Abstract
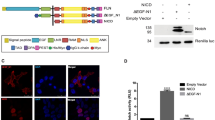
The Notch genes encode single-pass transmembrane receptors that transduce the extracellular signals responsible for cell fate determination during several steps of metazoan development. The mechanism by which extracellular signals affect gene transcription and ultimately cell fate decisions is beginning to emerge for the Notch signalling pathway. One paradigm is that ligand binding to Notch triggers a Presenilin1-dependent proteolytic release of the Notch intracellular domain from the membrane1, resulting in low amounts of Notch intracellular domain which form a nuclear complex with CBF1/Su(H)/Lag1 to activate transcription of downstream targets2. Not all observations clearly support this processing model, and the most rigorous test of it is to block processing in vivo and then determine the ability of unprocessed Notch to signal. Here we report that the phenotypes associated with a single point mutation at the intramembranous processing site of Notch1, Val1,744→Gly, resemble the null Notch1 phenotype3,4. Our results show that efficient intramembranous processing of Notch1 is indispensable for embryonic viability and proper early embryonic development in vivo.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
References
Chan, Y. M. & Jan, Y. N. Roles for proteolysis and trafficking in Notch maturation and signal transduction. Cell 94 , 423–426 (1998).
Jarriault, S. et al. Signalling downstream of activated mammalian Notch. Nature 377, 355–358 ( 1995).
Conlon, R. A., Reaume, A. G. & Rossant, J. Notch1 is required for the coordinate segmentation of somites. Development 121, 1533– 1545 (1995).
Swiatek, P. J., Lindsell, C. E., Amo, F. F. d., Weinmaster, G. & Gridley, T. Notch1 is essential for postimplantation development in mice. Genes Dev. 8, 707– 719 (1994).
Kidd, S., Lieber, T. & Young, M. W. Ligand-induced cleavage and regulation of nuclear entry of Notch in Drosophila melanogaster embryos. Genes Dev. 12, 3728–3740 ( 1998).
Schroeter, E. H., Kisslinger, J. A. & Kopan, R. Notch-1 signalling requires ligand-induced proteolytic release of intracellular domain. Nature 393, 382–386 (1998).
De Strooper, B. et al. A presenilin-1-dependent gamma-secretase-like protease mediates release of Notch intracellular domain. Nature 398, 518–521 (1999).
Lecourtois, M. & Schweisguth, F. Indirect evidence for Delta -dependent intracellular processing of Notch in Drosophila embryos. Curr. Biol. 8, 771–774 (1998).
Struhl, G. & Adachi, A. Nuclear access and action of Notch in vivo. Cell 93, 649–660 (1998).
Donoviel, D. B. et al. Mice lacking both presenilin genes exhibit early embryonic patterning defects. Genes Dev. 13, 2801– 2810 (1999).
Herreman, A. et al. Presenilin 2 deficiency causes a mild pulmonary phenotype and no changes in amyloid precursor protein processing but enhances the embryonic lethal phenotype of presenilin 1 deficiency. Proc. Natl Acad. Sci. USA 96, 11872–11877 ( 1999).
Li, X. & Greenwald, I. HOP-1, a Caenorhabditis elegans presenilin, appears to be functionally redundant with SEL-12 presenilin and to facilitate LIN-12 and GLP-1 signaling. Proc. Natl Acad. Sci. USA 94, 12204–12209 (1997).
Westlund, B., Parry, D., Clover, R., Basson, M. & Johnson, C. D. Reverse genetic analysis of Caenorhabditis elegans presenilins reveals redundant but unequal roles for sel-12 and hop-1 in Notch-pathway signaling. Proc. Natl Acad. Sci. USA 96, 2497–2502 (1999).
Artavanis-Tsakonas, S., Rand, M. D. & Lake, R. J. Notch signaling: cell fate control and signal integration in development. Science 284, 770– 776 (1999).
Ye, Y. H., Lukinova, N. & Fortini, M. E. Neurogenic phenotypes and altered Notch processing in Drosophila Presenilin mutants. Nature 398 , 525–529 (1999).
