Abstract
Many cells die during mammalian development and are engulfed by macrophages. In DNase II−/− embryos, the TUNEL-positive DNA of apoptotic cells is left undigested in macrophages, providing a system for studying programmed cell death during mouse development. Here, we showed that an Apaf-1-null mutation in the DNase II−/− embryos greatly reduced the number of macrophages carrying DNA at E11.5. However, at later stages of the embryogenesis, a significant number of macrophages carrying undigested DNA were present in Apaf-1−/− embryos, indicating that cells died and were engulfed in an Apaf-1-independent manner. In most tissues of the Apaf-1−/− embryos, no processed caspase-3 was detected, and the DNA of dead cells accumulated in the macrophages appeared intact. Many nonapoptotic dead cells were found in the tail of the Apaf-1−/− embryos, suggesting that the Apaf-1-independent programmed cell death occurred, and these dead cells were engulfed by macrophages. In contrast, active caspase-3 was detected in E14.5 thymus of Apaf-1−/− embryos. Treatment of fetal thymocytes with staurosporine, but not etoposide, induced processing of procaspases 3 and 9, indicating that the E14.5 thymocytes have the ability to undergo caspase-dependent apoptosis in an Apaf-1-independent manner. Thus, programmed cell death in mouse development, which normally proceeds in an efficient Apaf-1-depenent mechanism, appears to be backed up by Apaf-1-independent death systems.
Similar content being viewed by others
Main
Many useless and toxic cells are generated during animal development and removed by programmed cell death, which is mainly mediated by apoptosis.1, 2 Apoptosis is accompanied by morphological changes, such as the blebbing and condensation of cells, loss of membrane symmetry, and nuclear condensation.3 Late in apoptosis, dying cells are fragmented into small ‘apoptotic bodies,’ which are recognized by phagocytes for engulfment. This morphological change in the cells and their rapid disposal are distinct from necrosis, in which cells swell, the plasma membranes are ruptured, and the cellular contents are believed to be released.
The signal transduction involved in apoptotic cell death has been intensively studied4, 5 and shown to proceed by two major pathways, an extrinsic and an intrinsic one. In the extrinsic pathway, signals from death receptors, such as Fas and TNF receptor, activate an initiator caspase, caspase-8, that activates downstream effector caspases, mainly caspase-3, leading to the cleavage of more than 300 cellular proteins.6, 7 One of the caspase-3 substrates is the inhibitor of caspase-activated DNase (CAD), and its cleavage by caspase-3 inactivates the ability to associate with CAD, allowing CAD to cause the fragmentation of chromosomal DNA into nucleosomal units.8 In the intrinsic pathway, death signals cause the release of cytochrome c from mitochondria through ‘BH3-only’ proteins of the Bcl-2 family.9, 10 The cytochrome c binds to Apaf-1 and activates caspase-9, which leads to the activation of caspase-3 and CAD, resulting in the cleavage of cellular substrates and the fragmentation of chromosomal DNA. Apoptotic cells that die by either the intrinsic or the extrinsic pathway expose phosphatidylserine on their surface.11 Macrophages and immature dendritic cells recognize the phosphatidylserine and rapidly engulf the apoptotic cells.12 The engulfed dead cells are transferred to lysosomes, where all their components are degraded.
The role of the intrinsic apoptotic pathway in programmed cell death has been studied by establishing mice that lack genes involved in apoptotic signal transduction.13, 14, 15, 16, 17 The results obtained with these knockout mice have not always been consistent among groups. For example, Yoshida et al.14 and Kuida et al.13 reported that Apaf-1 and caspase-9 are indispensable for the cell death induced by cytotoxic agents or γ-radiation in various tissues, whereas Marsden et al.18 reported that lymphocytes undergo apoptosis without Apaf-1 or caspase-9 on cytokine deprivation. Furthermore, except for craniofacial abnormalities and the hyperproliferation of neuronal cells, Apaf-1-deficient mice are apparently normal, especially in the C57BL/6 background,19 suggesting the presence of a back-up system for the Apaf-1-mediated apoptosis in mouse development.
Because dying cells are swiftly engulfed and degraded by macrophages, the detection of apoptotic cells in animals is not an easy task. DNase II is responsible for digesting the DNA of apoptotic cells after macrophages engulf them, and many macrophages carrying engulfed DNA are present in DNase II-deficient embryos.20, 21, 22 Here, we used DNase II-deficient mice to detect the programmed cell death that occurs during mouse development, and found that Apaf-1 seems to be dispensable for this process. In the thymus of Apaf-1−/− embryos at E14.5, caspases were activated in an Apaf-1-independent manner. In most other tissues, cells died by an Apaf-1-independent nonapoptotic mechanism, and were engulfed by macrophages. These results indicate that Apaf-1-independent systems, with or without caspase activation, function as back-up systems for the programmed cell death in mammalian development.
Results
Apaf-1-dependent programmed cell death in mouse embryos at E11.5
When we used the TUNEL staining to detect apoptotic cells in E11.5 wild-type mouse embryos, it yielded barely detectable signals (data not shown). A mutation of nuc-1, a homologue of DNase II, enhances the TUNEL positivity in C. elegans.23 Similarly, the mouse DNase II−/− embryos showed very strong TUNEL signals (Figure 1a), which is due to the accumulation of DNA fragmented by CAD.24 TUNEL-positive spots were frequent in several regions, such as the diencephalon at the cerebrum, the lamina terminalis, somites, and mesonephric ducts. Observations at higher magnification showed TUNEL positivity in these regions was much stronger in the DNase II−/− than the wild-type embryos. The TUNEL-positive spots in the DNase II−/− embryos were clustered into foci that were strongly DAPI positive (Figure 1b). The foci were located inside F4/80-positive macrophages, suggesting that they were the DNA of engulfed apoptotic cells. The engulfment of corpses in C. elegans requires the activation of caspase; a mutation in Ced-3 or Ced-4, homologues of caspase and Apaf-1, respectively, prevents the engulfment of cell corpses.25 Accordingly, a null mutation of Apaf-1 blocked the generation of TUNEL-positive foci (Figure 1a and b), indicating that the apoptotic cell death and engulfment of dead cells in the E11.5 mouse embryos were apparently Apaf-1 dependent.
