methylated GSTP1 (mGSTP1) in plasma DNA is associated with chemotherapy response and overall survival (OS).
Methods:
Plasma samples were collected prospectively from a Phase I exploratory cohort of 75 men with castrate-resistant PC (CRPC) and a Phase II independent validation cohort (n=51). mGSTP1 levels in free DNA were measured using a sensitive methylation-specific PCR assay.
Results:
The Phase I cohort identified that detectable baseline mGSTP1 DNA was associated with poorer OS (HR, 4.2 95% CI 2.1–8.2; P<0.0001). A decrease in mGSTP1 DNA levels after cycle 1 was associated with a PSA response (P=0.008). In the Phase II cohort, baseline mGSTP1 DNA was a stronger predictor of OS than PSA change after 3 months (P=0.02). Undetectable plasma mGSTP1 after one cycle of chemotherapy was associated with PSA response (P=0.007).
Conclusions:
We identified plasma mGSTP1 DNA as a potential prognostic marker in men with CRPC as well as a potential surrogate therapeutic efficacy marker for chemotherapy and corroborated these findings in an independent Phase II cohort. Prospective Phase III assessment of mGSTP1 levels in plasma DNA is now warranted.
Similar content being viewed by others
Main
Prostate cancer (PC) remains the third most common cause of cancer death in men in the developed world (Jemal et al, 2009). Docetaxel chemotherapy offers both symptomatic and survival benefits in men with metastatic castrate-resistant PC (mCRPC), however, only 48–50% of men treated with Docetaxel benefit from treatment (Petrylak et al, 2004; Tannock et al, 2004). A high proportion of patients experience significant toxicity; 42% nausea, vomiting or both, 30% sensory neuropathy and 26% more than or equal to one serious adverse event (Petrylak et al, 2004; Tannock et al, 2004). The newer second-line cytotoxic, Cabazitaxel, similarly improves survival, but remains toxic and benefits <50% of patients (de Bono et al, 2010). Clearly, patients would benefit from knowing early in the treatment schedule, who will benefit from chemotherapy to avoid the morbidity in an elderly population.
Glutathione S-transferases (GSTs) are a family of enzymes that catalyse intracellular detoxification of a variety of electrophiles, including a number of xenobiotics and carcinogens (Rushmore and Pickett, 1993). Human GSTs are classified into distinct families; five of the gene families encode cytosolic GSTs, (alpha, mu, pi, sigma and theta), whereas single genes encode membrane-bound forms of the enzyme (Hayes and Pulford, 1995). Among the isozymes, the pi class enzyme Glutathione S-transferase 1 (GSTP1) is the most widely distributed (Ruzza et al, 2009) and many groups including our own have shown that GSTP1 expression is inactivated in >90% of all PCs(Lee et al, 1994; Millar et al, 1999) in association with aberrant DNA methylation of its CpG island associated promoter and first exon (Millar et al, 1999). PC can be diagnosed by the detection of methylated GSTP1 DNA (mGSTP1) in urine and semen samples (Bryzgunova et al, 2008). Detectable mGSTP1 in the pre-operative serum of men with localised PC is associated with biochemical relapse (Bastian et al, 2005), whereas detectable serum mGSTP1 is present in 28–32% of men with metastatic PC (Bastian et al, 2005; Reibenwein et al, 2007). However, there are no longitudinal data assessing the changes in levels during the progression and treatment of individual patients (Bastian et al, 2005).
Although the level of circulating tumour cells (CTCs) is prognostic and associated with chemoresponse in CRPC, quantitation is cumbersome and prone to technical challenges including cell lysis (Danila et al, 2007a; de Bono et al, 2008; Scher et al, 2009a). Detection of methylated GSTP1 in free DNA from plasma provides a stable marker to identify tumour DNA in the circulation and offers the potential to assess response to therapy. This study aimed to determine whether quantitative measurement of plasma mGSTP1 free DNA is associated with response to chemotherapy and overall survival (OS) in men with mCRPC.
Materials and methods
This study was reported in accordance with REMARK guidelines (McShane et al, 2005).
