Abstract
Brca1 mRNA was detectable in female mouse mammary gland tissue from adult virgins, during pregnancy and early lactation. It was associated with phases of mammary epithelial cell proliferation and differentiation but the pattern of Brca1 expression was dissociable from that of a true differentiation marker, β-casein, by virtue of its significant expression in the virgin gland and termination of its expression in early lactation. In a primary cell culture model, association of a laminin-rich extracellular matrix (ECM) with mammary epithelial cells was required for cell survival and cell differentiation and suppressed Brca1 expression in these cells. ECM-association may significantly contribute to the final restriction in Brca1 expression in the lactating gland in vivo. Interestingly, our results suggest that mammary epithelial cells undergo apoptosis both when expressing and when not expressing Brca1, depending on whether the dying cell populations had been terminally differentiated or not.
Similar content being viewed by others
Introduction
Mutations at the human breast cancer susceptibility gene (BRCA1) locus account for ∼45% of early-onset familial breast cancer and 80–90% of inherited co-occurring breast and ovarian malignancies.1 The murine homologue, Brca1, encodes a 1812 amino acid protein which displays 58% homology and similar charge and hydrophilic character to its human counterpart.2,3 BRCA1 has been variously localised to the nucleus of normal breast epithelial cells,4,5,6,7,8 to cylinders derived from cytoplasmic structures which anastamose through the nucleus9 and to the secretory apparatus10 and associated secretions.11,12 In addition, it has been reported to both remain in the nucleus7 and to be present in the cytoplasm of breast cancer cell4,5,6 (reviewed in13,14). Some or all of these observations may be reconciled by the hypothesis that BRCA1 transcripts are differentially spliced,15,16,17 and that the different mature proteins are directed to specific subcellular sites. For instance, nuclear localisation signals have been identified as encoded in exon 11 of human BRCA115,18 but these are obviously absent in the common exon 11-minus splice variant (reviewed in13). The occurrence in BRCA1 (and BRCA2) of a DNA-binding RING finger domain19 in the N-terminal region2,3,20 and of a transactivation domain at the C-terminus21,2223 indicates that these proteins act as transcription factors.24 This hypothesis is supported by observations which implicate BRCA1 in chromatin remodelling,25 and reports that the protein has been associated with the histone deacetylase complex.26 BRCA1 also appears to have a direct involvement with the process of gene transcription: it has been identified as the hSRB7 component of the holoenzyme RNA polymerase II,27 and the BRCA1 C-terminal (BRCT) domain has been reported to associate with RNA helicase A.28 Another function attributed to BRCA1 homologues is in the maintenance of genomic integrity, possibly through an involvement in transcription-coupled DNA repair.29,30 A functional relationship has been demonstrated between BRCA1 and cellular agents which mediate the resolution of double-stranded DNA breaks during recombination31,32,33,34 (see35), and BRCA1 has been localised to the centrosome during mitosis.36 It has also been suggested that the mouse homologue, Brca1, may interact with the DNA-repair protein p53 which shows a similar pattern of expression.37,38,39
Brca1 carries out one or more indispensable functions in early embryogenesis and appears to be required for embryonic proliferation and the development of germ layers and neural structures.38,40,41,42 That a deficit in cell proliferation underlies the failure of the Brca1 (and Brca2) null embryos to survive is further suggested by their overexpression of the p21/Waf1 cyclin dependent kinase inhibitor.41,43 The expression of Brca1 at multiple sites of organogenesis in later embryogenesis, and in a diversity of adult tissues including the thymus, testis, breast epithelium and ovarian follicles,37,44 suggests an involvement for the product(s) of this gene in one or more fundamental processes of development, and that it is expressed at sites of proliferation where differentiation will soon follow. Furthermore, experimental inhibition of BRCA1 expression produced accelerated growth of normal and malignant mammary epithelial cells but not of non-mammary epithelial cells45 and over-expression of BRCA1 produced growth retardation in vitro, specifically of breast and ovarian tumour cell lines.46
The reported expression of Brca1 in postnatal mammary epithelium during puberty and pregnancy37,44 suggests that the gene product functions in proliferating and/or differentiated cells. Mammary transcription of the Brca1 gene has thus far been reported to be high in mid- and late-pregnancy, appearing to wane upon lactation and early involution, but becoming prominent again 5–7 days post-weaning.37,44 Brca1 expression is upregulated in cultured cells from G1/S to M47 again implying a role in proliferation. The expression of the human homologue, BRCA1, and BRCA2 would seem to be similarly cell cycle regulated with mRNA levels being increased at late G1/early S-phase, preceding DNA synthesis, in cultured cells.15,48,49 The kinetics of upregulation of Brca1 expression during mammary epithelial activity in puberty and pregnancy are echoed by Brca2.39,47
We have investigated the levels of Brca1 mRNA in two model systems of mammary epithelial cell proliferation, differentiation and apoptosis. We show that Brca1 mRNA is expressed in the virgin, pregnant and for the first time in the early lactating gland which coincides with the time of a final burst of alveolar epithelial cell proliferation. Notably, Brca1 was not expressed in early involution when a wave of epithelial cell apoptosis occurs. In the virgin gland and at late lactation, Brca1 expression diverged sharply from that of the differentiation marker milk protein β-casein transcript, which is detected during pregnancy, lactation and early involution. In primary cultures of epithelial cells from mid-pregnant mammary gland tissue we found Brca1 mRNA in populations of proliferating cells and of quiescent cells which were predisposed to apoptosis. However, overlaying the quiescent population with a laminin-rich extracellular matrix which ensured cell survival and permitted cell differentiation was sufficient to suppress Brca1 expression. We hypothesise that Brca1 expression is primarily associated with proliferation, and is restricted in fully differentiated mammary epithelial cells by an ECM-integrin receptor-mediated regulatory mechanism.