Apaf-1-dependent programmed cell death in DNase II−/− mouse embryos at E11.5. (a) TUNEL staining of the E11.5 whole embryos. Paraffin sections from E11.5 Apaf-1+/−DNase II−/− and Apaf-1−/−DNase II−/− embryos were stained with TUNEL, followed by counterstaining with methyl green. Scale bar=500 μm. (b) TUNEL staining of various tissues from E11.5 embryos. Paraffin sections of the diencephalon, lamina terminalis, and somites from E11.5 Apaf-1+/−DNase II+/+, Apaf-1+/−DNase II−/−, and Apaf-1−/−DNase II−/− embryos, were stained with TUNEL, counterstained with methyl green, and observed by microscopy. For staining of F4/80, cryosections from E11.5 Apaf-1+/−DNase II−/− mouse embryos were stained with DAPI and a rat mAb against mouse F4/80, followed by incubation with peroxidase-conjugated rabbit anti-rat Ig, and detected by Cy3-labeled tyramide. Staining profiles with DAPI and anti-F4/80 are merged, and shown in the middle panels. Polarity of the embryos is indicated in left panels. a, anterior; p, posterior; d, dorsal; v, ventral. Yellow arrowheads indicate the macrophages carrying undigested DNA. Scale bar=100 μm
Apaf-1-independent programmed cell death in mouse embryos
Many TUNEL-positive foci were observed throughout the DNase II−/− embryos at E14.5 as well (data not shown). DNase II−/− embryos produce IFNβ which is lethal, but the lethality can be rescued by a deficiency of IFN type I receptor.26 To exclude the possible cell death caused by IFNβ, we carried out the TUNEL staining with DNase II−/−IFN-IR−/− embryos. Many TUNEL-positive foci still could be detected in various tissues of DNase II−/−IFN-IR−/− embryos (Figure 2a ; data not shown), suggesting that the death of the cells in DNase II−/− embryos was intrinsic to mouse embryogenesis, and not due to the secondary effect by IFNβ. The null mutation of Apaf-1 blocked the generation of TUNEL-positive foci in most tissues of the E14.5 DNase II−/−IFN-IR−/− embryos. For example, many TUNEL-positive foci were present at the interdigits of the hind paw of DNase II−/−IFN-IR−/− embryos, and activated caspase-3 could be detected in this region (Figure 2a and b). The deficiency of Apaf-1 prevented both the caspase activation and the appearance of TUNEL-positive foci. Probably due to the inhibition of apoptotic cell death, the formation of interdigits was retarded in the Apaf-1−/−DNase II−/−IFN-IR−/− embryos compared with Apaf-1+/+DNase II−/−IFN-IR−/− embryos, as reported for Apaf-1-null mice.14
Programmed cell death in the interdigit. (a and b) Programmed cell death at E14.5. Paraffin sections (a) or cryosections (b) of the hind paws from E14.5 Apaf-1+/+DNase II−/−IFN-IR−/− (upper) and Apaf-1−/−DNase II−/−IFN-IR−/− (lower) embryos were stained with DAPI and TUNEL (a), or with DAPI and an anti-processed caspase-3 mAb (b). Scale bar=100 μm. (c and d) Programmed cell death at E17.5. Cryosections of the hind paws from E17.5 Apaf-1+/+DNase II−/−IFN-IR−/− (left) and Apaf-1−/−DNase II−/−IFN-IR−/− (right) embryos were stained with DAPI (c). The boxed areas in (c) are enlarged in (d). Sections from E17.5 Apaf-1+/+DNase II−/−IFN-IR−/− (upper) and Apaf-1−/−DNase II−/−IFN-IR−/− (lower) mouse embryos were stained with DAPI and a rat mAb against mouse F4/80, followed by incubation with peroxidase-conjugated rabbit anti-rat Ig, and detected by Cy3-labeled tyramide. The staining profiles with DAPI and anti-F4/80 are merged in the right panels. Arrowheads indicate macrophages carrying engulfed DNA. Scale bars=50 μm (d). (e) Electron microscopy. Sections from the E17.5 hind paws of Apaf-1+/+DNase II−/−IFN-IR−/− (upper) and Apaf-1−/−DNase II−/−IFN-IR−/− (lower) mice were analyzed by electron microscopy. Macrophages are indicated by yellow lines. The areas surrounded by black and red boxes are enlarged in the middle and right panels. N, nuclear DNA from the engulfed dead cells. Scale bar=1 μm
At E17.5, fewer DAPI-positive foci were detected in the Apaf-1+/+DNase II−/−IFN-IR−/− embryos than at E14.5, as seen, for example, in the interdigits (Figure 2c), which have completely formed by E17.5. In contrast, the interdigits of the Apaf-1−/−DNase II−/−IFN-IR−/− embryos carried more DAPI-positive foci at E17.5 than at E14.5, and these DAPI-positive materials were located within F4/80-positive macrophages (Figure 2d). Electron microscopy showed that the DNA accumulated in the macrophages at the interdigits of the E17.5 Apaf-1+/+DNase II−/−IFN-IR−/− embryos is fragmented, whereas the DNA in the Apaf-1−/−DNase II−/−IFN-IR−/− embryos was apparently intact (Figure 2e). The active caspase-3 could be detected in the interdigits of Apaf-1+/+ but not Apaf-1−/− embryos (data not shown). These results suggested that in late embryogenesis, the cells in the interdigits could die in an Apaf-1-independent manner, and the dead cells were engulfed by macrophages.
Apaf-1-independent nonapoptotic cell death
At the ventral ectodermal ridge of the tail in mouse embryos, many cells undergo programmed cell death, which was detected as DAPI-positive foci in the DNase II−/−IFN-IR−/− embryos at E14.5 (Figure 3a and b). In the Apaf-1+/+ embryos, the DNA that accumulated in the ridge was TUNEL positive, and many processed caspase-3-positive cells could be detected in this region. On the other hand, the DNA that accumulated in the Apaf-1−/− embryos was TUNEL negative, and processed caspase-3-positive cells were hardly detectable. Because the DAPI-positive foci were relatively closely clustered in the ventral ectodermal ridge of the tail (Figure 3c), we analyzed this region by electron microscopy. As shown in Figure 3d, many unengulfed, nonapoptotic dying cells were detected in the E14.5 Apaf-1−/− embryos. That is, the unengulfed dying cells showed mottled chromatin condensation, nuclear membrane detachment and rupture, and dilated mitochondria. Approximately 5% of the cells in the tail ridge of E14.5 Apaf-1−/− embryos had the nonapoptotic morphology (Figure 3e), but apoptotic dying cells were hardly detectable. Nonapoptotic dead cells in the tail ridge of Apaf-1+/+ embryos were rare, about 1%, but cells with apoptotic characteristics, such as condensed and fragmented nuclei, were relatively abundant (1.72%). These results indicated that the cells in the tail ridge underwent nonapoptotic cell death in the absence of Apaf-1, and that these dead cells were engulfed by macrophages.