Patient population and plasma samples
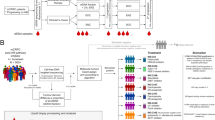
The phase I exploratory cohort (Figure 1A) consisted of 75 men with mCRPC, who had blood samples obtained at the time of presentation to Medical Oncology (Royal Prince Alfred Hospital, Concord Hospital and Royal North Shore Hospital) between June 2006 and July 2009. Patients were seen every 6–12 weeks and had blood samples obtained each visit. Men who subsequently had chemotherapy (Docetaxel 75 mg m−2 3-weekly or Mitoxantrone 12 mg m−2 3-weekly) had blood samples drawn prior to each cycle of treatment. The phase II independent validation cohort (Figure 1B) consisted of 51 men who were receiving chemotherapy (Docetaxel 75 mg m−2 3-weekly, Cabazitaxel 25 mg m−2 3-weekly or Mitoxantrone 12 mg m−2 3-weekly) for mCRPC, who had blood samples obtained prior to commencing chemotherapy and just prior to their second cycle of treatment and were enrolled between April 2009 and September 2012 from hospitals in the PRIMe (Pharmacogenomic Research for Individualized Medicine) network: Sydney Cancer Centre (Royal Prince Alfred Hospital and Concord Hospital), the SAN Clinic, Westmead Hospital and the Mater Misericorde Newcastle Hospital. Blood samples were collected according to a standardised operating procedure using 0.109 M buffered sodium citrate for the CTC preparation, BD Vacutainer tubes containing K2EDTA for plasma samples and BD Vacutainer tubes containing clot activator and gel separation for serum samples. For the Phase I exploratory cohort, 7.5 ml of whole blood was also obtained for each assay and free DNA was collected using the QIAamp MinElute Virus Spin Kit, according to the manufacturer’s directions. For the Phase II validation cohort, 1 ml of plasma was assayed for each time point. Samples were centrifuged and couriered at 4 °C to a central laboratory (Garvan Institute of Medical Research). Within 12 h of collection, CTCs were analysed and plasma was stored at −80 °C.
PSA partial response (PR) was defined as a 50% or greater decrease in serum PSA compared with the last value assessed prior to initiation of chemotherapy (Petrylak et al, 2004; Tannock et al, 2004). PSA progressive disease (PD) was defined as an increase in PSA value by >25% over the baseline, and stable disease (SD) was defined as a value >0.5 but <1.25 times the baseline value (Petrylak et al, 2004; Tannock et al, 2004). Overall survival was calculated from the date of the baseline biomarker blood draw to the date of death or last follow-up. Eastern Cooperative Oncology Group performance status (ECOG PS) assessment, serum haemoglobin (Hb) and serum alkaline phosphatase (ALP) data were collected prospectively, but not available for all participants. Radiological assessments were according to standard practice, were variable and unsuitable for determining objective tumour response.
All patients provided written informed consent, and the study was approved by human research ethics review committees at all participating institutions, and was registered on the Australian New Zealand Clinical Trials Registry.
Methylation-specific head-loop PCR (MS-HL PCR) GSTP1 assay
Bisulphite treatment was carried out on all plasma-free DNA samples for 4 h (Zymo EZ-96 DNA Methylation-Gold Kit #D5008). The GSTP1 methylation-sensitive headloop (MS-HL) PCR assay was performed as previously described to measure the absolute amount of methylated GSTP1 DNA in the DNA samples (Rand et al, 2002). Forward primer: 5′ACACAACCCACATCCCCAAAATGTTGGGAGTTTTGAGTTTTATTTT; Reverse primer: 5′ AAAACCICIAAACCTTCICTAAAATTTC; Probe: 5′ VIC-TCG CCG CCG CAA T-MGBNFQ. The control GSTP1 PCR reaction was used to measure the amount of total DNA in the sample. Forward primer: 5′GGG ATT ATT TTT ATA AGG TTY GGA GGT; Reverse primer: 5′ AAA ACC CRA ACC TAA TAC TAC RAA TTA A; Probe 5′ FAM-CCC CAT ACT AAA AAC TCT AAA CCC CAT CCC. PCR reaction conditions: 95 °C for 120 s, 50 cycles at 95 °C for 15 s, 60 °C for 60 s. Real-time PCR was carried out in triplicate using an ABI PRISM ABI7900 Sequence Detection System. The average triplicate Ct values from the GSTP1 MS-HL PCR assay were used to estimate the quantity of methylated DNA in the plasma and serum DNA samples using the standard curve (Supplementary Figure 1), generated using known (1 ng, 5 ng, 10 ng, 50 ng, 100 ng, 500 ng) concentrations of SssI fully methylated DNA into unmethylated DNA, as described previously(Rand et al, 2002). Detectable mGSTP1 was defined as >1 ng per plasma sample. These assays were performed blinded to the clinical outcomes of the patients.
