Abstract
The amyloid precursor protein (APP) has been suggested to regulate gene expression. GeneChip® analysis and in vitro kinase assays revealed potent APP-dependent repression of c-Jun, its target gene SPARC and reduced basal c-Jun N-terminal kinase (JNK) activity in PC12 cells overexpressing APP. UV-induced activation of the JNK signalling pathway and subsequent apoptosis were likewise reduced by APP and this effect could be mimicked by the indirect JNK inhibitor CEP-11004. Treatment with a B⃗-secretase inhibitor did not affect APP-mediated downmodulation of the JNK signalling pathway, suggesting that the effects might be mediated via α-secretase processing of APP. In support of these data, overexpression of the Swedish mutant of APP did not inhibit SPARC expression, UV-induced JNK activation and cell death. Our data suggest an important physiological role of APP and α-secretase activity in the control of JNK/c-Jun signalling, target gene expression and cell death activation in response to cytotoxic stress.
Similar content being viewed by others
Introduction
In recent years, significant progress has been achieved in elucidating the molecular mechanisms leading to neurodegeneration in Alzheimeŕs disease (AD). The β-amyloid precursor protein (APP), a type 1 transmembrane protein, plays a pivotal role in the pathophysiology of AD.1, 2 The majority of APP is cleaved by the activity of α-secretase at the cell surface1, 3 leading to secretion of sAPPα, which might act as a neuroprotective cytokine and regulate gene expression.4 In an alternative processing pathway, APP is cleaved by β-secretase at the N-terminus of the Aβ domain. The generated cleavage fragment can be further processed by the activity of γ-secretase to yield Aβ, the major constituent of Alzheimer plaques.1, 3 Generation of Aβ peptides can be prevented by α-secretase activity, which cuts APP within the Aβ sequence. Therefore, α- and β-secretase are concurring activities, the latter being favored by a number of missense mutations identified to date within three genes associated with familial AD (FAD): APP, presenilin 1 and presenilin 2. Presenilins are believed to be the active components of γ-secretase activity.5, 6 However, at present, little is known about the biological function of the APP C-terminal region from which the APP intracellular domain (AICD) is cleaved off by γ-secretase.2, 7 Recent findings indicate that AICD may serve as a membrane-to-nucleus messenger;8, 9 thus, both sAPPα and AICD might link APP to regulation of transcriptional processes. The YENPTY motif of the C-terminal domain of APP has been shown to bind to several interaction partners, among them Fe65 and c-Jun N-terminal kinase (JNK)-interacting protein-1b (JIP-1b).8, 10, 11, 12 JIP-1b has been shown to serve as a scaffold molecule for JNKs and their upstream kinases13, 14 and thus may couple APP to the JNK mitogen-activated protein (MAP) kinase signal transduction pathway. The JNK signalling pathway is a central stress signalling pathway implicated in neuronal plasticity, regeneration and cell death of both neuronal and non-neuronal cells.15 It can be activated by a large variety of cytotoxic stresses, such as oxidative stress, genotoxic stress, ischemia, heat shock and proinflammatory cytokines. In the present study, we investigated the effect of APP on basal gene expression using microarray technology, and demonstrate an important function of APP in the regulation of the JNK/c-Jun signalling pathway and JNK-dependent apoptosis.