Apaf-1-independent nonapoptotic cell death. (a–c) Programmed cell death in the tail. Paraffin sections (a) or cryosections (b) of the tail from E14.5 Apaf-1+/+DNase II−/−IFN-IR−/− (upper) and Apaf-1−/−DNase II−/−IFN-IR−/− (lower) embryos were stained with DAPI and TUNEL (a), and with DAPI and a rabbit mAb against processed caspase-3 (b). Scale bar=100 μm. In (c), a transverse section of the tail from an Apaf-1+/−DNase II−/−IFN-IR−/− embryo at E14.5 was stained with toluidine blue. Scale bars=100 μm. (d) Electron microscopy. Sections from the E14.5 tail of Apaf-1+/+DNase II+/−IFN-IR−/− (left) and Apaf-1−/−DNase II+/+IFN-IR−/− (middle and right) embryos were analyzed by electron microscopy. Apoptotic cells in the Apaf-1+/+ embryos (left) and nonapoptotic dead cells (middle and right) in the Apaf-1−/− embryos are shown. Arrowheads indicate dilated mitochondria. Scale bar=1 μm. (e) Increase in nonapoptotic dead cells in the Apaf-1−/− tail. Electron micrographs (55 × 55 μm) with low magnification of the ventral parts (boxed area in c) of the tail of Apaf-1+/+ or +/− and Apaf-1−/− embryos at E14.5 were assembled, and the percentages of apoptotic and nonapoptotic dead cells out of 950 cells were determined. Average numbers obtained from three embryos are plotted
Apaf-1-independent apoptosis in the thymus
Many TUNEL- and DAPI-positive foci were observed in the thymus of DNase II−/−IFN-IR−/− embryos at E14.5 (Figure 4a). In contrast to most other tissues, such as the interdigits and tails at this stage of embryogenesis, the TUNEL-positive foci were present in the Apaf-1−/− thymus as well, where processed caspase-3 was also detected. Electron microscopy indicated that the DAPI-positive material in the E14.5 thymus was inside macrophages, and the DNA that accumulated in the macrophages was fragmented in both the Apaf-1+/+ and the Apaf-1−/− thymuses (Figure 4b). These results indicated that the thymocytes at E14.5 could activate caspase-3 and CAD using an Apaf-1-independent mechanism.
Programmed cell death in the fetal thymus. (a) Programmed cell death at E14.5. Paraffin sections (left) or cryosections (right) of the thymus from E14.5 Apaf-1+/+DNase II−/−IFN-IR−/− (upper) and Apaf-1−/−DNase II−/−IFN-IR−/− (lower) embryos were stained with DAPI and TUNEL using the digoxigenin/anti-digoxigenin fluorescein system (left), or with an anti-processed caspase-3 mAb (right). Scale bars=100 μm. (b) Electron microscopy of the E14.5 thymus. Sections from the thymus of E14.5 Apaf-1+/+DNase II−/−IFN-IR−/− (upper) and Apaf-1−/−DNase II−/−IFN-IR−/− (lower) embryos were analyzed by electron microscopy. Boxed areas were enlarged. N, nuclear DNA from the engulfed dead cells; MN, macrophage nuclei. Scale bar=1 μm. (c) Programmed cell death at E17.5. Adjacent cryosections of the thymus from E17.5 embryos with Apaf-1+/+DNase II−/−IFN-IR−/− (upper) and Apaf-1−/−DNase II−/−IFN-IR−/− (lower) embryos were stained with DAPI and TUNEL (left), or with an anti-processed caspase-3 mAb. Scale bars=100 μm. In the right, the sections were stained with DAPI and a rat mAb against mouse F4/80, and their staining profiles were merged. Yellow arrowheads indicate the macrophages carrying engulfed DNA. Scale bar=100 μm. (d) Electron microscopy of the E17.5 thymus. Sections from the E17.5 thymus of Apaf-1+/+DNase II−/−IFN-IR−/− (upper) and Apaf-1−/−DNase II−/−IFN-IR−/− (lower) embryos were analyzed by electron microscopy. Boxed areas in the left panels were enlarged in the adjacent right panels. N, nuclear DNA from the engulfed dead cells; MN, macrophage nuclei. Scale bars=2 μm
The E17.5 Apaf-1−/− thymus also carried DAPI-positive foci (Figure 4c). But, in contrast to the E14.5 Apaf-1−/− thymus, the TUNEL reactivity of the foci in the E17.5 Apaf-1−/−DNase II−/−IFN-IR−/− thymus was very weak, and processed caspase-3 was barely detected. Staining with F4/80 indicated that the DAPI-positive foci in the Apaf-1−/− thymus were inside macrophages (Figure 4c), and the electron microscopy showed that the DNA that accumulated in the macrophages was intact (Figure 4d). These properties of the E17.5 Apaf-1−/− thymus were different from those of the E14.5 thymus, but similar with those found in other tissues (interdigits, and tail) at E14.5 and 17.5.
Apaf-1-independent caspase activation
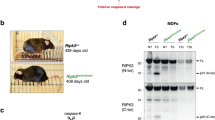
To examine whether thymocytes can undergo caspase-dependent apoptotic cell death in the absence of Apaf-1, we prepared thymocytes from the wild-type and Apaf-1−/− E14.5 embryos. They were treated with two different apoptosis inducers, etoposide and staurosporine, and the caspase-3 activity in the cell lysates was determined with a fluorescent substrate, Ac-DEVD-AMC. As shown in Figure 5a, a strong caspase-3 activity (about 5600 U) could be detected in the cell lysates from the wild-type but not Apaf-1-deficient thymocytes treated with 50 μM etoposide. The cell lysates from the thymocytes treated with 10 μM staurosporine also showed a caspase-3-activity (about 1300 U). Different from the treatment with etoposide, the cell lysates from the staurosporine-treated Apaf-1−/− thymocytes carried a significant caspase-3 activity (500 U). The active caspase-3 is often released from the cells into supernatant.27 In fact, the supernatant from staurosporine-treated Apaf-1+/+ and Apaf-1−/− thymocytes carried a high caspase-3 activity (10 000–15 000 U). Conversely, the treatment with etoposide produced the caspase-3 activity in the supernatant with Apaf-1+/+ thymocytes, but not with Apaf-1−/− thymocytes. These results indicated that etoposide could activate caspase-3 in E14.5 thymocytes in an Apaf-1-dependent manner, whereas Apaf-1 is dispensable for the staurosporine-induced activation of caspase-3. Similar results were obtained with E17.5 thymocytes, that is, staurosporine but not etoposide activated caspase-3 in E17.5 Apaf-1−/− thymocytes as efficiently as Apaf-1+/+ thymocytes, although the extent of the activated caspase-3 per cell is significantly lower than that found with E14.5 thymocytes (Figure 5a).