Results
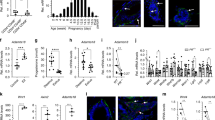
Whole mount analysis50 of adult mouse mammary gland (Figure 1A) reveals the growth of the ductal and alveolar systems from the virginal state, through pregnancy to full development in lactation: by mid-pregnancy abundant ductal branching and strong initial alveolar development has become evident. In lactation full alveolar development has been achieved and maximum colonisation of the entire fat pad is apparent. These events reflect waves of epithelial cell proliferation during pregnancy and as late as early in lactation.51 Weaning triggers the onset of involution that involves, sequentially, milk stasis in the alveoli, a phase of intense alveolar epithelial cell apoptosis and finally a phase of reductive glandular remodelling that is associated with significant local matrix metalloprotease activity.52,53 The whole mount of the 4-day involuting gland shows significant alveolar regression and this continues with time (Figure 1A and results not shown).
(A) Development and regression of ductal and alveolar elements in the adult mammary gland. Carmine stained whole mounts of (left to right): virgin (5 weeks), pregnant (14 days), lactating and involuting (4 days) mammary gland. (B) RT–PCR analysis of Brca 1, β-casein and gpdh mRNA levels in virgin (5 weeks), pregnant (9, 14, 15, 17 and 19 days), lactating (2, 4 and 8 days) and involuting (2, 4 and 8 days) mammary gland. Water blanks (Q) are shown with each analytical set; the analysis of gpdh amplification in the absence of RT addition to the RNA samples is also shown; 2 μg of total mammary gland RNA was used in each analysis. The data presented are representative of three independent experiments. (C) Verification of Brca1 PCR analysis. (Top) Cartoon mapping primer and nesting primer/probe positions; the Brca1 exon structure; the site of the Ear1 restriction site; and the relative sizes of the amplified (primers 1/3), semi-nested (primers 2/3) and Ear1 restricted fragments are shown. (Bottom, left) Typical PCR product (primers 1/3) unrestricted (lane 1) and restricted with Ear1; (middle) Southern blot of partial Ear1 restriction of typical PCR product probed with primer 2; (left) Typical PCR product (primers 1/3 lanes 1, 2) and after semi-nesting with primers 2/3 yield expected product sizes of 535 and 366 bp, respectively (lanes 1, 3: pregnant mammary gland; lanes 2, 4 proliferating primary cultured mid-pregnant mammary epithelial cells; and lanes 3, 6: water blanks)
The proliferation of the alveolar mammary epithelial cells during pregnancy is accompanied by gradual asynchronous differentiation of the cells as pregnancy progresses.54 The differentiation of the alveolar element of the mammary gland can be followed by following the appearance of transcripts of milk protein genes as the gland develops.52 Figure 1B charts the analysis of β-casein transcripts in total mRNA isolated from virgin, pregnant, lactating, and involuting mammary gland by RT–PCR using a limited cycle-number PCR. Under these conditions no β-casein transcripts were detectable in the virgin gland, but transcript was detectable from 8 days of pregnancy onwards through lactation and early involution eventually becoming undetectable after 8 days of involution (see also52). It has been of interest to attempt to associate Brca1 expression with either the proliferating and/or proliferating and differentiated states of the mammary epithelial cell.37,39,47 RT–PCR analysis of Brca1 transcripts (Figure 1B) show their presence to be dissociated from that of β-casein transcripts. Brca1 transcripts are detectable in adult virgin glands where we did not detect β-casein; generally, Brca1 and β-casein expression parallel each other during pregnancy. However, we here and others37 report relatively low levels of Brca1 expression at day 14 of pregnancy. At this time β-casein expression would seem maintained. The event that dictates this reduction in Brca1 transcript levels remains unclear. In this report, for the first time, the presence of Brca1 transcripts has been investigated in early lactation, and are found to be present. Knight and Peaker51 have shown that a final surge in epithelial cell proliferation occurs early in lactation. It is not known what drives this phase of epithelial cell proliferation but Brca1 expression may very well accompany it. Thereafter, Brca1 and β-casein expression diverge. Brca1 transcripts were not again detectable until 4 and 8 days after weaning when they were detectable at trace levels. β-casein expresion on the other hand is maintained throughout lactation and is eventually lost at day 8 of involution. Notably, Brca1 was not expressed in early involution (day 2) when a wave of epithelial cell apoptosis occurs.51
Figure 1C demonstrates the specificity of the Brca1 PCR measurement; the first panel shows the digestion of the RT–PCR product at a predicted unique internal Ear1 site (see cartoon); the second panel shows the result of a Southern blot of the Ear1 digestion products and target DNA using an internal probe which detects the target DNA band and the 303 bp product, as predicted. The third panel shows the result of a hemi-nested PCR with: the primary PCR product (535 bp) as template; the ‘Southern probe’ oligonucleotide as forward primer and using the primary PCR reverse primer. This generates a single band of 366 bp, as predicted (lanes 4 and 5). We therefore judge our Brca1 RT–PCR products from the tissue samples and the primary cell cultures to validly reflect cellular Brca1 transcripts.
