Abstract
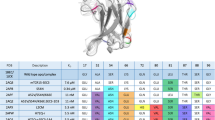
Antigen recognition by T lymphocytes is mediated by cell-surface glycoproteins known as T-cell antigen receptors (TCRs). These are composed of α and β, or γ and δ, polypeptide chains with variable (V) and constant (C) regions. In contrast to αβ TCRs, which recognize antigen only as peptide fragments bound to molecules of the major histocompatibility complex (MHC), γδ TCRs appear to recognize proteins directly, without antigen processing, and to recognize MHC molecules independently of the bound peptide1,2,3,4. Moreover, small phosphate-containing non-peptide compounds have also been identified as ligands for certainγδ T cells5,6. These studies indicate that antigen recognition by γδ TCRs may be fundamentally different from that by αβ TCRs. The three-dimensional structures of several αβ TCRs and TCR fragments7,8,9,10, and their complexes with peptide–MHC or superantigens9,10,11, have been determined. Here we report the crystal structure of the Vδ domain of a human γδ TCR at 1.9 Å resolution. A comparison with antibody and αβ TCR V domains reveals that the framework structure of Vδ more closely resembles that of VHthan of Vα, Vβ or VL(where H and L refer to heavy and light chains), whereas therelative positions and conformations of its complementarity-determining regions (CDRs) share features of both Vα and VH. These results provide the first direct evidence that γδ TCRs are structurally distinct from αβ TCRs and, together with the observation that the CDR3 length distribution of TCR δ chains is similar to that of immunoglobulin heavy chains12, are consistent with functional studies suggesting that recognition of certain antigens by γδ TCRs may resemble antigen recognition by antibodies1,2,3.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Schild, H.et al. The nature of major histocompatibility complex recognition by γδ T-cells. Cell 76, 29–37 (1994).
Chien, Y., Jores, R. & Crowley, M. P. Recognition by γ/δ T cells. Annu. Rev. Immunol. 14, 511–532 (1996).
Crowley, M. P., Reich, Z., Mavaddat, N., Altman, J. D. & Chien, Y. The recognition of the nonclassical major histocompatibility complex (MHC) class I molecule, T10, by the γδ T cell, G8. J. Exp. Med. 185, 1223–1230 (1997).
Sciammas, R.et al. Unique antigen recognition by a herpesvirus-specific TCR-γ-δ cell. J. Immunol. 152, 5392–5397 (1994).
Constant, P.et al. Stimulation of human γδ T-cells by nonpeptidic mycobacterial ligands. Science 264, 267–270 (1994).
Tanaka, Y.et al. Natural and synthetic non-peptide antigens recognized by human γδ T-cells. Nature 375, 155–158 (1995).
Bentley, G. A., Boulot, G., Karjalainen, K. & Mariuzza, R. A. Crystal structure of the β chain of a T-cell antigen receptor. Science 267, 1984–1987 (1995).
Fields, B. A.et al. Crystal structure of the Vα domain of a T-cell antigen receptor. Science 270, 1821–1824 (1995).
Garcia, K. C.et al. An αβ T cell receptor structure at 2.5 Å resolution and its orientation in the TCR-MHC complex. Science 274, 209–219 (1996).
Garboczi, D. N.et al. Structure of the complex between human T-cell receptor, viral peptide and HLA-A2. Nature 384, 134–141 (1996).
Fields, B. A.et al. Crystal structure of the β chain of a T-cell receptor complexed with a superantigen. Nature 384, 188–192 (1996).
Rock, E. P., Sibbald, P. R., Davis, M. M. & Chien, Y. CDR3 length in antigen-specific immune receptors. J. Exp. Med. 179, 323–328 (1994).
Spits, H., Paliard, X., Engelhard, V. H. & de Vries, J. E. Cytotoxic activity and lymphokine production of T cell receptor (TCR)-αβ+ and TCR-γδ+ cytotoxic T lymphocyte (CTL) clones recognizing HLA-A2 and HLA-A2 mutants. J. Immunol. 144, 4156–4162 (1990).
Fan, Z.et al. Three-dimensional structure of an Fv from a human IgM immunoglobulin. J. Mol. Biol. 228, 188–207 (1992).