Statistical analysis
Plasma and serum levels of mGSTP1 were not normally distributed, so we used McNemar’s test and linear regression after log-transformation to determine relationships between the different DNA assays. Reported correlations are based on regression of the log-transformed values. ANOVA was used to assess the relationship between plasma mGSTP1 levels, the sites of metastases and Gleason score. The relationship between plasma mGSTP1 levels (detectable vs undetectable) and PSA response to chemotherapy (PR and SD vs PD) was assessed with the χ2-test. Kaplan–Meier curves and the log-rank test were used to assess the association between increased plasma mGSTP1 levels and OS. Cox proportional hazards regression was used for multivariable analyses assessing the relationship between change in plasma mGSTP1 levels (detectable vs undetectable) and previously identified clinicopathologic variables. All reported P-values are two-sided. A P-value of 0.05 was used as the criterion for statistical significance without any adjustment for multiple testing. All statistical analyses were performed using Statview 4.5 software (Abacus Systems, Berkeley, CA, USA).
Results
Efficacy of mGSTP1 assays
In the phase I exploratory cohort, we initially used immunomagnetic bead separation of CTCs to assess levels of mGSTP1 in blood from men with CRPC. Isolation of CTCs is expensive and labour-intensive, so we assessed whether free DNA isolated from plasma would be a more sensitive technique for assessment of mGSTP1 levels. For the sequential samples from the first 34 patients (172 paired samples), DNA was isolated from both CTC and plasma samples obtained from ficoll preparations (Supplementary Figure 2). In the paired DNA samples, mGSTP1 was detected approximately twice as frequently in plasma-free DNA 74/172 (43%) than in DNA from CTCs 35/172 (20%) (McNemar’s test, P<0.0001), with a moderate correlation between the log-transformed values measured by the two techniques (R2=0.42, Figure 2A). Owing to the higher sensitivity for mGSTP1 detection, free plasma DNA was used for the remainder of the study.
The relationship between different techniques for extracting DNA assessment of mGSTP1 levels in the Phase I exploratory cohort. (A) A regression analysis of the relationship between mGSTP1 results from DNA extracted from CTC vs free plasma DNA from a ficoll preparation (n=172 paired blood samples). (B) A regression analysis of the relationship between mGSTP1 results from free plasma DNA from a ficoll preparation vs free plasma DNA from a spin preparation (n=98 paired blood samples). (C) A regression analysis of the relationship between mGSTP1 results from free plasma DNA from a spin preparation vs free serum DNA from a spin preparation (n=178 paired blood samples).
Based on the clinical need that a practical assay is required across a wide range of clinical settings, paired plasma samples were collected to assess further refinements in the technique (Supplementary Figure 2). We compared free plasma DNA from Ficoll preparation and DNA from plasma spin from 28 patients (98 paired samples) and found that they correlated well (R2=0.73, Figure 2B). We also compared serum and plasma samples from 32 patients (178 paired samples) using spin protocol and found significant correlation (R2=0.81, Figure 2C). These data indicated that the mGSTP1 assay could be reliably performed on DNA obtained using a simple spin protocol in either serum or plasma samples. All the results described subsequently used quantitative mGSTP1 levels from free DNA from plasma spin.
mGSTP1 levels as prognostic marker in Phase I exploratory cohort
The Phase I exploratory patient cohort consisted of 75 men with CRPC, of whom 89% had chemotherapy during the course of their disease. The baseline characteristics are summarised in Table 1. The median follow-up was 16 months (range 1–44 months) with 51 deaths (68%).
mGSTP1 was detectable in free plasma DNA in two thirds (50/75) of men at presentation to Medical Oncology with CRPC. The undetectable mGSTP1 cohort (25/75) represented 88% (7/8) of those with no metastases, 29% (12/42) with bone only metastases and 29% (6/21) with both bone and visceral metastases. Men with higher Gleason score 9/10 cancers were more likely to have detectable mGSTP1 compared with men with Gleason score 7/8 cancers (82% vs 65%). There was a non-significant trend towards increased mGSTP1 levels in men with bone metastases (Supplementary Figure 3A) and the levels of mGSTP1 were higher in men with Gleason scores of 9 or 10 vs 8 or lower (P=0.07; Supplementary Figure 3B).