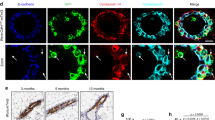
A series of studies have used cell culture systems to model the proliferative, differentiative/surviving and apoptotic fates of the alveolar mammary epithelial cell.55,56,57 These studies emphasise the capacity of laminin-rich extracellular matrix to support both the survival and differentiation of these mammary epithelial cells (in the presence of lactogenic hormones). Figure 2A shows how overlay of primary cultures of mid-pregnant mammary epithelial cells with laminin rich-ECM (Matrigel) suppresses the apoptotic loss of the cells when cultured on plastic, as judged by the suppression of the apoptosis-associated sub-G0/G1 peak (compare panel (c) to panels (b1) and (b2)). This analysis is corroborated by the measurement of genomic DNA fragmentation by the TUNEL end-labelling technique using flow cytometric analysis of fluorescence (Figure 2B): ECM overlay suppressed the extent of DNA end-labelling detectable in cells not overlaid with ECM. As there was a significant and obvious loss of cells into the medium from the populations cultured in the absence of ECM-overlay, FACS and TUNEL/FACS analysis was carried out both on the ‘floating cells’ in the medium and those remaining adherent, in this study. The population of proliferating cells showed the expected FACS cell cycle pattern of DNA-associated PI fluorescence (Figure 2A) and a low level of genomic DNA fragmentation. Survival of the cells could be ensured by substituting laminin overlay for EHS-Matrigel matrix but not by culturing the cells on a collagen matrix.58 As well as providing a survival stimulus, it is well known that association of ECM with such mammary epithelial cells is permissive for differentiation responses (e.g. the expression of milk protein genes in the presence of lactogenic hormones):55,57 In Figure 2C it can be seen that transcripts of milk protein genes, β-casein and WAP (panels 3 and 4), are detectable in the population of cells cultured in the presence of ECM overlay and lactogenic hormones but not in the proliferating cells or those cultured in the absence of ECM overlay (panels 3 and 4). We could additionally define that expression of transcripts encoding the oncoprotein receptor, neu, was only detectable in the proliferating cells (Figure 2C, panel 1). Furthermore, transcripts for the apoptosis associated protein, Bax, were detectable in the proliferating and apoptotic cell populations but were undetectable after ECM-overlay (panel 2). Against this background we analyzed Brca1 transcripts in the primary cultured cells (panel 5): transcripts were detected in the proliferating cells, and in the population of cells not overlaid with ECM but were not detectable in the ECM-overlaid cells. (gpdh transcripts were equally detectable in all three cell populations (panel 6)). These results suggest that overlay of primary mammary epithelial cells with laminin-rich ECM which ensures cell survival and is permissive for cell differentiation is also sufficient to suppress Brca1 gene expression. We hypothesise that ECM (laminin) interactions with β1-integrin receptors55,59,60,61 on the alveolar epithelial cell membrane contributes significantly to the suppression of Brca1 expression in later lactation (Figure 1B).
Brca1 expression in proliferating, apoptoic and differentiated/surviving (ECM-overlaid) mouse mammary epithelial cells in primary culture. (A) FACS/cell cycle analysis of proliferating (a), apoptotic (‘pellet’ (adherent cells; (b1)) and ‘supernatant’) non-adherent population; (b2)) and ECM-overlaid (c) primary mammary epithelial cells. Propidium iodide-induced fluorescence (X-axis) is plotted against cell counts Y-axis. HIP, hydrocortisone, insulin and prolactin. Results shown are representative of three independent experiments. (B) FACS/TUNEL analysis of proliferating, apoptotic (adherent and non-adherent populations, as in (A) above) and ECM-overlaid cells. Graph shows exent of terminal transferase (TdT) fluorescent labelling of free 3′-OH ends generated by DNA-fragmentation. Data are presented as fold induction in incorporation of FITC label over control (samples from the same culture plate to which no TdT was added). Data for each culture condition were generated from three independent experiments; and (C) Brca1 expression in primary cultured mouse mammary epithelial cells. RT–PCR analysis of neu, Bax, β-casein, WAP, Brca 1 and gpdh in total RNA samples from proliferating (P), apoptotic (−E) or ECM-overlaid (+E) cultured primary mammary epithelial cells. Results of gpdh analysis at 25 cycles of amplification and where RT was omitted from the first strand synthesis step are also presented. The Brca1 RT–PCR analysis presented is representative of six independent experiments
In summary our results lead us to three major conclusions: (1) That Brca1 expression in both model systems studied, in vivo and in vitro (primary cell culture), is primarily linked to cell proliferation; (2) That Brca1 expression in both model systems studied is repressed in the fully differentiated, ‘surviving’, mammary epithelial cell; with cell interaction with laminin-rich ECM being shown for the first time to be central to this repression; and (3) That Brca1 expression in populations of mammary epithelial cells committed to apoptosis may differ depending on the cells physiological status: where the apoptotic population has been previously associated with ECM no Brca1 expression is seen; where the apoptotic population has not been associated with matrix Brca1 expression is maintained.
Discussion
The data reported here for both the in vivo and cell culture studies suggest that Brca1 expression is primarily linked to mammary epithelial cell proliferation. In vivo, expression is seen in the virgin animals, during a period reported to exhibit active epithelial cell proliferation in the terminal end buds, structures associated with the establishment of the mature mammary tree.62,63 Expression was also seen in early and late pregnancy; the former period associated with intense branched ductal expansion, the latter with alveologenesis and differentiation. Additionally, expression is reported for the first time during the final proliferative pulse,51 at the beginning of lactation. Finally, we show Brca1 expression in primary cultures of mid-pregnant mammary epithelial cells proliferating in the presence of foetal calf serum and EGF (Figure 2C). The pattern of Brca1 expression reported here for the mouse mammary gland is similar (but extends to early lactation) that reported by Marquis et al37 and Rajan et al47 and parallels the expression pattern reported for Brca2.39,47 These authors have also reported Brca1 expression in proliferating mammary epithelial HC11 cells.37,47 Particularly interesting is the finding in this and the latter studies that Brca1 expression would seem to accompany mammary epithelial cell proliferation whenever it occurs, and it occurs in the gland under a range of differing humoral and local influences (i.e. in the TEB of the virgin, during early and late pregnancy, and early in lactation). That the Brca1 expressed in the mammary gland during periods of avid proliferation is absolutely required for mammary gland development has been demonstrated by Xu et al64 who selectively restricted full-length Brca1 isoform production in pregnant mammary gland using the Cre–loxP recombination system under the control of the Wap and MMTV promoters. Without the full-length Brca1 isoform, mammary tissue failed to fully develop during pregnancy: The ductal tree formed incompletely and failed to fully penetrate the mammary fat pad and there was a frequent increase in apoptosis. The authors felt that cellular loss of full-length Brca1 isoform deprived the cells of a crucial genome ‘caretaker’. This would result in frequent genetic alterations and cell loss by apoptosis, triggered by ‘gatekeepers’ such as p53. That these animals also succumbed to mammary tumours strengthened their case. The Brca1 molecular machinery may also contribute significantly to alveologenesis: This is suggested by the observation that BARD-1-deficient mammary epithelial cell clones lost the capacity to form luminal structures in three-dimensional collagen gels. BARD1 (BRCA1-associated RING domain protein) was identified as a protein which functionally interacts with BRCA1.20
Brca1 expression in both model systems studied was repressed in the fully differentiated cells (Figures 1B and 2C); with cell interaction with the laminin-rich ECM being most likely responsible for this repression. The association of the mammary epithelial cell with the laminin-rich ECM through occupation of β1-integrin receptors provides a strong survival signal to the mammary epithelial cell56,57 at least in part through activating an insulin/IGF-1 modulated anti-apoptotic PI-3 kinase/Akt transduction cascade.65,66,67 Mammary epithelial cells in primary culture require both ECM-association and insulin (or IGF-1) for survival.56,57 Farrelly et al61 have demonstrated the ECM-survival stimulus to act through insulin/IGF-1-regulated PI 3 kinase pathway-dependent signalling. The pathway would presumably regulate the activity of pro- or anti-apoptotic members of the Bcl 2 family of proteins. Additionally, ECM-cell contacts are suggested to affect nuclear shape (and gene expression) by determining stress fibre tensility through the formation of focal adhesions.68 This may impact on rates of transcription of specific genes. For instance, it has been possible to define elements within the promoters of the β-casein and TGF-β genes that are ECM-sensitive, the former mediating a positive response,69,70 the latter a negative response71 to mammary epithelial cell-ECM association. Perhaps it is by such a mechanism that ECM suppression of Brca1 expression occurs.