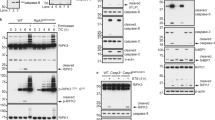
Apaf-1-independent caspase activation. (a) Activation of caspase-3. Thymocytes were prepared from E14.5 or E17.5 embryos with Apaf-1+/+ or Apaf-1+/− (C) and Apaf-1−/− (K) genotypes, and treated with 10 μM staurosporine or 50 μM etoposide for 4 h. The caspase activity in the cell lysates and supernatant was determined with Ac-DEVD-AMC as described in Materials and Methods. The caspase activity is expressed in an arbitrary unit. One ng recombinant human caspase-3 gave 14 100 U under the same conditions. Experiments were performed twice with different fetal thymus, and the average values are shown. (b) Processing of caspase-3. In left panel, thymocytes from E17.5 embryos with Apaf-1+/+ or Apaf-1+/− (C) and Apaf-1−/− (K) genotypes were treated with 10 μM staurosporine (STS) for 2 h or with 50 μM etoposide (ETO) for 4 h, and the cell lysates were subjected to western blotting with rabbit mAb against mouse caspase-3. The cell lysates from untreated thymocytes were also analyzed (0 h). Bands for the proform and cleaved form of caspase-3 are indicated by arrows. As a loading control, the membrane was re-blotted for BiP, and shown in the bottom. In right panel, thymocytes were treated for 4 h with staurosporine (STS) or etoposide (ETO). The supernatant was immunoprecipitated with rabbit mAb against the cleaved caspase-3, and subjected to western blotting with the same mAb, followed by incubation with HRP-conjugated Protein A. The bands for cleaved caspase-3 are indicated by an arrow. (c) Processing of caspase-9. Thymocytes from E17.5 embryos with Apaf-1+/+ or Apaf-1+/− (C) and Apaf-1−/− (K) genotypes were treated 10 μM staurosporine (STS) for 2 h or with 50 μM etoposide (ETO) for 4 h. The cell lysates from the STS- and ETO-treated and untreated (0 h) thymocytes were subjected to western blotting with mouse mAb against mouse caspase-9. Bands for the proform and cleaved form of caspase-9 are indicated by arrows. The membrane was re-blotted for BiP, and is shown in the bottom
To confirm the caspase-3 activity detected with Ac-DEVD-AMC is due to the processed caspase-3, we analyzed the cell lysates using western blot method with the antibody against caspase-3. As shown in Figure 5b, the lysates from the staurosporine-treated Apaf-1+/+ and Apaf-1−/− thymocytes showed a 17 kDa band for the processed caspase-3. This band was detected in the lysates from the etoposide-treated Apaf-1+/+ but not Apaf-1−/− thymocytes, and its intensity correlated with the caspase-3 activity detected (Figure 5a ). When the culture supernatants from staurosporine- or etoposide-treated E17.5 Apaf-1+/+ thymocytes were immunoprecipitated with the antibody against processed caspase-3, they showed the 17 kDa band by western blot with the antibody against the processed caspase-3 (Figure 5b). However, the 17 kDa band was not detected in the immunoprecipitate of the etoposide-treated E17.5 Apaf-1−/− thymocytes, which agrees with the little caspase-3 activity in the supernatant (Figure 5a).
In the intrinsic apoptotic pathway, Apaf-1 works as a scaffold to activate caspase-9, which induces processing of procaspase-3.10 To examine an involvement of caspase-9 in the Apaf-1-independent activation of caspase-3, we carried out western blot analysis with anticaspase-9. As shown in Figure 5c, the 37 kDa processed caspase-9 could be detected in the lysates from the staurosporine-treated Apaf-1+/+ as well as Apaf-1−/− thymocytes. In contrast, the processing of procaspase-9 in the etoposide-treated thymocytes was observed in the Apaf-1+/+ thymocytes, but not in the Apaf-1−/− thymocytes. These results indicate that staurosporine but not etoposide can activate caspase-9 in the absence of Apaf-1.
Discussion
Many cells die during mammalian development. However, as the dead cells are swiftly engulfed by macrophages for degradation, it is difficult to detect them. For example, more than 90% of thymocytes undergo programmed cell death during the development of the thymus. But, only a small percentage of the cells can be stained by TUNEL at a given time point.28 Many neurons also undergo programmed cell death in the brain, but the reported number differs greatly among researchers,29, 30, 31 probably due to the difficulty in detecting dead cells that are quickly eliminated. Here, we used DNase II-null mice to detect the programmed cell death that occurs during mouse embryogenesis. The DNase II-null mice carried in various tissues macrophages that contained undigested DNA, suggesting that cells died surrounding the macrophages and were engulfed. Although it is difficult to determine which and how many cells died, the DNase II−/− mice were nonetheless useful for localizing the regions where programmed cell death occurs. The macrophages carrying undigested DNA eventually disappeared, probably because those with a heavy load of DNA could not survive, suggesting that the DNase II−/− mice can also be used to estimate the time when programmed cell death takes place. In fact, we have been able to show that specific layers of cerebral neurons undergo programmed cell death at specific stages of mouse embryogenesis (AN, KK, and SN, manuscript in preparation).