Survival analysis demonstrated that detection of plasma mGSTP1 at presentation to Medical Oncology with CRPC was associated with a shorter OS (HR 4.2, 95% CI 2.1–8.2; P<0.0001; Figure 3A). The 2-year survival for men with no detectable plasma mGSTP1 was 71% compared with 23% for men with detectable plasma mGSTP1 levels (Figure 3A). Receiver operating characteristic curve analysis confirmed that detectable mGSTP1 at baseline was associated with death within 12 months (AUC 0.78, 95% CI 0.67–0.89; P<0.001; Figure 3B). Baseline levels of plasma mGSTP1, serum PSA, serum Hb, Gleason score and serum ALP were prognostic for OS (Table 2) on univariable analysis. Performance status was not significant though there were missing data from 40% of patients. Serum LDH was not recorded. On multivariable analysis, detectable mGSTP1 at baseline (HR 3.3, 95% CI 1.4–7.9; P=0.006) and higher levels of baseline serum PSA (HR 1.001, 95% CI 1.0–1.001; P=0.002) were significantly associated with poorer survival (Table 2).
The relationship between plasma mGSTP1, overall survival and response to chemotherapy in men with CRPC in the Phase I exploratory cohort ( n =75). (A) Kaplan–Meier analysis of the relationship between plasma mGSTP1 levels (detectable vs undetectable) at presentation to Medical Oncology with CRPC and overall survival demonstrating that detectable plasma mGSTP1 is associated with a poorer prognosis. (B) Receiver operating characteristic curve assessing the association between baseline mGSTP1 and death within 12 months. (C) Plasma mGSTP1 levels in men with CRPC treated with chemotherapy in the Phase I exploratory cohort (n=35). The relationship between the change in mGSTP1 levels after chemotherapy and the response to chemotherapy is shown in a Waterfall plot assessing the differences between responders (PR) and non-responders (PD/SD). Fold changes >500 were capped at 500 to assist with the presentation of the data. The association between the change in mGSTP1 levels and response to chemotherapy was assessed using a χ2 analysis.
mGSTP1 levels as a therapeutic efficacy marker in Phase I exploratory cohort
Although 67 patients in the cohort were treated with chemotherapy, only 35 patients had sequential plasma samples drawn pre/post the first cycle of chemotherapy. Docetaxel treatment was used in 30 patients and the remainder received Mitoxantrone (Figure 1A). The patients’ characteristics are summarised in Table 3. The median follow-up was 15 months (range 2–38 months), with 22 deaths (63%). Of the 35 patients, 30 had detectable plasma mGSTP1 DNA prior to starting chemotherapy. Detectable plasma mGSTP1 at baseline was associated with shorter OS (HR 6.8, 95% CI 1.5–30.4; P=0.01), but was not associated with subsequent PSA response to therapy (P=0.3). An increase in plasma mGSTP1 DNA levels after the first cycle of chemotherapy was associated with subsequent PSA progression (P=0.008, Figure 3C).
Phase II independent validation cohort
The Phase II validation cohort consisted of 51 men with mCRPC, all of whom received chemotherapy and had paired blood samples available (pre/post one cycle of chemotherapy) (Figure 1B). The baseline clinical characteristics are detailed in Table 3. There were 28 deaths (55%) after a median follow-up of 12 months (range 2–33 months). Prior to chemotherapy, 59% of patients (30/51) had detectable plasma mGSTP1 DNA, and it was detectable after one cycle of chemotherapy in 37% of patients (19/51). Most patients (96%) were treated with docetaxel (Table 3). The PSA PR rate was 33/51 (65%), with SD in 13 (25%), and PD in 5 (10%) (Table 3).
The phase II validation cohort confirmed the prognostic significance of plasma mGSTP1 DNA levels. Detectable plasma mGSTP1 prior to chemotherapy was associated with shorter OS (HR 2.4, 95% CI 1.0–5.6; P=0.039; Figure 4B, Table 4) as was a detectable mGSTP1 after one cycle of chemotherapy (HR 3.7, 95% CI 1.6–8.2; P=0.001; Supplementary Figure 4). In concordance with the TAX327 study (Armstrong et al, 2007), we found that a ⩾30% decline in serum PSA at 3 months was associated with longer OS (P=0.002, Figure 4C). Receiver operating characteristic curve analysis demonstrated that detectable plasma mGSTP1 at baseline and after one cycle of chemotherapy were better predictors of death within 12 months than change in PSA after 3 months (Figure 4D).