A comparison of Brca1 expression in the two model systems studied revealed that Brca1 expression in populations of mammary epithelial cells committed to apoptosis seems to differ depending on the cell populations physiological status. In the population of fully differentiated alveolar epithelial cells which predominate in lactation we observed that Brca1 expression becomes repressed after the final wave of proliferation of early lactation (Figure 1B). This is most likely regulated by ECM-cell interaction (Figure 2C). In this, the population of cells lost by apoptosis in early involution, Brca1 expression remains suppressed. It is interesting that while the expression of a variety of cell cycle/ G1 associated genes become activated in early involution-associated apoptosis52,72,73 Brca1 expression remains repressed. Although there is no gross destruction of ECM as yet in the early involuting gland that exhibits high levels of apoptosis (e.g. the two day involuting gland52), it is likely that there might be change in the status of laminin-integrin receptor interactions at this time, thus, it is not clear that ECM-cell associations are responsible for maintaining this repression of Brca1 expression. The mammary gland of early parturition of the conditional Brca1 knockout mouse exhibits high levels of apoptosis.64 This parallels our findings in early involution and might support a hypothesis that the Brca1-minus state is ‘pro-apoptotic’ for differentiated mammary alveolar epithelial cells. However, the increased apoptosis seen in the knockout mouse at early parturition might reflect the clearing of a population of cells with DNA-damage that arose during the last wave of ‘proliferation without an intact cell cycle surveillance system.’ In the cell culture model (Figure 2) Brca1 expression persisted after cell proliferation has ceased, in a population of cells that are quiescent, exposed to ‘lactogenic hormones (prolactin, hydrocortisone and the required survival factor, insulin) but are undergoing extensive apoptosis because of a lack of association with a suitable laminin-rich ECM. As the Brca1 expression measurements were made 48 h after removal of the growth factors (serum and EGF) we feel the persistent Brca1 expression is not residual to the effects of growth factor exposure during proliferation. Other growth factor responses, e.g. Erk1/Erk2 activation, are lost within 6 h of growth factor removal (D Finlay and F Martin, unpublished observation). It is important to note that the cell culture experiment (Figure 2) does not model a transition of mammary epithelial cells from differentiated to apoptotic state (equivalent to the lactation to involution transition) but rather demonstrates the rescue by suitable ECM-association of a cell population that will otherwise die due to lack of this survival influence. This experiment shows that apoptosis of mammary epithelial cells can also occur in the presence of Brca1 expression, but, in cells in which Brca1 expression has not previously been repressed by terminal differentiation. This apoptotic population expressing Brca1, were it being considered to directly reflect a physiological population may parallel those cells in the terminal end-bud of the young adult virgin gland which when ‘non-differentiated’ in the bud, die by apoptosis while another population proliferate and survive to contribute to branching of the mammary tree.63 Interestingly these latter cells would die at a time when the glandular Brca1 expression is high (Figure 1B). Thus, our study suggests that Brca1 expression is most likely associated with proliferation in the mammary gland; it may be permitted in the dying population of the terminal end bud but it is finally repressed in the gland at lactation which is quantitatively dominated by a population of alveolar epithelial cells most of which are not again going to re-enter the cell cycle but will die by apoptosis.