Results
Gene expression microarray analysis of APP-mediated transcriptional changes
To analyze APP-mediated transcriptome alterations, we used high-density oligonucleotide microarrays for global analysis of gene expression in PC12 cells. Comparison of APP-dependent changes in basal gene expression (i.e. in the absence of cytotoxic stress) was performed with two independent PC12 cell lines, PC12 APP M5 and PC12 APP N10, overexpressing equal amounts of the human brain-specific isoform APP695,16, 17, 18 which lacks the the Kunitz-type protease inhibitor (KPI) domain.1 Empty vector-transfected PC12 cells (PC12neo O1) served as control. Tables 1 and 2 provide the lists of differentially expressed genes (for a detailed description of data analysis, see Materials and Methods) with an at least 1.5-fold expression increase or decrease in both APP-overexpressing cell clones. A total of 34 upregulated and 13 downregulated genes were identified in APP-overexpressing cells. APP significantly activated expression of the genes encoding the mitochondrial antioxidant manganese superoxide dismutase (MnSOD, 2.3- and 1.7-fold), cytochrome c (2.0- and 1.7-fold), the presenilin-interacting protein Sel-10 (2.1- and 2.1-fold), the developmentally expressed protein TPO1 (2.8- and 3.0-fold) and the synaptosomal-associated protein 25 kDa (SNAP-25B, 2.6- and 4.6-fold). Other targets found to be transcriptionally activated by APP included calnexin and the mitochondrial heat shock protein gene mt-hsp70/mortalin (Table 1). Increased expression of selective APP-induced genes was confirmed by semiquantitative RT-PCR analysis (Figure 1a).
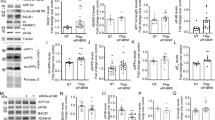
Semiquantitative RT-PCR analysis of transcriptional alterations induced by overexpression of APP. (a) Basal expression of putative APP target genes c-Jun, SPARC/osteonectin, MnSOD and Sel-10 was analyzed in PC12 APPwt M5 and PC12 APPwt N10 cells, with PC12 neo O1 (neo) serving as a negative control. Expression of GAPDH served as an internal control. Expression levels of individual genes in PC12 APPwt M5 and PC12 APPwt N10 relative to the PC12 neo O1 control were analyzed with the ImageJ software. (b) Expression of c-Jun target gene SPARC/osteonectin is not altered by application of a γ-secretase inhibitor. RT-PCR analysis in PC12 neo O1, PC12 APPwt M5, PC12 APPwt N10, PC12 APPsw Q8 and PC12 APPsw Q7 cells left untreated (upper panel) or treated with 10 μM γ-secretase inhibitor (γ-SI, lower panel) for 24 h. Expression of GAPDH served as an internal control
Inhibition of expression of c-Jun/AP-1 target genes and reduced basal JNK activity in APP-overexpressing cells
Interestingly, among the genes most prominently downregulated in APP-overexpressing cells, we found SPARC/osteonectin and the transcription factor c-Jun (Table 2). SPARC/osteonectin (SPARC: secreted protein, acidic and rich in cysteine) has been shown to be induced in response to c-Jun overexpression and is a proposed c-Jun/activator protein-1 (AP-1) target gene.19 It is to be noted that, c-Jun also is a c-Jun/AP-1 target gene, and is believed to amplify c-Jun-dependent transcription.20 Downregulation of both c-Jun and SPARC/osteonectin was confirmed by RT-PCR analysis in the APP-overexpressing cells (Figure 1a). To analyze the role of the APP metabolism in APP-dependent transcriptional regulation, we employed two PC12 cell lines stably overexpressing KM670/671NL, the Swedish double mutant of APP (APPsw) (Figure 1b), which is expressed at equal levels as APP wild type (APPwt).16 KM670/671NL is preferentially processed via the amyloidogenic β-secretase pathway and is known to be associated with enhanced Aβ levels and abrogation of sAPPα formation. In comparison to APPwt-overexpressing cells, expression of SPARC/osteonectin was significantly enhanced in cells overexpressing APPsw (Figure 1b). A fenchylamine sulfonamide γ-secretase inhibitor21 had no discernible effect in any of the cell lines (Figure 1b), indicating that APP processing by α-secretase might be required for the APP-mediated inhibition of SPARC/osteonectin expression. We subsequently investigated whether APP exerts a regulatory effect on the JNK signalling pathway. To this end, we analyzed basal JNK activity using an in vitro kinase assay (Figure 2a). Incubation with cellular extracts of PC12 neo cells and PC12 APPsw cells led to detectable amounts of phosphorylated recombinant GST-c-Jun, which were greatly diminished after incubation with extracts of PC12 APPwt cells.