The relationship between plasma mGSTP1 levels, overall survival and response to chemotherapy in men with CRPC in the Phase II validation cohort ( n =51). (A) Receiver operating characteristic curve assessing the association between plasma mGSTP1 after one cycle of chemotherapy (detectable vs undetectable) and chemoresistance (PD vs SD+PR). (B) Kaplan–Meier analysis of the relationship between plasma mGSTP1 levels prior to chemotherapy (detectable vs undetectable) and overall survival demonstrating that detectable baseline plasma mGSTP1 is associated with a poorer prognosis. (C) Kaplan–Meier analysis of the relationship between a ⩾30% decline in PSA at 3 months after commencing chemotherapy and overall survival. (D) Receiver operating characteristic curve assessing the association between plasma mGSTP1 (detectable vs undetectable) at baseline, plasma mGSTP1 after one cycle of chemotherapy, decline in PSA at 3 months and death within 12 months.
Univariable analysis revealed that baseline detectable plasma mGSTP1 (P=0.049), Gleason score (P=0.04) and high baseline ALP (P=0.002) were significantly associated with a poorer OS (Table 4). Detectable mGSTP1 at baseline was an independent predictor of OS in multivariable analyses adjusting for baseline PSA, Gleason Score and Hb concentration (P<0.05; Table 4). Serum LDH and ECOG performance status were not available.
High mGSTP1 levels after one cycle of chemotherapy were associated with PSA progression on ROC curve analysis (P=0.006; Figure 4A). Baseline levels of mGSTP1 were not significantly associated with PSA response (P>0.05).
Discussion
This study identifies plasma mGSTP1-free DNA as a potential epigenetic marker of prognosis and therapeutic response in CRPC. Detectable plasma levels of mGSTP1 at presentation with CRPC were associated with poorer survival irrespective of cytotoxic treatment. In addition, detectable plasma mGSTP1 DNA after cycle 1 of chemotherapy was associated with progressive disease and a poorer OS irrespective of response to chemotherapy. Furthermore, the assay for mGSTP1 based on circulating DNA was more sensitive than that based on DNA extraction from CTC.
Assessment of treatment efficacy in the setting of CRPC remains a challenge. The TAX327 and SWOG9916 trials used PSA response, objective tumour response, improvements in pain and quality of life as secondary measures of treatment effect (Petrylak et al, 2004; Tannock et al, 2004). However, 20% of men, who eventually had a PSA response to chemotherapy, had an initial rise in their serum PSA and the median time to PSA response was 44 days (range 26–68 days), so most patients received two or more cycles of treatment before their response status could be ascertained (Tannock et al, 2004). Trials of Sipuleucel-T have demonstrated improvements in OS, without demonstrating effects on other outcomes measures (Kantoff et al, 2010). The identification of earlier and more reliable markers of disease response are needed.
The assessment of CTC numbers to determine prognosis and monitor treatment efficacy is the most developed biomarker in CRPC. The Cellsearch system (Johnson & Johnson, New Brunswick, NJ, USA) is the only CTC technology that has been FDA approved for the monitoring of disease status in PC (Danila et al, 2011). Baseline numbers of CTC before treatment were independently prognostic for survival analysed as a continuous variable (Danila et al, 2007b; Scher et al, 2009b) or using a cutoff (⩾5 CTC per 7.5 ml blood vs 4 CTC) (de Bono et al, 2008). Furthermore, changes in the number of CTC after treatment were more strongly associated with survival than were changes in PSA at 4, 8 and 12 weeks post treatment (de Bono et al, 2008; Scher et al, 2009b). Our assay for plasma mGSTP1 levels at baseline, and after 1 cycle of chemotherapy, were also strongly predictive of OS. However, our study is limited by its lack of direct comparison with CTC enumeration (CellSearch platform) as this technology was not available in Australia at the time. Despite this, the plasma mGSTP1 assay has several advantages: (1) the blood for a mGSTP1 assay can be collected in a standard EDTA tube rather than a specialised CTC tube (CellSave Preservative Tube, Johnson & Johnson), (2) the sample for a mGSTP1 assay can be spun and frozen for later assaying, as DNA methylation is a stable marker, as opposed to the CTC assay that must be run within 96 h of blood collection and (3) the mGSTP1 assay uses standard RT-PCR assay equipment available in most diagnostic laboratories, rather than requiring a specialised machine.