Materials and Methods
Mammary tissue dissected from virgin (5-week-old), pregnant, lactating and weaned CD1 mice was snap frozen in liquid nitrogen and stored at −70°C for subsequent RNA preparation. A fourth (inguinal) mammary gland was used for methacarn fixed, carmalum stained whole mount preparation, as described by Edwards et al.50 Primary mammary epithelial cell cultures59,74,75 were prepared from mammary glands from 14–19 day pregnant CD-1 mice, as recently described.57 After growing under ‘proliferating’ conditions for 48 h, populations of ‘apoptotic cells (‘minus ECM’)’, cultured in the absence of serum and EGF and the absence of ECM (Matrigel) overlay but in the presence of lactogenic hormones and ‘surviving/differentiated cells (‘plus ECM’)’ cultured in the absence of serum and EGF, but in the presence of ECM (EHS tumour-derived ECM, Matrigel (Becton Dickinson)) overlay and lactogenic hormones, were prepared.57 These were subsequently harvested for FACS-cell cycle analysis, FACS/TUNEL and total RNA isolation and RT–PCR analysis. FACS-cell cycle analysis was carried out as recently described.57
For FACS/TUNEL76 analysis cultured cells were recovered from the medium by centrifugation or gently scraped off into PBS and pelleted. They were resuspended in a 1% solution of formaldehyde in PBS and fixed on ice for 15 min, washed in PBS, resuspended in ice-cold 70% ethanol and left at 4°C for at least 30 min. The cells were washed in PBS and resuspended in 50 μL TdT reaction medium (37.8 μL H2O; 5 μL 10×TdT buffer (300 mM Tris base, 1.4 M sodium cacodylate (pH 7.2)) and 10 mM cobalt chloride); 5 μL cobalt chloride (25 mM); 2 μL biotinylated-16-dUTP (1 nM) and 0.2 μL TdT (25 U.μL−1). The samples were incubated at 37°C for 30 min, then washed in PBS and resuspended in staining buffer (54.2 μL MilliQ water; 25 μL 20×SSC; 20 μL 25% BLOTTO; 0.7 μL FlTC-conjugated Extravidin (Sigma) and 0.1 μL Triton X-100) and stained in the dark at 37°C for 30 min. The cells were then washed and stored in PBS at 4°C, in the dark, until analyzed in a FACStar Plus flow cytometer. Total RNA was prepared by the method of Chomczynski and Sacchi77 from snap frozen mouse mammary tissue and primary cultured cells, treated as outlined above. Aliquots of total RNA were treated with RNase-free DNase I (Boehringer Mannheim) and the RNA concentration determined spectrophotometrically. 2–10 μg of DNase I-treated total RNA was transcribed with Superscript™ II RNase H-minus reverse transcriptase (GibCo BRL) in the presence of 100 ng–1 μg of the appropriate reverse primer. Ten per cent of the first strand cDNA synthesis reaction was used as template for PCR in a Taq catalysed amplification using the appropriate primer pairs: for Brca1, fwd: 5′ TCGGCGCTTGCAAGTACGGATCT 3′ ; rev: 5′ AAGGTTAGACAGCTGGACACCTA 3′ (see Figure 1); for β-casein, fwd: 5′ GCCTTGCCAGTCTTGCTA 3′ ; rev: 5′ GGAATGTTGTGAGTGGC 3′;78 for WAP, fwd: 5′ AAAAGCCAGCCCCATTGA 3′; rev: 5′ AGGGTTATCACTGGCACT 3′;78 for Bax, fwd: 5′ GGTTTCATCCAGGATCGAGCAG 3′; rev: 5′ CCGTCTGGCACTGGTAGA 3′;56 for gpdh, fwd: 5′ CCTTCATTGACCTCAACTAC 3′; rev: 5′ TTCACACCCATCACAAAC 3′;78 and for neu, fwd: 5′ CGGAACCCACATCAGGCC 3′; rev: 5′ TTTCCTGCAGCAGCCTAC 3′.79 Taq polymerase addition was witheld until after a ‘hot start’ cycle, 95°C for 2 min and cooling to 80°C after which the enzyme was added. Each PCR cycle consisted of a denaturing step at 95°C for 30 s, an annealing step at 52–60°C (optimised for individual primer pairs) for 45 s and a primer elongation step at 72°C for 80 s. Depending on the abundance of target mRNA present in the starting sample, 18–40 cycles of PCR amplification were carried out.
Abbreviations
- BRCA1:
-
breast cancer susceptibility gene 1
- ECM:
-
extracellular matrix
- FACS/TUNEL:
-
fluorescence activated cell sorting/TdT-mediated biotinylated dUTP nick end-labelling
- IGF:
-
insulin-like growth factor
- PI:
-
propidium iodide
- RT–PCR:
-
reverse transcriptase-polymerase chain reaction
- TdT:
-
terminal deoxynucleotidyl transferase
- WAP:
-
whey acidic protein
References
Miki Y, Swensen J, Shattuck-Evans D, Futreal AP, Harshman K, Tavtigian S, Liu Q, Cochran C, Bennett ML, Ding W, Bell R, Rosenthal J, Hussey C, Tran T, McClure M, Frye C, Hattier T, Phelps R, Haugen-Strano A, Katcher H, Yakumo K, Gholami Z, Shaffer D, Stone S, Bayer S, Wray C, Bogden R, Dayananth P, Ward J, Tonin P, Narod S, Bristow PK, Norris FH, Helvering L, Morrison P, Rosteck P, Lai M, Barrett C, Lewis C, Neuhausen S, Cannon-Albright L, Goldgar D, Wiseman R, Kamb A and Skolnick MH . (1994) A strong candidate for the breast and ovarian cancer susceptibility gene BRCA1. Science 266: 66–71
Sharan SK, Wims M and Bradley A . (1995) Murine Brca1: sequence and significance for human missense mutations. Hum. Mol. Genet. 4: 2275–2278
Abel KJ, Xy J, Yin GY, Lyons RH, Meisler MH, and Weber BL, . (1995) Mouse Brca1: localization sequence analysis and identification of evolutionary conserved domains. Hum. Mol. Genet. 4: 2265–2273
Chen Y, Chen C-F, Riley DJ, Allred DC, Chen P-L, von Hoff D, Osborne CK and Lee W-H . (1995) Aberrant subcellular localisation of BRCA1 in breast cancer. Science 270: 789–791