APPwt inhibits both basal and stress-induced JNK activity. (a) PC12 neo O1, PC12 APPwt M5 and PC12 APPsw Q8 cultures were UV-irradiated (UV) or left untreated (Basal). JNK activity was analysed by a GST-c-Jun in vitro kinase assay (upper panel). To confirm equal c-Jun levels in the samples, the residual protein in the gel was stained with Simply Blue Safe Stain (Invitrogen Life Technologies) after blotting (lower panel). (b, c) APPwt inhibits UV-triggered JNK phosphorylation. For analysis of JNK activation, cultures of PC12 neo O1, PC12 APP M5 and PC12 APPsw Q8 (b, left panel) or PC12 neo O3, PC12 APPwt N10 and PC12 APPsw Q7 (c) were subjected to UV irradiation (UV) or left untreated (Basal). At 4 h postirradiation, cultures were lysed and subjected to Western blotting with the anti-ACTIVE® JNK antiserum. Effect of a γ-secretase inhibitor (γ-SI) on UV-induced JNK activation (b, right panel) is shown. PC12 neo O1, PC12 APP M5 and PC12 APPsw Q8 cells were preincubated with γ-SI for 24 h at 10 μM, UV-irradiated and continuously incubated in the presence of γ-SI for 4 h postirradiation. (d) APPwt inhibits basal and UV-induced c-Jun expression. For analysis of c-Jun levels, cultures of PC12 neo O1 and PC12 APP M5 were subjected to UV irradiation (UV) or left untreated (Basal). At 4 h postirradiation, cultures were lysed and subjected to Western blotting with the c-jun (Ab-3) antibody. JNK1 served as an internal loading control
APP inhibits UV-induced JNK activation and cell death
We also determined whether APP modulated stress-induced JNK activity. UV irradiation triggers activation of the JNK signalling pathway in a wide array of cell types, including neurons.22 UV induced a marked elevation of the active forms of JNK1/JNK2 in two independent vector-transfected PC12 cell lines (PC12 neo O1 and PC12 neo O3) (Figure 2b, left panel, and Figure 2c) as detected with a phospho-specific antibody binding to the dually phosphorylated Thr/Pro/Tyr region of the JNKs. Remarkably, UV-triggered JNK phosphorylation was potently inhibited in the two APPwt-overexpressing cell lines (PC12 APPwt M5 and PC12 APPwt N10), but not in the APPsw-overexpressing cell lines (PC12 APPsw Q8 and PC12 APPsw Q7) (Figure 2b, left panel, and Figure 2c). APPwt also potently downmodulated UV-induced JNK activity detected with the in vitro kinase assay (Figure 2a). The γ-secretase inhibitor had no effect on stress-induced JNK activation in APPwt and APPsw cells (Figure 2b, right panel). In addition to the inhibitory effect of APPwt on c-Jun transcription (Figure 1a), we observed significantly lowered c-Jun protein levels in PC12 APPwt cells, both under basal conditions and after UV irradiation (Figure 2d).
In light of the strong inhibitory effect of APP on UV-triggered JNK activation, we investigated whether APP overexpression affected cell survival after UV irradiation. The JNK signalling pathway is known to play a fundamental role in UV-triggered, caspase-dependent apoptosis.23, 24 To elucidate the potential role of APP in modulation of UV-induced apoptosis, we analyzed the activation of the effector caspase caspase 3 by Western blotting. In comparison to neo cells, we found significantly lowered amounts of the active caspase 3 subunit in extracts of UV-treated PC12 APPwt cells, but not in PC12 APPsw cells (Figure 3a). To investigate nuclear alterations during UV-induced apoptosis, we performed Hoechst staining 16 h postirradiation. A large number of nuclei were condensed and/or fragmented in UV-treated PC12 neo cells. In contrast, the number of apoptotic nuclei of UV-treated PC12 APP cells was markedly lower (Figure 3b). Analysis of effector caspase activation with an enzymatic assay revealed that APPwt not only suppressed UV-triggered JNK activation but also significantly reduced UV-triggered activation of DEVDases in comparison to PC12 neo controls (Figure 3c). Since APPsw cells showed no reduction of JNK activation after UV irradiation, we also compared the effect of overexpressed APPwt and APPsw on UV-triggered apoptosis. In contrast to APPwt, APPsw did not enhance cellular resistance against apoptosis (Figure 3c). Since Aβ has been implicated in caspase-independent cell death,25 we also performed a quantitative comparison of apoptosis and (primary and secondary) necrosis in UV-treated cultures by simultaneous staining with Hoechst and propidium iodide (PI) (Figure 3d). In concordance with the DEVDase assays, overexpression of APPwt reduced the percentage of apoptotic cells after UV irradiation compared with vector-transfected controls. It is to be noted that overexpression of APPsw significantly increased necrotic cell death in the UV-treated cultures.