The identification of hypermethylated GSTP1 DNA in plasma in cancer patients is well documented (Thompson et al, 2009; Jeronimo et al, 2011). Hypermethylation of GSTP1 can be detected in samples of prostate, urine and blood. GSTP1 hypermethylation has high specificity (86.8–100%), but low sensitivity for PC detection in both urine (18.8–38.9%) and serum/plasma (13.0–72.5%)(Goessl et al, 2000; Cairns et al, 2001; Jeronimo et al, 2002; Gonzalgo et al, 2003; Roupret et al, 2007). In the advanced PC setting, elevated plasma mGSTP1 levels have been correlated with higher Gleason score at diagnosis (Reibenwein et al, 2007). A recent study of the relationship between CTC levels and several methylated plasma markers, including GSTP1, in 76 men with CRPC demonstrated a significant correlation between ⩾5 CTCs and the detection of mGSTP1 (P<0.001) (Okegawa et al, 2010). Furthermore, men with detectable CTCs and elevated methylated markers in plasma had a poorer prognosis than those with either marker alone (Okegawa et al, 2010). Our study is the first to assess changes in plasma mGSTP1 after chemotherapy. The detection of mGSTP1 can now be done routinely in the clinic with recent commercialisation of PCR-based assays.
Conclusions
We have identified plasma mGSTP1 levels as a potential prognostic marker in men with CRPC, raised the hypothesis that plasma mGSTP1-free DNA levels could be a potential surrogate measure of therapeutic efficacy for cytotoxic chemotherapy and corroborated these findings in an independent Phase II cohort. Assessment of methylated DNA can be performed in a routine pathology laboratory and, unlike CTC, is a stable marker that can still be measured after long-term storage. These data strongly suggest that plasma mGSTP1 levels warrant further investigation, comparison with CTC enumeration, and assessment within prospective clinical trials to determine its clinical utility as a biomarker for response to chemotherapy in CRPC.
Change history
28 October 2014
This paper was modified 12 months after initial publication to switch to Creative Commons licence terms, as noted at publication
References
Armstrong AJ, Garrett-Mayer E, Ou Yang YC, Carducci MA, Tannock I, de Wit R, Eisenberger M (2007) Prostate-specific antigen and pain surrogacy analysis in metastatic hormone-refractory prostate cancer. J Clin Oncol 25: 3965–3970.
Bastian PJ, Palapattu GS, Lin X, Yegnasubramanian S, Mangold LA, Trock B, Eisenberger MA, Partin AW, Nelson WG (2005) Preoperative serum DNA GSTP1 CpG island hypermethylation and the risk of early prostate-specific antigen recurrence following radical prostatectomy. Clin Cancer Res 11: 4037–4043.
Bryzgunova OE, Morozkin ES, Yarmoschuk SV, Vlassov VV, Laktionov PP (2008) Methylation-specific sequencing of GSTP1 gene promoter in circulating/extracellular DNA from blood and urine of healthy donors and prostate cancer patients. Ann N Y Acad Sci 1137: 222–225.
Cairns P, Esteller M, Herman JG, Schoenberg M, Jeronimo C, Sanchez-Cespedes M, Chow NH, Grasso M, Wu L, Westra WB, Sidransky D (2001) Molecular detection of prostate cancer in urine by GSTP1 hypermethylation. Clin Cancer Res 7: 2727–2730.
Danila DC, Fleisher M, Scher HI (2011) Circulating tumor cells as biomarkers in prostate cancer. Clin Cancer Res 17: 3903–3912.
Danila DC, Heller G, Gignac GA, Gonzalez-Espinoza R, Anand A, Tanaka E, Lilja H, Schwartz L, Larson S, Fleisher M, Scher HI (2007a) Circulating tumor cell number and prognosis in progressive castration-resistant prostate cancer. Clin Cancer Res 13: 7053–7058.