Crossing the Apaf-1−/− mice with DNase II−/− mice, we showed that most of the apoptotic cell death during mouse embryonic development takes place through an Apaf-1-dependent pathway. This agrees with previous reports that deletion of the Apaf-1 or caspase-9 gene blocks apoptotic cell death in the brain, and ear.13, 14 Similarly, the apoptotic cell death in E17.5 thymus was also blocked by deficiency of Apaf-1, indicating that the negative and positive selections at CD4+CD8+ thymocytes proceed in an Apaf-1-dependent pathway. In contrast, very surprisingly, we found that caspase-3 could be activated without Apaf-1 in the E14.5 thymus, in which most thymocytes are CD4−CD8−.32 Furthermore, staurosporine but not etoposide activated caspase-3 in the fetal thymocytes without Apaf-1. Etoposide and staurosporine are inhibitors for topoisomerase and kinase C, respectively, and both of them have been reported to activate caspase-3 by releasing cytochrome c from mitochondria.33, 34 Thus, no requirement of Apaf-1 in the staurosporine-induced activation of caspase-3 in the fetal thymocytes was unexpected. This result suggests that staurosporine can activate caspase-3 in thymocytes through a pathway that does not use cytochrome c, and this pathway may be similar to that used in the Apaf-1-independent apoptotic cell death of E14.5 thymocytes. We observed that caspase-3 can be activated without Apaf-1 not only in the fetal thymus but also in the fetal liver (AN, KK, and SN, unpublished observation), which may be consistent with reports that hemopoietic cells reconstituted with fetal liver cells can activate caspase without Apaf-1.18, 35, 36 To understand this apoptosis, it will be necessary to determine how and which caspases are activated by staurosporine without Apaf-1.
Even when the caspase was not activated in Apaf-1-deficient mice, macrophages carrying undigested DNA were present in the embryo, in particular at the late stage of development, indicating that cells could die without Apaf-1, and were engulfed by macrophages. This agrees with the previous report showing that myeloid cells die without Apaf-1 or caspase-9.37 Several Apaf-1- or caspase-independent cell death processes have been proposed, including autophagic cell death and necrosis.38 As reported for the interdigits of Apaf-1−/− embryos,39 we found in the ectodermal ridge of the tails of Apaf-1−/− mice many cells exhibiting mottled chromatin condensation, nuclear membrane detachment and rupture, and dilated mitochondria. Whereas, no cells with the characteristics of autophagy (double-membraned vacuoles), indicating that the cells probably died because of necrosis. One possible cause of nonapoptotic cell death is that the stimuli that induce programmed cell death in mouse embryogenesis cause cytochrome c to be released, inactivating mitochondrial function, which leads to cell death.37 The mice lacking both Bak and Bax that are essential for the release of cytochrome c40 show the persistence of interdigital webs,17 which suggests that Bak/Bax can cause necrotic cell death by releasing cytochrome c. It will be interesting to examine whether the null mutation of Bax and Bak can prevent the generation of macrophages carrying undigested DNA in DNase II-null embryos.
In Apaf-1−/− embryos, dead cells were found in macrophages, indicating that the nonapoptotic dead cells were recognized by macrophages for engulfment. In C. elegans, a common set of genes mediates the removal of both apoptotic and necrotic cell corpses.41 Conversely, different mechanisms have been proposed to clear apoptotic and necrotic cells in mammalian system.42 Apoptotic cells expose phosphatidylserine, which is recognized by specific receptors or opsonins for engulfment.12 Because necrotic cells also expose phosphatidylserine at a late stage,38 it is possible that similar receptors or opsonins mediate the engulfment of apoptotic and necrotic cells. The complement system is another candidate for the engulfment of late apoptotic or necrotic cells.43 It will be interesting to study the engulfment of necrotic cells by crossing the Apaf-1−/− mice with mice deficient in the engulfment of apoptotic or necrotic cells.44, 45 Unengulfed apoptotic cells are difficult to find in mouse tissues. However, we frequently detected unengulfed dead cells with nonapoptotic morphology in the Apaf-1−/− embryos, suggesting that the engulfment of nonapoptotic dead cells is inefficient compared with that of apoptotic cells. This may allow noxious materials to be released from dying cells. In addition, the engulfment of necrotic cells but not apoptotic ones is known to cause the release of inflammatory cytokines.46 In this regard, it will be interesting to study whether the normally developing Apaf-1−/− mice suffer from inflammatory disease.
Materials and Methods
Mice
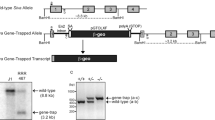
C57BL/6 mice were purchased from Nippon SLC (Hamamatsu, Japan) or Shimizu Laboratory (Kyoto, Japan) Supplies. DNase II−/− mice20 were backcrossed to C57BL/6 at lease six times. IFN-IR−/− mice47 in the C57BL/6 background were obtained from Michel Aguet (Swiss Institute for Experimental Cancer Research, Epalinges, Switzerland). Apaf-1−/− mice in the C57BL/6 background were described previously.14, 19 DNase II−/−IFN-IR−/− mice and Apaf-1−/−DNase II−/−IFN-IR−/− embryos were generated by crossing Apaf-1+/−DNase II−/−IFN-IR−/− parents, respectively. Mice were housed in specific pathogen-free facilities at Osaka University Medical School and Oriental Bio Service. All animal experiments were conducted according to the Guidelines for Animal Experiments of Osaka University or Kyoto University. To determine the genotype of DNase II and IFN-IR alleles, we prepared DNA from embryonic tissues or adult tail-snip tissue, as described,48 and analyzed using PCR. For the DNase II gene, a sense primer specific for the wild-type (5′-GCCCATCTAGACTAACTTTC-3′) or mutant allele (5′-GATTCGCAGCGCATCGCCTT-3′); sequence in the neomycin-resistant gene) was used with a common antisense primer (5′-GAGTCTTAGTCCTTTGCTCCG-3′). The wild-type and mutant alleles for IFN-IR were examined with a wild-type (5′-AAGATGTGCTGTTCCCTTCCTCTGCTCTGA-3′) or mutant-specific (5′-CCTGCGTGCAATCCATCTTG-3′) antisense primer and a common sense primer (5′-ATTATTAAAAGAAAAGACGAGGCGAAGTGG-3′). For the Apaf-1 allele, a wild-type (5′-CTCAAACACCTCCTCCACAA-3′) or mutant-specific (5′-GGGCCAGCTCATTCCTC-3′) sense primer was used with a common antisense primer (5′-GTCATCTGGAAGGGCAGCGA-3′).
Histochemical analysis with paraffin sections
Embryos were fixed with 4% PFA in 0.1 M Na-phosphate buffer (pH 7.2) containing 4% sucrose at 4°C for more than 1 day with shaking. They were gradually dehydrated by dipping into 50, 70, 80, and 90% ethanol at room temperature for 12 h each, and then twice in 100% ethanol for 1 h. The samples were soaked twice in 100% xylene at room temperature for 1 h, in a 1 : 1 mixture of xylene and paraffin for 2 h at 60°C. They were embedded in paraffin by successive incubations in paraffin for 12 and 4 h at 60°C, and sectioned at 4 μm using a microtome (RM2245; Leica, Solms, Germany).