Chen CF, Li S, Chen Y, Chen PL, Sharp DZ and Lee WH . (1996) The nuclear localization sequences of the BRCA1 protein interact with the importin-α subunit of the nuclear transport signal. J. Biol. Chem. 271: 32863–32868
Chen Y, Chen P-L, Riley DJ, Lee W-H, Allred DC, and Osborne CK . (1996) Response to Scully et al. Science 272: 125–126
Scully R, Ganesan S, Brown M, de Caprio JA, Cannistra SA, Feuteun J, Schnitt S and Livingston DM . (1996) Location of BRCA1 in human breast and ovarian cancer cells. Science 272: 123–125
Thomas JE, Smith M, Rubinfeld B, Gutowski M, Beckmann RP and Polakis P . (1996) Subcellular localisation and analysis of apparent 180-KDa and 220-kDa proteins of the breast cancer susceptibility gene, BRCA1. J. Biol. Chem. 271: 28630–28635
Coene E, van Oostveld P, Willems K, van Emmelo J and de Potter CR . (1997) BRCA1 is localised in cytoplasmic tube-like invaginations in the nucleus. Nat. Genet. 16: 122–124
Jensen RA, Thompson ME, Jetton TL, Szabo CI, van der Meer R, Helou B, Tronick SR, Page DL, King M-C and Holt J. . (1996) BRCA1 is secreted and exhibits properties of a granin. Nat. Genet. 12: 303–308
Bernard-Gallon DJ, Peffault de Latour M, de Olivera F, Rio P, Hizel C, Favy D, Bignon YJ . (1998) BRCA1 is localized in ductal secretions of normal breast tissues: detection by antibodies raised against the amino terminus of BRCA1 (K-18). Int. J. Cancer 77: 803–806
Bernard-Gallon DJ, Dechelotte P, Rio PG and Bignon YJ . (1999) BRCA1 and BRCA2 proteins are expressed in milk fat globules. Int. J. Cancer 81: 839–843
Bertwistle D and Ashworth A . (1998) Functions of the BRCA1 and BRCA2 genes. Curr. Opin. Genet. Dev. 8: 14–20
De Potter CR, Coene ED and Schelfhout VRJ . (1998) Localisation of BRCA 1 protein at the cellular level. J. Mammary Gland Biol. and Neoplasia 3: 423–430
Gudas JM, Li T, Nguyen H, Jensen D, Rauscher III FJ and Cowan KH . (1996) Growth retardation and tumour inhibition by BRCA1. Nat. Genet. 12: 298–302
Lu M, Conzen SD, Cole CN and Arrick BA . (1996) Characterisation of functional messenger RNA splice variants of BRCA1 expressed in nonmalignant and tumour-derived breast cells. Cancer Res. 56: 4578–4581
Thakur S, Zhang HB, Peng Y, Le H, Carroll B, Ward T, Yao J, Farid LM, Couch FJ, Wilson RB and Weber BL. . (1997) Localization of BRCA1 and a splice variant identifies the nuclear localization signal. Mol. Cell. Biol. 17: 444–452
Wilson CA, Payton MN, Elliot GS, Buaas FW, Cajulis EE, Groshaans D, Ramos L, Reese DM, Slamon DJ and Calzone FJ . (1997) Differential subcellular localisation, expression and biological toxicity of BRCA 1 and the splice variant BRCA 1-Δ 11b. Oncogene 14: 1–16
Saurin AJ, Borden KL, Boddy MN and Freemont PS . (1996) Does this have a familiar RING? Trends Biochem. Sci. 21: 208–214
Wu LC, Wang ZW, Tsan JT, Spillman MA, Phung A, Xu XL, Yang M-CW, Hwang L-Y, Bowcock AM and Baer R . (1996) Identification of a RING protein that can interact in vivo with the BRCA1 gene product. Nat. Genet. 14: 430–440
Chapman MS and Verma IM . (1996) Transcriptional activation by BRCA1. Nature 382: 678–679
Monteiro A, August A and Hanafusa H . (1996) Evidence for a transcriptional activation function of BRCA1 C-terminal region. Proc. Natl. Acad. Sci. USA 93: 13595–13599
Milner J, Ponder B, Hughes-Davies L, Seltmann M and Kouzarides T . (1997) Transcriptional activation functions in BRCA2. Nature 386: 772–773
Holt JT, Thompson ME, Szabo C, Robinson-Benion C, Arteaga CL, King MC and Jensen RA . (1997) Transcriptional activation functions in BRCA2. Nature 386: 772–773
Hu YF, Hao ZL and Li R . (1999) Chromatin remodelling and activation of chromosomal DNA replication by an acidic transcriptional activation domain from BRCA1. Genes Dev. 13: 637–642
Yarden RI and Brody LC . (1999) BRCA1 interacts with components of the histone deacetylase complex. Proc. Natl. Acad. Sci. USA 96: 4983–4988
Scully R, Anderson SF, Chao DM, Wei W, Ye L, Young RA, Livingston DM and Parvin JD . (1997) BRCA1 is a component of the RNA polymerase II holoenzyme. Proc. Natl. Acad. Sci. USA 94: 5605–5610
Anderson SF, Schlegel BP, Nakajima T, Wolpin ES and Parvin JD . (1998) BRCA1 protein is linked to the RNA polymerase II holoenzyme comples via RNA helicase A. Nat. Genet. 19: 254–256
Gowen LC, Avrutskaya AV, Latour AM, Koller BH and Leadon SA . (1998) BRCA1 required for transcription-coupled repair of oxidative DNA damage. Science 281: 1009–1012
Abbott DW, Thompson ME, Robinson-Benion C, Tomlinson G, Jensen RA and Holt JT . (1999) BRCA1 expression restores radiation resistance in BRCA1-defective cancer cells through enhancement of transcription-coupled DBA repair. J Biol. Chem. 274: 18808–18812
Scully R, Chen J, Plug A, Xiao Y, Weaver D, Feunteun J, Ashley T and Livingston D . (1997) Association of BRCA1 with Rad51 in mitotic and meiotic cells. Cell 88: 265–275
Scully R, Chen J, Ochs RL, Keegan K, Hoekstra M, Feunteun J and Livingston DM . (1997) Dynamic changes of BRCA1 subnuclear location and phophorylation state are initiated by DNA damage. Cell 90: 425–435
Zhong Q, Chen CF, Li S, Chen Y, Wang CC, Xiao J, Chen PL, Sharp ZD and Lee WH . (1999) Association of BRCA1 with the hRad50-hMre11-p95 complex and the DNA damage response. Science 285: 747–750