Danila DC, Heller G, Gignac GA, Gonzalez-Espinoza R, Anand A, Tanaka E, Lilja H, Schwartz L, Larson S, Fleisher M, Scher HI (2007b) Circulating tumor cell number and prognosis in progressive castration-resistant prostate cancer. Clin Cancer Res 13: 7053–7058.
de Bono JS, Oudard S, Ozguroglu M, Hansen S, Machiels JP, Kocak I, Gravis G, Bodrogi I, Mackenzie MJ, Shen L, Roessner M, Gupta S, Sartor AO (2010) Prednisone plus cabazitaxel or mitoxantrone for metastatic castration-resistant prostate cancer progressing after docetaxel treatment: a randomised open-label trial. Lancet 376: 1147–1154.
de Bono JS, Scher HI, Montgomery RB, Parker C, Miller MC, Tissing H, Doyle GV, Terstappen LW, Pienta KJ, Raghavan D (2008) Circulating tumor cells predict survival benefit from treatment in metastatic castration-resistant prostate cancer. Clin Cancer Res 14: 6302–6309.
Goessl C, Krause H, Muller M, Heicappell R, Schrader M, Sachsinger J, Miller K (2000) Fluorescent methylation-specific polymerase chain reaction for DNA-based detection of prostate cancer in bodily fluids. Cancer Res 60: 5941–5945.
Gonzalgo ML, Pavlovich CP, Lee SM, Nelson WG (2003) Prostate cancer detection by GSTP1 methylation analysis of postbiopsy urine specimens. Clinical Cancer Res 9: 2673–2677.
Hayes JD, Pulford DJ (1995) The glutathione S-transferase supergene family: regulation of GST and the contribution of the isoenzymes to cancer chemoprotection and drug resistance. Crit Rev Biochem Mol Biol 30: 445–600.
Jemal A, Siegel R, Ward E, Hao Y, Xu J, Thun MJ (2009) Cancer Statistics, 2009. CA Cancer J Clin 59: 225–249.
Jeronimo C, Bastian PJ, Bjartell A, Carbone GM, Catto JW, Clark SJ, Henrique R, Nelson WG, Shariat SF (2011) Epigenetics in prostate cancer: biologic and clinical relevance. Eur Urol 60: 753–766.
Jeronimo C, Usadel H, Henrique R, Silva C, Oliveira J, Lopes C, Sidransky D (2002) Quantitative GSTP1 hypermethylation in bodily fluids of patients with prostate cancer. Urology 60: 1131–1135.
Kantoff PW, Higano CS, Shore ND, Berger ER, Small EJ, Penson DF, Redfern CH, Ferrari AC, Dreicer R, Sims RB, Xu Y, Frohlich MW, Schellhammer PF (2010) Sipuleucel-T immunotherapy for castration-resistant prostate cancer. N Engl J Med 363: 411–422.
Lee WH, Morton RA, Epstein JI, Brooks JD, Campbell PA, Bova GS, Hsieh WS, Isaacs WB, Nelson WG (1994) Cytidine methylation of regulatory sequences near the pi-class glutathione S-transferase gene accompanies human prostatic carcinogenesis. Proc Natl Acad Sci USA 91: 11733–11737.
McShane LM, Altman DG, Sauerbrei W, Taube SE, Gion M, Clark GM (2005) REporting recommendations for tumour MARKer prognostic studies (REMARK). Br J Cancer 93: 387–391.
Millar DM, Ow K, Paul C, Russell P, Molloy P, Clark SJ (1999) Detailed methylation analysis of the glutathione S-transferase pi (GSTP1) gene in prostate cancer. Oncogene 18: 1313–1324.
Okegawa T, Nutahara K, Higashihara E (2010) Association of circulating tumor cells with tumor-related methylated DNA in patients with hormone-refractory prostate cancer. Int J Urol 17: 466–475.
Petrylak DP, Tangen CM, Hussain MH, Lara PNJ, Jones JA, Taplin ME, Burch PA, Berry D, Moinpour C, Kohli M, Benson MC, Small EJ, Raghavan D, Crawford ED (2004) Docetaxel and estramustine compared with mitoxantrone and prednisone for advanced refractory prostate cancer. N Engl J Med 351: 1513–1520.
Rand K, Qu W, Ho T, Clark S, Molloy P (2002) A Companion to Methods in Enzymology Methods 27: 114–120.
Reibenwein J, Pils D, Horak P, Tomicek B, Goldner G, Worel N, Elandt K, Krainer M (2007) Promoter hypermethylation of GSTP1, AR, and 14-3-3sigma in serum of prostate cancer patients and its clinical relevance. Prostate 67: 427–432.