Overexpression of APP protects PC12 cells from UV-triggered apoptosis. (a) Analysis of UV-induced procaspase 3 cleavage in PC12 neo O1 (neo) and PC12 APP M5 (APP) cells treated with UV. Active caspase 3 p17 subunits were detected by Western blotting. (b) Nuclear apoptosis visualized by Hoechst staining. Cultures of PC12 neo O1 (neo) and PC12 APP M5 cells (APP) were UV-irradiated or left untreated. Images were taken 20 h postirradiation. Scale bar: 25 μm. (c) PC12 neo O1 (neo), PC12 APPwt M5 (M5), PC12 APPwt N10 (N10), PC12 APPsw Q8 (Q8) and PC12 APPsw Q7 (Q7) cells were UV-irradiated (UV). Control cultures were left untreated (Con). At 20 h postirradiation, the cultures were lysed and caspase 3-like activity was monitored over 1 h. Data are means±S.E.M. from n=4 cultures per treatment. Experiments were repeated two times with similar results. AU=arbitrary fluorescence units. *P<0.05 difference from vehicle-treated controls.  , P<0.05: difference from respective control cell line PC12 neo. (d) Parallel quantification of apoptosis and (primary and secondary) necrosis after UV irradiation. PC12 neo O1 (neo), PC12 APPwt M5 (APP) and PC12 APPsw Q8 (sw) cells were UV-irradiated (UV) or left untreated (Con). At 20 h postirradiation, the percentage of apoptotic and necrotic cells in the different cultures was analyzed as described in Materials and Methods. Data are means±S.E.M. from n=4 cultures per treatment. Experiments were repeated two times with similar results. *P<0.05: difference from vehicle-treated controls.
, P<0.05: difference from respective control cell line PC12 neo. (d) Parallel quantification of apoptosis and (primary and secondary) necrosis after UV irradiation. PC12 neo O1 (neo), PC12 APPwt M5 (APP) and PC12 APPsw Q8 (sw) cells were UV-irradiated (UV) or left untreated (Con). At 20 h postirradiation, the percentage of apoptotic and necrotic cells in the different cultures was analyzed as described in Materials and Methods. Data are means±S.E.M. from n=4 cultures per treatment. Experiments were repeated two times with similar results. *P<0.05: difference from vehicle-treated controls.  , P<0.05: difference from respective control cell line PC12 neo
, P<0.05: difference from respective control cell line PC12 neo
The death-inhibiting function of APP can be mimicked by an MLK inhibitor
Further proof for the critical involvement of the JNK signalling pathway and its regulation by APP in UV-triggered apoptosis was obtained from experiments with the synthetic and specific mixed-lineage kinase (MLK) inhibitor CEP-1100426, 27 (Figure 4). MLKs have been shown to bind to JIP-1b and are known upstream activators of JNKs. CEP-1004 almost completely abolished UV-induced Ser63 phosphorylation of c-Jun after UV irradiation in PC12 neo cells (Figure 4a). Application of CEP-11004 downmodulated apoptosis in PC12 neo cells to a level comparable to that observed in PC12 APPwt cells, while it had only a very moderate protective effect in the presence of overexpressed APP (Figure 3b).