Kabat, E. A., Wu, T. T., Perry, H. M., Gottesman, K. S. & Foeller, C. Sequences of Proteins of Immunological Interest5th edn (Public Health Services, National Institutes of Health, Washington DC, 1991).
Arden, B., Clark, S. P., Kabelitz, D. & Mak, T. W. Human T-cell receptor variable gene segment families. Immunogenetics 42, 455–500 (1995).
Haas, W., Pereira, P. & Tonegawa, S. Gamma/delta cells. Annu. Rev. Immunol. 11, 637–685 (1993).
Bork, P., Holm, L. & Sander, C. The immunoglobulin fold: structural classification, sequence patterns and common core. J. Mol. Biol. 242, 309–320 (1994).
Chothia, C.et al. Conformations of immunoglobulin hypervariable regions. Nature 342, 877–883 (1989).
Bhat, T. N.et al. Bound water molecules and conformational stabilization help mediate an antigen-antibody association. Proc. Natl Acad. Sci. USA 91, 1089–1093 (1994).
Bentley, G. A., Boulot, G., Riottot, M.-M. & Poljak, R. J. Three-dimensional structure of an idiotope-anti-idiotope complex. Nature 348, 254–257 (1990).
Lebedeva, M. I.et al. Cloning, expression, and crystallization of the Vδ domain of a human γδ T-cell receptor. Protein Sci. 5, 2638–2642 (1996).
Krangel, M. S., Yssel, H., Brocklehurst, C. & Spits, H. Adistinct wave of human T cell receptor γ/δ lymphocytes in the early fetal thymus: evidence for controlled rearrangement and cytokine production. J. Exp. Med. 172, 847–859 (1990).
Correa, I.et al. Most γδ cells develop normally in β2-microglobulin-deficient mice. Proc. Natl Acad. Sci. USA 89, 653–657 (1992).
Bigby, M.et al. Most γδ T cells develop normally in the absence of MHC class II molecules. J. Immunol. 9, 4465–4475 (1993).
Howard, A. J.et al. Use of an imaging proportional counter in macromolecular crystallography. J.Appl. Crystallogr. 20, 383–387 (1987).
Messerschmidt, A. & Pflugrath, J. W. Crystal orientation and X-ray pattern prediction routines for area-detector diffractometer systems in macromolecular crystallography. J. Appl. Crystallogr. 20, 306–315 (1987).
Otwinowski, Z. & Minor, W. Processing X-ray diffraction data collected in oscillation mode. Methods Enzymol. 276, 307–326 (1997).
Collaborative Computational Project No. 4. Acta Crystallogr.D 50, 760–763 (1994).
Cowtan, K. Joint CCP4 and ESF-EACBM Newsl. Protein Crystallogr. 31, 34–38 (1994).
Acknowledgements
We thank X. Ysern and B. C. Braden for valuable discussions, X. Ji for help in map interpretation, R. J. Poljak for critical reading of the manuscript, and the staff at Brookhaven National Laboratory for assistance with data collection. We also thank I. A. Wilson for providing coordinates of the 2C TCR and D. N. Garboczi and D. C. Wiley for coordinates of the A6 TCR. This work was supported by grants from the NIH (R.A.M. and M.B.B.) and the National Multiple Sclerosis Society (R.A.M.).
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Li, H., Lebedeva, M., Llera, A. et al. Structure of the Vδ domain of a human γδ T-cell antigen receptor. Nature 391, 502–506 (1998). https://doi.org/10.1038/35172
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1038/35172
This article is cited by
-
Expression, Purification, and Crystallization of the Vγ9Vδ2 T-cell Receptor Recognizing Protein/Peptide Antigens
The Protein Journal (2023)
-
Epitope-Based Vaccine Designing of Nocardia asteroides Targeting the Virulence Factor Mce-Family Protein by Immunoinformatics Approach
International Journal of Peptide Research and Therapeutics (2020)
-
Long-term use of interferon-β in multiple sclerosis increases Vδ1−Vδ2−Vγ9− γδ T cells that are associated with a better outcome
Journal of Neuroinflammation (2019)
-
The burgeoning family of unconventional T cells
Nature Immunology (2015)
-
Identification of an Ideal-like Fingerprint for a Protein Fold using Overlapped Conserved Residues based Approach
Scientific Reports (2014)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.