For TUNEL staining, sections were treated at room temperature for 20 min with 20 μg/ml Proteinase K, and stained with an Apoptag kit (Millipore, Bedford, MA, USA), according to the manufacturer's instructions, except that the amount of terminal transferase was reduced to 10% of the recommended concentration. Sections were mounted with Mount-Quick (Daido Sangyo, Toda, Saitama, Japan) or Fluoromount (Diagnostic BioSystems, Pleasanton, CA, USA), and observed by fluoroescence microscopy (IX-70; Olympus, Tokyo, Japan or BioRevo BZ-9000; Keyence, Osaka, Japan).
Histochemical analysis with cryosections
Embryos were fixed at 4°C in 4% PFA containing 4% sucrose in 0.1 M Na-phosphate buffer (pH 7.2) for 2 h, successively immersed in 10 and 20% sucrose-containing 0.1 M Na-phosphate buffer (pH 7.2) for 4 h and overnight each, embedded in OCT compound (Sakura Finetek, Tokyo, Japan), and frozen in liquid nitrogen. Sections (10 μm) were prepared using a cryostat (CM3050 S; Leica) in the cold (−16 to −25°C).
To detect active caspase-3, we fixed sections with 4% PFA in PBS at room temperature for 10 min, and blocked with 5% normal goat serum in PBS containing 0.3% Triton X-100 at room temperature for 1 h. They were stained at 4°C overnight with a 100-fold diluted rabbit monoclonal antibody (mAb) against active caspase-3 (clone 5A1E; Cell Signaling, Danvers, MA, USA), followed by incubation at room temperature for 1 h with Cy3-conjugated goat anti-rabbit IgG (Jackson Laboratories, West Grove, PA, USA).
For the staining of F4/80 antigen, a rat hybridoma against mouse F4/8049 was grown in serum-free GIT medium (Nihon Pharmaceutical, Tokyo, Japan). Cryosections from mouse embryos were fixed at room temperature for 10 min with 1% PFA in PBS. After blocking with 10% normal rabbit serum and 0.5% BSA in PBS at room temperature for 1 h, sections were incubated at 4°C overnight with the supernatant of the F4/80 hybridoma, and washed with PBS containing 0.5% BSA. The endogenous peroxidase was quenched by incubation at 4°C for 20–30 min with 3% H2O2 in methanol, and incubated at room temperature for 45 min with peroxidase-conjugated rabbit anti-rat Ig (Dako, Copenhagen, Denmark). The signals were detected by incubation at room temperature for 5 min with Cy3-labeled tyramide (PerkinElmer, Boston, MA, USA).
TUNEL staining was performed as described above, except that the concentration of terminal transferase was reduced to 0.4–0.5% of that recommended by the manufacturer. Sections were mounted with mounting reagent containing 1–2 μg/ml DAPI (Dojin Laboratories, Kumamoto, Japan), and observed by microscopy as described earlier.
Electron microscopy
Embryonic tissues were fixed by incubation at 4°C for 2 h in 0.1 M Na-phosphate buffer (pH 7.2) containing 2% PFA and 2% glutaraldehyde. After being washed five times with 0.1 M Na-phosphate buffer (pH 7.2) at 4°C for 20 min each, the samples were post-fixed at 4°C for 2 h with 1% OsO4 in the same buffer, and dehydrated at 4°C by dipping into a graded series (50, 60, 70, 80, 90, 99%) of ethanol for 10 min each. After two 20-min immersions in 100% ethanol, the samples were incubated at 35°C twice in propylene oxide for 20 min, in a 3 : 1 mixture of propylene oxide and epoxide for 1 h, in a 1 : 3 mixture of propylene oxide and epoxide for 1 h, and in epoxide overnight. They were then embedded in epoxide by incubation at 60°C for 3 days. Ultrathin sections (80–90 nm) were prepared with an ultramicrotome (Ultracut N; Reichert-Nissei, Kumamoto, Japan), stained with uranyl acetate and lead citrate, and observed with a Hitachi H-7650 microscope (Hitachi High-Technologies, Tokyo, Japan).
Assay for caspase-3
Thymocytes (4 × 104 cells per 100 μl) from E14.5 embryos or thymocytes (5 × 105 cells per 200 μl) from E17.5 embryos were treated with 10 μM staurosporine or 50 μM etoposide in DMEM containing 10% FCS. The caspase-3 activity in the cell lysates and medium was determined using Caspase-3 Cellular Assay Kit PLUS (Enzo Life Sciences, Farmingdale, NY, USA) according to the instructions provided by the manufacturer. In brief, cells were collected by centrifugation at 5000 r.p.m. for 5 min, washed with PBS, and lysed with 50 μl Lysis Buffer (50 mM HEPES-NaOH buffer (pH 7.4), 0.1% CHAPS, 5 mM DTT, and 0.1 mM EDTA). Aliquots (10 μl) of the medium or cell lysates were incubated at 37°C for 3 h with 30 μM Ac-DEVD-AMC in 100 μl Assay Buffer (50 mM HEPES-NaOH buffer (pH 7.4), 100 mM NaCl, 0.1% CHAPS, 10 mM DTT, 1 mM EDTA, and 10% glycerol), and the fluorescence was detected with an excitation wavelength of 360 nm and emission wavelength of 460 using a microplate reader (Infinite M200; Tecan, Männedorf, Switzerland). The specific caspase-3 activities were determined by subtracting the values obtained in the presence of 0.1 μM Ac-DEVD-CHO.
Western blot analysis
For western blotting, cells (5 × 105) were directly lysed by heating at 85°C for 30 min in 15 μl of sample buffer (30 mM Tris-HCl (pH 6.8), 1% SDS, 5% glycerol, 2.5% 2-mercaptoethanol, and 0.0005% BPB). Proteins were separated by 10–20% gradient SDS–PAGE, and transferred to PVDF membranes (Millipore). After blocking with blocking buffer (Tris-buffered saline containing 0.05% Tween 20 (TBST) supplemented with 5% non-fat dry milk or 4% Block Ace (DS Pharma Biomedical, Suita, Osaka, Japan)), the membranes were incubated at 4°C overnight with rabbit mAb against caspase-3 (clone 8G10; Cell Signaling) or mouse mAb against caspase-9 (clone 5B4; MBL, Nagoya, Japan) in blocking buffer. After washing with TBST, the membranes were incubated with HRP-conjugated anti-rabbit Ig or anti-mouse Ig antibody at room temperature for 1 h, and the proteins recognized by the antibody were visualized by a chemiluminescence reaction (Immobilon Western, Millipore). Western blotting with mouse mAb against BiP (Grp78) (clone 40; BD Biosciences) was performed as a loading control.