(a) CEP-11004 significantly inhibits c-Jun phosphorylation. PC12 neo O1 cells were UV-irradiated (UV) or left untreated (Con). Where indicated, CEP-11004 was added at a final concentration of 200 nM 2 h prior to the irradiation time point. c-Jun phosphorylation was analyzed by Western blotting with the polyclonal phospho-c-Jun (Ser63) II antibody with JNK1 serving as a loading control. (b) The synthetic MLK inhibitor CEP-11004 can mimic the neuroprotective function of APP. PC12 neo O1 (neo) and PC12 APPwt M5 (APP) cells were UV-irradiated (UV) or left untreated (Con). Where indicated, CEP-11004 was added at a final concentration of 200 nM 2 h prior to the irradiation time point. At 20 h postirradiation, the cultures were lysed and caspase 3-like activity was monitored over one hour. Data are means±S.E.M. from n=4 cultures per treatment. Experiments were repeated two times with similar results. AU=arbitrary fluorescence units. *P<0.05: difference from vehicle-treated controls
Discussion
Here, we analyzed the effect of APP on global gene expression in PC12 cells. Our data demonstrate that APP induces a specific subset of neuroprotective and antioxidative genes, genes regulating mitochondrial function, as well as developmental genes. GeneChip analysis also demonstrated that APP potently downmodulated basal expression of c-Jun and the c-Jun/AP-1 target gene SPARC/osteonectin. Furthermore, we established an inhibitory role of APP in the regulation of the stress-induced JNK signalling cascade.
At least two proteolytic cleavage products of APP, sAPPα and AICD, which are released by γ-secretase cleavage, have been previously implicated in the regulation of transcriptional processes by APP.4, 8 It has been reported that sAPPα may exert its neuroprotective effect by activation of the nuclear factor-kappa B (NF-κB) signalling pathway.4 Our microarray analysis of global changes in gene expression revealed APP-mediated induction of MnSOD, which has been described as an NF-κB target gene.28 In addition, APP activated expression of TPO1, genes regulating protein folding (mt-hsp70/mortalin) and ATP production (cytochrome-C) in mitochondria, as well as the ER chaperone calnexin. Interestingly, TPO1 was suggested to be a key player in myelin biogenesis,29 linking APP to protection from white matter degeneration known to occur in AD.30 Disturbance of the oxidative stress response (MnSOD) and mitochondrial function (mt-hsp70/mortalin) has been suggested to play an important role in the pathogenesis of AD.31 Calnexin serves both as a molecular chaperone and a calcium ion storage protein.32 In light of the fact that ER Ca2+ store depletion has been implicated in UV-triggered apoptosis33 and the pathophysiology of AD,34 induction of proteins regulating ER Ca2+ homeostasis might be an underlying mechanism of the observed neuroprotective effects of APP.
It is to be noted that our microarray analysis suggested that overexpression of APP provoked a pronounced inhibition of c-Jun/AP-1 target gene expression. This downregulation was associated with a potent reduction in JNK activity. Interestingly, it was recently demonstrated that inhibition of c-Jun by a dominant-negative c-Jun peptide also leads to activation of MnSOD expression.35 This suggests that in addition to its implication in the NF-κB pathway (see above), APP-mediated activation of MnSOD might be causally linked to the observed inhibition of the JNK signalling pathway. The proposed c-Jun/AP-1 target gene SPARC/osteonectin belongs to the group of matricellular proteins. Components of the extracellular matrix (ECM) mediate neuron–glia interactions, thus stabilizing the neuronal–astroglial network. It is to be noted that expression of SPARC/osteonectin has been shown to weaken cellular adhesion to the ECM by counteracting integrin–ECM interactions.36 By controlling the level of SPARC/osteonectin expression, APP may counteract neurodegeneration and neuronal cell death induced by matrix detachment (anoikis) and loss of cell/cell contacts. It is tempting to speculate that downmodulation of SPARC/osteonectin may be linked to AD-associated neuronal atrophy and progressive loss of neurons during the disease.