Chen JJ, Silver D, Cantor S, Livingston DM and Scully R . (1999) BRCA1, BRCA2 and Rad51 operate in a common DNA damage response pathway. Cancer Res. 59: (Suppl.) 1752s–1756s
Edelmann W and Kucherlapati R . (1996) Role of recombination enzymes in mammalian cell survival. Proc. Natl. Acad. Sci. USA 93: 6225–6227
Hsu LC and White RL . (1998) BRCA1 is associated with the centrosome during mitosis. Proc. Natl. Acad. Sci. USA 95: 12983–12988
Marquis ST, Rajan JV, Wynshaw-Boris A, Xu J, Yin G-Y, Abel KJ, Weber BL and Chodosh LA . (1995) The developmental pattern of Brca1 expression implies a role in differentiation of the breast and other tissues. Nat. Genet. 11: 17–26
Ludwig T, Chapman DL, Papaioannou VE and Efstratiadis A . (1997) Targeted mutation of breast cancer susceptibility gene homologues in mice: lethal phenotypes of Brca1, Brca2, Brca1/Brca2, Brca1/p53 and Brca2/p53 nullizygous embryos. Genes Dev. 11: 1226–1241
Rajan JV, Marquis ST, Gardner HP and Chodosh LA . (1997) Developmental expression of Brca2 colocalizes with Brca1 and is associated with proliferation and differentiation in multiple tissues. Dev. Biol. 184: 385–401
Gowen LC, Johnson BL, Latour AM, Sulik KK and Koller BH . (1996) Brca1 deficiency results in early embryonic lethality characterized by neuroepithelial abnormalities. Nat. Genet. 12: 191–194
Hakem R, de la Pompa JL, Sirard C, Mo R, Woo M, Hakem A, Wakeham A, Potter J, Reitmar A, Billia F, Firpo E, Hui CC, Roberts J, Rossant J and Mak TW . (1996) The tumour suppressor gene Brca1 is required for embryonic cellular proliferation in the mouse. Cell 85: 1009–1023
Liu C-Y, Flesken-Nikitin A, Li S, Zeng Y and Lee W-H . (1996) Inactivation of the mouse Brca1 gene leads to failure in the morphogenesis of the egg cylinder in early postimplantation development. Genes Dev. 10: 1835–1843
Suzuki A, de la Pompa JL, Hakem R, Elia A, Yoshida R, Mo R, Nishina H, Chuang T, Wakeham A, Itie A, Koo W, Billia P, Ho A, Fukumoto M, Hui CC and Mak TW . (1997) Brca2 is required for embryonic cellular proliferation in the mouse. Genes. Dev. 11: 1242–1252
Lane TF, Deng C, Elson A, Lyu M, Kozak CA and Leder P . (1995) Expression of Brca1 is associated with terminal differentiation of ectodermally and mesodermally derived tissue in mice. Nat. Genet. 9: 2712–2722
Thompson ME, Jensen RA, Obermiller PS, Page DL and Holt JT . (1996) Decreased expression of BRCA1 accelerates growth and is often present during sporadic breast cancer progression. Nat. Genet. 9: 444–450
Holt JT, Thompson ME, Szabo C, Robinson-Benion C, Arteaga CL, King MC and Jensen RA . (1996) Growth retardation and tumour inhibition by BRCA1. Nat. Genet. 12: 298–302
Rajan JV, Wang M, Marquis ST and Chodosh LA . (1996) Brca2 is coordinately regulated with Brca1 during proliferation and differentiation in mammary epithelial cells. Proc. Natl. Acad. Sci. USA 93: 13078–13083
Vaughn JP, Cirisano FD, Huper G, Berchuck A, Futreal AP, Marks JR and Iglehart JD . (1996) Cell cycle control of BRCA2. Cancer Res. 56: 4590–4594
Vaughn JP, Davis PL, Jarboe MD, Huper G, Evans AC, Wiseman RW, Berchuck A, Iglehart JD, Futreal AP and Marks JR . (1996) BRCA1 expression is induced before DNA synthesis in both normal and tumour-derived breast cells. Cell Growth Differ. 7: 711–715
Edwards PA, Hiby SE, Papkoff J and Bradbury JM . (1992) Hyperplasia of mouse mammary epithelium induced by expression of the Wnt-1(int-1) oncogene in reconstituted mammary gland. Oncogene 7: 2041–2051
Knight CH and Peaker M . (1982) Mammary cell proliferation in mice during pregnancy and lactation in relation to milk yield. Q. J. Exp. Physiol. 67: 165–177
Strange R, Li F, Saurer S, Burkhardt A and Friis RR . (1992) Apoptotic cell death and tissue remodelling during mouse mammary gland involution. Development 115: 49–58
Talhouk RS, Bissell MJ and Werb Z . (1992) Coordinated expression of extracellular matrix-degrading proteins and their inhibitors regulates mammary epithelial function during involution. J. Cell Biol. 118: 1271–1282
Robinson GW, McKnight RA, Smith GH and Henighausen L . (1995) Mammary epithelial cells undergo secretory differentiation in cycling virgins but require pregnancy for the establishment of terminal differentiation. Development 121: 2079–2090
Streuli CH, Edwards GM, Delcomenne M, Whitelaw BA, Burdon TG, Schindler C and Watson CJ . (1995) Stat5 as a target for regulation by extracellular matrix. J. Biol. Chem. 270: 21639–21644
Pullan S, Wilson J, Metcalfe A, Edwards GM, Goberdhan N, Tilly J, Hickman JA, Dive C, Streuli CH . (1996) Requirement of basement membrane for the suppression of programmed cell death in mammary epithelium. J. Cell Sci. 109: 631–642
Furlong EEM, Keon NK, Thornton F, Rein T and Martin F . (1996) Expression of a 74kD NF-1 protein is induced in mouse mammary gland involution. Involution enhanced occupation of a twin NF-1 binding element in the TRPM-2/clusterin promoter. J. Biol. Chem. 271: 29688–29697
Finlay D, Healy V, Furlong F, O'Connell FC, Keon NK and Martin F . (2000) MAP kinase pathway signalling is essential for extracellular matrix determined mammary epithelial cell survival. Cell Death Differ. in press