For immunoprecipitation, the culture supernatant was precleared by incubating twice at 4°C with a 1 : 1 mixture of nProtein A-Sepharose 4 Fast Flow (GE Healthcare, Uppsala, Sweden) and Protein G-Sepharose 4 Fast Flow (GE Healthcare) for 1.5 h. After removing Sepharose by centrifugation, the supernatant was incubated at 4°C overnight with rabbit mAb against the cleaved caspase-3 (clone 5A1E; Cell Signaling), followed by the incubation with Protein A-Sepharose. After washing with PBS containing 1% Triton X-100, the proteins bound to the beads were eluted by boiling in SDS sample buffer, separated by 10–20% SDS–PAGE, transferred onto a PVDF membrane, and subjected to western blotting with rabbit mAb against the cleaved caspase-3, followed by the incubation with HRP-conjugated Protein A (Bio-Rad, Hercules, CA, USA).
Abbreviations
- CAD:
-
caspase-activated DNase
- mAb:
-
monoclonal antibody
References
Jacobson MD, Weil M, Raff MC . Programmed cell death in animal development. Cell 1997; 88: 347–354.
Vaux DL, Korsmeyer SJ . Cell death in development. Cell 1999; 96: 245–254.
Kerr JF, Wyllie AH, Currie AR . Apoptosis: a basic biological phenomenon with wide-ranging implications in tissue kinetics. Br J Cancer 1972; 26: 239–257.
Adams JM . Ways of dying: multiple pathways to apoptosis. Genes Dev 2003; 17: 2481–2495.
Danial NN, Korsmeyer SJ . Cell death: critical control points. Cell 2004; 116: 205–219.
Nagata S . Apoptosis by death factor. Cell 1997; 88: 355–365.
Strasser A, Jost PJ, Nagata S . The many roles of FAS receptor signaling in the immune system. Immunity 2009; 30: 180–192.
Nagata S . DNA degradation in development and programmed cell death. Annu Rev Immunol 2005; 23: 853–875.
Jiang X, Wang X . Cytochrome c-mediated apoptosis. Annu Rev Biochem 2004; 73: 87–106.
Schafer ZT, Kornbluth S . The apoptosome: physiological, developmental, and pathological modes of regulation. Dev Cell 2006; 10: 549–561.
Fadok VA, Bratton DL, Frasch SC, Warner ML, Henson PM . The role of phosphatidylserine in recognition of apoptotic cells by phagocytes. Cell Death Differ 1998; 5: 551–562.
Ravichandran KS, Lorenz U . Engulfment of apoptotic cells: signals for a good meal. Nat Rev Immunol 2007; 7: 964–974.
Kuida K, Haydar TF, Kuan CY, Gu Y, Taya C, Karasuyama H et al. Reduced apoptosis and cytochrome c-mediated caspase activation in mice lacking caspase 9. Cell 1998; 94: 325–337.
Yoshida H, Kong YY, Yoshida R, Elia AJ, Hakem A, Hakem R et al. Apaf1 is required for mitochondrial pathways of apoptosis and brain development. Cell 1998; 94: 739–750.
Hao Z, Duncan GS, Chang CC, Elia A, Fang M, Wakeham A et al. Specific ablation of the apoptotic functions of cytochrome c reveals a differential requirement for cytochrome c and Apaf-1 in apoptosis. Cell 2005; 121: 579–591.
Cecconi F, Alvarez-Bolado G, Meyer BI, Roth KA, Gruss P . Apaf1 (CED-4 homolog) regulates programmed cell death in mammalian development. Cell 1998; 94: 727–737.
Lindsten T, Ross AJ, King A, Zong WX, Rathmell JC, Shiels HA et al. The combined functions of proapoptotic Bcl-2 family members bak and bax are essential for normal development of multiple tissues. Mol Cell 2000; 6: 1389–1399.
Marsden VS, O’Connor L, O’Reilly LA, Silke J, Metcalf D, Ekert PG et al. Apoptosis initiated by Bcl-2-regulated caspase activation independently of the cytochrome c/Apaf-1/caspase-9 apoptosome. Nature 2002; 419: 634–637.
Okamoto H, Shiraishi H, Yoshida H . Histological analyses of normally grown, fertile Apaf1-deficient mice. Cell Death Differ 2006; 13: 668–671.
Kawane K, Fukuyama H, Kondoh G, Takeda J, Ohsawa Y, Uchiyama Y et al. Requirement of DNase II for definitive erythropoiesis in the mouse fetal liver. Science 2001; 292: 1546–1549.
Kawane K, Fukuyama H, Yoshida H, Nagase H, Ohsawa Y, Uchiyama Y et al. Impaired thymic development in mouse embryos deficient in apoptotic DNA degradation. Nat Immunol 2003; 4: 138–144.
Krieser RJ, MacLea KS, Longnecker DS, Fields JL, Fiering S, Eastman A . Deoxyribonuclease IIalpha is required during the phagocytic phase of apoptosis and its loss causes lethality. Cell Death Differ 2002; 9: 956–962.
Wu YC, Stanfield GM, Horvitz HR . NUC-1, a Caenorhabditis elegans DNase II homolog, functions in an intermediate step of DNA degradation during apoptosis. Genes Dev 2000; 14: 536–548.
Mukae N, Yokoyama H, Yokokura T, Sakoyama Y, Nagata S . Activation of the innate immunity in Drosophila by endogenous chromosomal DNA that escaped apoptotic degradation. Genes Dev 2002; 16: 2662–2671.
Reddien PW, Horvitz HR . The engulfment process of programmed cell death in Caenorhabditis elegans. Annu Rev Cell Dev Biol 2004; 20: 193–221.
Yoshida H, Okabe Y, Kawane K, Fukuyama H, Nagata S . Lethal anemia caused by interferon-beta produced in mouse embryos carrying undigested DNA. Nat Immunol 2005; 6: 49–56.
Denecker G, Vercammen D, Steemans M, Vanden Berghe T, Brouckaert G, Van Loo G et al. Death receptor-induced apoptotic and necrotic cell death: differential role of caspases and mitochondria. Cell Death Differ 2001; 8: 829–840.