Roupret M, Hupertan V, Yates DR, Catto JW, Rehman I, Meuth M, Ricci S, Lacave R, Cancel-Tassin G, de la Taille A, Rozet F, Cathelineau X, Vallancien G, Hamdy FC, Cussenot O (2007) Molecular detection of localized prostate cancer using quantitative methylation-specific PCR on urinary cells obtained following prostate massage. Clin Cancer Res 13: 1720–1725.
Rushmore TH, Pickett CB (1993) Glutathione S-transferases, structure, regulation, and therapeutic implications. J Biol Chem 268: 11475–11478.
Ruzza P, Rosato A, Rossi CR, Floreani M, Quintieri L (2009) Glutathione transferases as targets for cancer therapy. Anticancer Agents Med Chem 9: 763–777.
Scher HI, Jia X, de Bono JS, Fleisher M, Pienta KJ, Raghavan D, Heller G (2009a) Circulating tumour cells as prognostic markers in progressive, castration-resistant prostate cancer: a reanalysis of IMMC38 trial data. Lancet Oncol 10: 233–239.
Scher HI, Jia X, de Bono JS, Fleisher M, Pienta KJ, Raghavan D, Heller G (2009b) Circulating tumour cells as prognostic markers in progressive, castration-resistant prostate cancer: a reanalysis of IMMC38 trial data. Lancet Oncol 10: 233–239.
Tannock IF, de Wit R, Berry WR, Horti J, Pluzanska A, Chi KN, Oudard S, Theodore C, James ND, Turesson I, Rosenthal MA, Eisenberger MA (2004) Docetaxel plus prednisone or mitoxantrone plus prednisone for advanced prostate cancer. N Engl J Med 351: 1502–1512.
Thompson IM, Ankerst DP, Tangen CM (eds) (2009) Current Clinical Urology: Prostate Cancer Screening. Humana Press: Totowa, NJ, USA.
Acknowledgements
Cancer Institute New South Wales, National Health and Medical Research Council, Australian Cancer Research Foundation, Cancer Australia, Prostate Cancer Foundation of Australia, RT Hall Trust, Tour de Cure, Sydney Catalyst and Cancer Council NSW. We thank the late Robert L Sutherland for his role in the development and support of this project.
Author Contributions
Concept and design of study: KL Mahon, MJ Boyer, PL Molloy, LG Horvath, SJ Clark Data acquisition/analysis/interpretation: all authors. Draft article and revising it: all authors. Final approval of article: all authors. Agreement to be accountable for all aspects of the work in ensuring that questions related to the accuracy or integrity of any part of the work are appropriately investigated and resolved: all authors.
Author information
Authors and Affiliations
Consortia
Corresponding author
Ethics declarations
Competing interests
Diagnostic assay for methylation in the GSTP1 gene; International Patent Application No. PCT/AU99/00306; Inventors: Clark SJ & Molloy PL was licensed to Epigenomics AG.
Additional information
This work is published under the standard license to publish agreement. After 12 months the work will become freely available and the license terms will switch to a Creative Commons Attribution-NonCommercial-Share Alike 3.0 Unported License.
Supplementary Information accompanies this paper on British Journal of Cancer website
Rights and permissions
From twelve months after its original publication, this work is licensed under the Creative Commons Attribution-NonCommercial-Share Alike 3.0 Unported License. To view a copy of this license, visit http://creativecommons.org/licenses/by-nc-sa/3.0/
About this article
Cite this article
Mahon, K., Qu, W., Devaney, J. et al. Methylated Glutathione S-transferase 1 (mGSTP1) is a potential plasma free DNA epigenetic marker of prognosis and response to chemotherapy in castrate-resistant prostate cancer. Br J Cancer 111, 1802–1809 (2014). https://doi.org/10.1038/bjc.2014.463
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/bjc.2014.463
Keywords
This article is cited by
-
Blood-based liquid biopsies for prostate cancer: clinical opportunities and challenges
British Journal of Cancer (2022)
-
Genomic and phenotypic heterogeneity in prostate cancer
Nature Reviews Urology (2021)
-
Epigenetic Regulation in Prostate Cancer Progression
Current Molecular Biology Reports (2018)
-
Liquid biopsy: a step forward towards precision medicine in urologic malignancies
Molecular Cancer (2017)
-
Quantitative comparison of DNA methylation assays for biomarker development and clinical applications
Nature Biotechnology (2016)