Streuli CH, Bailey N and Bissell MJ . (1991) Control of mammary epithelial differentiation: basement membrane induces tissue-specific gene expression in the absence of cell-cell interaction and morphological polarity. J. Cell Biol. 115: 1383–1395
Streuli CH, Schmidhauser C, Bailey N, Yurchenco P, Skubitz AP, Roskelley C and Bissell MJ . (1995b) Laminin mediates tissue-specific gene expression in mammary epithelia. J. Cell Biol. 129: 591–603
Farrelly N, Lee YJ, Oliver J, Dive C and Streuli CH . (1999) Extracellular matrix regulates Apoptosis in mammary epithelium through a control on insulin signaling. J. Cell Biol. 144: 1337–1348
Williams JM and Daniel CW . (1983) Mammary ductal elongation: differentiation of myoepithelium and basal lamina during branching morphogenesis. Dev. Biol. 97: 274–290
Humphreys RC, Krajewska M, Krnacik S, Jaeger R, Weiher H, Krajewski S, Reed JC and Rosen JM . (1996) Apoptosis in the terminal endbud of the murine mammary gland: a mechanism of ductal morphogenesis. Development 122: 4013–4022
Xu X, Wagner K-U, Larson D, Weaver Z, Li C, Ried T, Hennighausen L, Wynshaw-Boris A and Deng C-X . (1999) Conditional mutation of Brca1 in mammary epithelial cells results in blunted ductal morphogenesis and tumour formation. Nat. Genet. 22: 37–43
Datta SR, Dudek H, Tao X, Masters S, Fu H, Gotoh Y and Greenberg ME . (1997) Akt phosphorylation of BAD couples survival signals to the cell-intrinsic death machinery. Cell 91: 231–241
Gajewski TF and Thompson CB . (1996) Apoptosis meets signal transduction: elimination of a BAD influence. Cell 87: 589–592
Khwaja A, Rodriguez-Viciana P, Wennstrom S, Warne PH and Downward J . (1997) Matrix adhesion and Ras transformation both activate a phosphoinositide 3-OH kinase and protein kinase B/Akt cellular survival pathway. EMBO J. 16: 2783–2793
Roskelley CD, Desprez PY and Bissell MJ . (1994) Extracellular matrix-dependent tissue-specific gene expression in mammary epithelial cells requires both physical and biochemical signal transduction. Proc. Natl. Acad. Sci. USA 91: 12378–12382
Schmidhauser C, Casperson GF, Myers CA, Sanzo KT, Bolten S and Bissell MJ . (1992) A novel transcriptional enhancer is involved in the prolactin- and extracellular matrix-dependent regulation of beta-casein gene expression. Mol. Biol. Cell 3: 699–709
Myers CA, Schmidhauser C, Mellentin-Michelotti J, Fragoso G, Roskelley CD, Casperson G, Mossi R, Pujuguet P, Hager G and Bissel MJ . (1998) Characterization of BCE-1, a transcriptional enhancer regulated by prolactin and extracellular matrix and modulated by the state of histone acetylation. Mol. Cell. Biol. 18: 2184–2195
Streuli CH, Schmidhauser C, Kobrin M, Bissell MJ and Derynck R . (1993) Extracellular matrix regulates expression of the TGF-beta 1 gene. J. Cell Biol. 120: 253–260
Marti A, Jehn B, Costello E, Keon N, Ke G, Martin F and Jaggi R . (1994) AP-1 (cFos/JunD) is induced during apoptosis of mouse mammary epithelial cells. Oncogene 9: 1213–1223
Marti A, Feng Z, Jehn B, Djonov V, Chicaiza G, Altermatt H and Jaggi R . (1995) Expression and activity of cell cycle regulators during proliferation and programmed death in the mammary gland. Cell Death Differ. 2: 277–283
Barcellos-Hoff MH, Aggeler J, Ram TG, and Bissell MJ . (1989) Functional differentiation and alveolar morphogenesis of primary mammary cultures on reconstituted basement membrane. Development 105: 223–235
Streuli CH and Bissell MJ . (1990) Expression of extracellular matrix components is regulated by substratum. J. Cell Biol. 110: 1405–1415
Gavrieli Y, Sherman Y and Ben-Sasson SA . (1992) Identification of programmed cell death in situ via specific labelling of nuclear DNA fragmentation. J. Cell Biol. 119: 493–501
Chomczynski P and Sacchi N . (1987) Single-step method of RNA isolation by acid guanidinium thiocyanate-phenol-chloroform extraction. Anal. Biochem. 162: 156–159
Hathaway HJ and Shur BD . (1996) Mammary gland morphogenesis is inhibited in transgenic mice that overexpress cell surface β 1,4-galactosyltransferase. Development 122: 2859–2872
Siegel PM, Dankort DL, Hardy WR and Muller WJ . (1994) Novel activating mutations in the neu proto-oncogene involved in induction of mammary tumors. Mol. Cell. Biol. 14: 7068–7077
Acknowledgements
We acknowledge financial support from Enterprise Ireland/Forbairt, The Irish Science and Technology Agency and from the Health Research Board, Ireland.
Author information
Authors and Affiliations
Corresponding author
Additional information
Edited by L Schwartz
Rights and permissions
About this article
Cite this article
O'Connell, F., Martin, F. Laminin-rich extracellular matrix association with mammary epithelial cells suppresses Brca1 expression. Cell Death Differ 7, 360–367 (2000). https://doi.org/10.1038/sj.cdd.4400647
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/sj.cdd.4400647
Keywords
This article is cited by
-
Tissue extracellular matrix hydrogels as alternatives to Matrigel for culturing gastrointestinal organoids
Nature Communications (2022)
-
CITED1 homozygous null mice display aberrant pubertal mammary ductal morphogenesis
Oncogene (2006)
-
Regulation of BRCA1 expression and its relationship to sporadic breast cancer
Breast Cancer Research (2002)