Surh CD, Sprent J . T-cell apoptosis detected in situ during positive and negative selection in the thymus. Nature 1994; 372: 100–103.
Verney C, Takahashi T, Bhide PG, Nowakowski RS, Caviness Jr VS . Independent controls for neocortical neuron production and histogenetic cell death. Dev Neurosci 2000; 22: 125–138.
Thomaidou D, Mione MC, Cavanagh JF, Parnavelas JG . Apoptosis and its relation to the cell cycle in the developing cerebral cortex. J Neurosci 1997; 17: 1075–1085.
Blaschke AJ, Staley K, Chun J . Widespread programmed cell death in proliferative and postmitotic regions of the fetal cerebral cortex. Development 1996; 122: 1165–1174.
Rodewald HR, Moingeon P, Lucich JL, Dosiou C, Lopez P, Reinherz EL . A population of early fetal thymocytes expressing Fc gamma RII/III contains precursors of T lymphocytes and natural killer cells. Cell 1992; 69: 139–150.
Kaufmann SH . Cell death induced by topoisomerase-targeted drugs: more questions than answers. Biochim Biophys Acta 1998; 1400: 195–211.
Maeno E, Ishizaki Y, Kanaseki T, Hazama A, Okada Y . Normotonic cell shrinkage because of disordered volume regulation is an early prerequisite to apoptosis. Proc Natl Acad Sci USA 2000; 97: 9487–9492.
Hara H, Takeda A, Takeuchi M, Wakeham AC, Itie A, Sasaki M et al. The apoptotic protease-activating factor 1-mediated pathway of apoptosis is dispensable for negative selection of thymocytes. J Immunol 2002; 168: 2288–2295.
Matsuki Y, Zhang HG, Hsu HC, Yang PA, Zhou T, Dodd CH et al. Different role of Apaf-1 in positive selection, negative selection and death by neglect in foetal thymic organ culture. Scand J Immunol 2002; 56: 174–184.
Ekert PG, Read SH, Silke J, Marsden VS, Kaufmann H, Hawkins CJ et al. Apaf-1 and caspase-9 accelerate apoptosis, but do not determine whether factor-deprived or drug-treated cells die. J Cell Biol 2004; 165: 835–842.
Kroemer G, Galluzzi L, Vandenabeele P, Abrams J, Alnemri ES, Baehrecke EH et al. Classification of cell death: recommendations of the Nomenclature Committee on Cell Death 2009. Cell Death Differ 2009; 16: 3–11.
Chautan M, Chazal G, Cecconi F, Gruss P, Golstein P . Interdigital cell death can occur through a necrotic and caspase-independent pathway. Curr Biol 1999; 9: 967–970.
Wei MC, Zong WX, Cheng EH, Lindsten T, Panoutsakopoulou V, Ross AJ et al. Proapoptotic BAX and BAK: a requisite gateway to mitochondrial dysfunction and death. Science 2001; 292: 727–730.
Chung S, Gumienny TL, Hengartner MO, Driscoll M . A common set of engulfment genes mediates removal of both apoptotic and necrotic cell corpses in C. elegans. Nat Cell Biol 2000; 2: 931–937.
Krysko DV, Denecker G, Festjens N, Gabriels S, Parthoens E, D’Herde K et al. Macrophages use different internalization mechanisms to clear apoptotic and necrotic cells. Cell Death Differ 2006; 13: 2011–2022.
Zwart B, Ciurana C, Rensink I, Manoe R, Hack CE, Aarden LA . Complement activation by apoptotic cells occurs predominantly via IgM and is limited to late apoptotic (secondary necrotic) cells. Autoimmunity 2004; 37: 95–102.
Hanayama R, Tanaka M, Miyasaka K, Aozasa K, Koike M, Uchiyama Y et al. Autoimmune disease and impaired uptake of apoptotic cells in MFG-E8-deficient mice. Science 2004; 304: 1147–1150.
Botto M, Dell’Agnola C, Bygrave AE, Thompson EM, Cook HT, Petry F et al. Homozygous C1q deficiency causes glomerulonephritis associated with multiple apoptotic bodies. Nat Genet 1998; 19: 56–59.
Fadok VA, Bratton DL, Guthrie L, Henson PM . Differential effects of apoptotic versus lysed cells on macrophage production of cytokines: role of proteases. J Immunol 2001; 166: 6847–6854.
Muller U, Steinhoff U, Reis LF, Hemmi S, Pavlovic J, Zinkernagel RM et al. Functional role of type I and type II interferons in antiviral defense. Science 1994; 264: 1918–1921.
Laird PW, Zijderveld A, Linders K, Rudnicki MA, Jaenisch R, Berns A . Simplified mammalian DNA isolation procedure. Nucleic Acids Res 1991; 19: 4293.
Austyn JM, Gordon S . F4/80, a monoclonal antibody directed specifically against the mouse macrophage. Eur J Immunol 1981; 11: 805–815.
Acknowledgements
We thank Dr. M Koike (Juntendo University) for his advice for the immunochemical staining, and M Fujii and M Harayama for secretarial assistance. This work was supported in part by Grants-in-Aid from the Ministry of Education, Science, Sports, and Culture in Japan, and by the Kyoto University Global COE Program (Center for Frontier Medicine). AN was a research assistant for the Osaka University Global COE Program (System Dynamics of Biological Function).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no conflict of interest.
Additional information
Edited by G Melino
Rights and permissions
About this article
Cite this article
Nagasaka, A., Kawane, K., Yoshida, H. et al. Apaf-1-independent programmed cell death in mouse development. Cell Death Differ 17, 931–941 (2010). https://doi.org/10.1038/cdd.2009.186
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/cdd.2009.186
Keywords
This article is cited by
-
Lysosomes, caspase-mediated apoptosis, and cytoplasmic activation of P21, but not cell senescence, participate in a redundant fashion in embryonic morphogenetic cell death
Cell Death & Disease (2023)
-
The effects of temperature variation treatments on embryonic development: a mouse study
Scientific Reports (2022)
-
Epiblast fragmentation by shedding—a novel mechanism to eliminate cells in post-implantation mouse embryos
Cell Death & Differentiation (2022)
-
In vivo detection of programmed cell death during mouse heart development
Cell Death & Differentiation (2020)
-
A cyclometalated iridium(III) complex induces apoptosis and autophagy through inhibition of the PI3K/AKT/mTOR pathway
Transition Metal Chemistry (2018)