Berechid, B. E., Thinakaran, G., Wong, P. C., Sisodia, S. S. & Nye, J. S. Lack of requirement for Presenilin1 in Notch1 signaling. Curr. Biol. 9, 1493 –1496 (1999).
Kopan, R., Schroeter, E. H., Weintraub, H. & Nye, J. S. Signal transduction by activated mNotch: importance of proteolytic processing and its regulation by the extracellular domain. Proc. Natl Acad. Sci. USA 93, 1683–1688 ( 1996).
Krebs, L. T. et al. Notch signaling is essential for vascular morphogenesis in mice. Genes Dev. 14, 1343– 1352 (2000).
Neidhardt, L. M., Kispert, A. & Herrmann, B. G. A mouse gene of the paired-related homeobox class expressed in the caudal somite compartment and in the developing vertebral column, kidney and nervous system. Dev. Genes Evol. 207, 330–339 (1997).
Bettenhausen, B., de Angelis, M. H., Simon, D., Guenet, J. -L. & Gossler, A. Transient and restricted expression during mouse embryogenesis of Dll1, a murine gene closely related to Drosophila Delta. Development 121, 2407 –2418 (1995).
de la Pompa, J. L. et al. Conservation of the Notch signalling pathway in mammalian neurogenesis. Development 124, 1139– 1148 (1997).
Reaume, A. G., Conlon, R. A., Zirngibl, R., Yamaguchi, T. P. & Rossant, J. Expression analysis of a Notch homologue in the mouse embryo. Dev. Biol. 154, 377–387 (1992).
Schweisguth, F. Dominant-negative mutation in the β2 and β6 proteasome subunit genes affect alternative cell fate decisions in the Drosophila sense organ lineage. Proc. Natl Acad. Sci. USA 96, 11382 –11386 (1999).
Carmeliet, P. et al. Abnormal blood vessel development and lethality in embryos lacking a single VEGF allele. Nature 380, 435–439 (1996).
Lewandoski, M. & Martin, G. R. Cre-mediated chromosome loss in mice. Nature Genet. 17, 223–225 (1997).
Yuan, W. et al. The mouse SLIT family: Secreted ligands for ROBO expressed in patterns that suggest a role in morphogenesis and axon guidance. Dev. Biol. 212, 290–306 (1999).
Milner, L. A. et al. Inhibition of granulocytic differentiation by mNotch1. Proc. Natl Acad. Sci. USA 93, 13014– 13019 (1996).
Girard, L. et al. Frequent provirus insertional mutagenesis of Notch1 in thymomas of MMTVD/myc transgenic mice suggests a collaboration of c-myc and Notch1 for oncogenesis. Genes Dev. 10, 1930– 1944 (1996).
Acknowledgements
We thank T. Ley, E. Ross, D. Ornitz and S. Hua for technical help; R. Conlon and T. Gridley for providing animals and communicating results before publication; R. Aguilera, D. Henrique, B. Herrmann, G. Martin, J. Sanes, M. Thayer and S. Troyanovsky for reagents; R. Cagan and O. Pourquie for reading and commenting on the manuscript; and members of the Kopan lab—X. Tian, A. Nichols, M.-H. Lin, J. Kisslinger, B. Hadland and J. Books. This work was supported by the NIH.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Huppert, S., Le, A., Schroeter, E. et al. Embryonic lethality in mice homozygous for a processing-deficient allele of Notch1. Nature 405, 966–970 (2000). https://doi.org/10.1038/35016111
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1038/35016111
This article is cited by
-
Class II phosphatidylinositol 3-kinase-C2α is essential for Notch signaling by regulating the endocytosis of γ-secretase in endothelial cells
Scientific Reports (2021)
-
A non-canonical Notch complex regulates adherens junctions and vascular barrier function
Nature (2017)
-
Characterization of activating mutations of NOTCH3 in T-cell acute lymphoblastic leukemia and anti-leukemic activity of NOTCH3 inhibitory antibodies
Oncogene (2016)
-
Taking inflammatory bowel disease up a Notch
Immunologic Research (2012)
-
Potent amyloidogenicity and pathogenicity of Aβ43
Nature Neuroscience (2011)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.