Disruption of the JNK/c-Jun signalling pathway has been shown to alleviate apoptosis triggered by UV, a classical stimulus of the JNK signalling pathway.23, 24 Following UV irradiation, we observed a potent APP-mediated inhibition of JNK activation, which was accompanied with repression of c-Jun and significant protection from UV-induced cell death. In another study, APP was shown to inhibit neuronal cell death induced by nerve growth factor (NGF)- deprivation,37 which is known to require activation of the JNK pathway.38 Collectively, these data suggest that following neuronal insults, APP may exert neuroprotective effects by blocking activation of the proapoptotic JNK signalling cascade. MLKs are known to be essential upstream components of UV-induced JNK activation.39 Indeed, application of the MLK inhibitor CEP-11004 almost completely abolished UV-induced c-Jun phosphorylation in PC12 neo cells. CEP-11004 also significantly reduced UV-induced apoptosis in PC12 neo cells to a level comparable to that observed in PC12 APPwt cells. Thus, our data suggest that APP-mediated protection from UV-triggered cell death may to a large extent be attributable to the observed inhibition of JNK activation. It is to be noted that JNKs can also activate apoptosis by transcription-independent mechanisms. For example, it has been shown that JNKs activate translocation of Bax and Bim to the mitochondria.40, 41
Increasing evidence points to a pivotal role of the JNK signalling pathway in the development of neurodegenerative diseases, such as AD.42, 43 In vitro exposure of neurons to Aβ peptides has been shown to induce cell death via activation of the JNK signalling cascade.44, 45 Enhanced JNK activation has also been demonstrated in degenerating neurons of post-mortem brain sections from AD patients.45, 46 It was recently demonstrated that the C-terminal domain of APP is physically bound to the adapter protein JIP-1b,10, 11, 12 which serves as a scaffolding protein for JNKs and upstream kinases such as MKK4, MKK7 and MLKs. Interestingly, our experiments revealed that the Swedish APP mutant did not inhibit stress-induced JNK signalling. Full-length APPsw was previously shown to bind JIP1b with a similar efficiency as APPwt,10 suggesting that APP-mediated JNK inhibition is not due to sequestration of JIP-1b by APP and prevention of scaffolding of JNKs and their upstream kinases. Furthermore, the inhibitory effect on JNK signalling could not be restored by application of a γ-secretase inhibitor in PC12 APPsw cells, indicating that extracellular sAPPα rather than AICD might be the proteolytic cleavage product responsible for antagonizing JNK signalling. Although sAPPα has been described as an injury-induced neurotrophic factor,47 the receptor that is required for transducing its neuroprotective effect has not been identified so far. However, it has been shown that APPwt, but not mutant APP, can activate the antiapoptotic PI3K/Akt pathway,48 which is known to block the JNK pathway via phosphorylation of MLK3.49 Furthermore, JNK activation increases non-amyloidogenic α-secretase cleavage of APP,50 indicating the possible existence of a negative feedback loop between APP and the JNK pathway after neuron damage. In light of the observed inhibitory effect of APPwt on JNKs, overactivation of the JNK signalling pathway in AD44, 45, 46 might reflect a potential loss of the physiological function of APP. Parallel to this loss of function, enhanced Aβ production might further enhance JNK activation, oxidative stress and mitochondrial dysfunction in AD.
Materials and Methods
Materials
Acetyl-DEVD-7-amido-4-methylcoumarin (Ac-DEVD-AMC) was purchased from Bachem (Heidelberg, Germany). The MLK inhibitor CEP-11004 was a generous gift from Cephalon Inc. (West Chester, USA).26 The γ-secretase inhibitor 1-(S)-endo-N-(1,3,3)-trimethylbicyclo[2.2.1]hept-2-yl)-4-fluorophenyl sulfonamide21 was from Merck Biosciences (Schwalbach, Germany). All other biochemicals and chemicals came in analytical grade purity from Roche Diagnostics (Mannheim, Germany) or Roth (Karlsruhe, Germany).
Cell culture
PC12 cell lines expressing equal amounts of the human wild-type APP695 isoform (PC12 APPwt, clones M5 and N10), PC12 cell lines expressing the Swedish mutant of APP, KM670/671NL (PC12 APPsw, clones Q8 and Q7), as well as vector-transfected control cells (PC12 neo, clones O1 and O3) have been described previously.16, 17, 18 Cell lines were cultivated in DMEM supplemented with 5% heat-inactivated fetal calf serum, 10% horse serum, 100 U/ml penicillin and 100 μg/ml streptomycin.
UV irradiation of cells
For UV irradiation of cells, the culture medium was aspirated, and cells were exposed to a dose of 100 J/m2 UV in a Stratalinker® UV Crosslinker (Stratagene, Amsterdam, The Netherlands). After UV irradiation, the culture medium was immediately returned to the respective cultures.
Measurement of caspase 3-like protease activity
Cells were lysed in 200 μl lysis buffer (10 mM HEPES, pH 7.4, 42 mM KCl, 5 mM MgCl2, 1 mM phenylmethylsulfonyl fluoride (PMSF), 0.1 mM EDTA, 0.1 mM EGTA, 1 mM dithiothreitol (DTT), 1 μg/ml pepstatin A, 1 μg/ml leupeptin, 5 μg/ml aprotinin, 0.5% 3-(3-cholamidopropyldimethylammonio)-1-propane sulfonate (CHAPS)). A 50 μl portion of this lysate was added to 150 μl reaction buffer (25 mM HEPES, 1 mM EDTA, 0.1% CHAPS, 10% sucrose, 3 mM DTT, pH 7.5) and 10 μM of the fluorigenic substrate Ac-DEVD-AMC. This substrate is preferentially cleaved by caspase 3, and to a lesser extent by caspases 6, 7, 8, 9, and 10.51 Accumulation of AMC fluorescence was monitored over 1 h using an HTS fluorescent plate reader (excitation 380 nm, emission 465 nm). Fluorescence of blanks containing no cell lysate was subtracted from the values. Protein content was determined using the Pierce Coomassie Plus Protein Assay reagent (KMF, Cologne, Germany). Caspase activity is expressed as change in fluorescence units per microgram protein per hour.
Quantification of apoptosis
Chromatin condensation and fragmentation, which are nuclear hallmarks of apoptosis, were visualized using the DNA-binding fluorescent dye Hoechst 33258 (Sigma, Deisenhofen, Germany). Cells cultured on 24-well plates were UV-irradiated and stained live with Hoechst 33258 at a final concentration of 1 μg/ml. After incubation for 10 min, nuclear morphology was observed using an Eclipse TE 300 inverted microscope (Nikon, Düsseldorf, Germany) and a × 20 dry objective with the following optics: excitation, 340–380 nm; dichroic mirror, 400 nm; emission, 435–485. Digital images were acquired with a SPOT-2 camera using SPOT software version 2.2.1 (Diagnostic Instruments, Sterling Heights, MI, USA). For each time point and treatment, a total number of 300 cells were analyzed for apoptotic morphology in three subfields of each culture. All experiments were performed at least three times with similar results.
Western blot
A 30 μg portion of whole cell lysate (lysis buffer: 68.5 mM Tris/HCl pH 6.8, 2% SDS, 10% glycerol, and protease inhibitors) was loaded onto 12 or 15% SDS–polyacrylamide gels. Proteins were separated at 120 V and then blotted to nitrocellulose membranes (Protean BA 83; 2 μm; Schleicher & Schuell, Dassel, Germany) in Towbin buffer (25 mM Tris, 192 mM glycine, 20% methanol (v/v) and 0.01% SDS) at 18 V for 50 min. The blots were blocked in blocking buffer (5% non-fat milk, 15 mM Tris-HCl pH 7.5, 200 mM NaCl and 0.1% Tween 20) at room temperature for 2 h. Major isoforms of JNKs are p46 JNK1 and p54 JNK2.
For detection of activated JNK1 and JNK2, we diluted the rabbit polyclonal anti-ACTIVE® JNK antiserum (Promega, Heidelberg, Germany) 1 : 5000 in Tris-buffered saline (TBS) containing 1% bovine serum albumin (BSA) and 0.1% Tween 20. Primary antibodies were detected by subsequent incubation with a biotinylated anti-rabbit antibody (1 : 2000; Pharmingen Becton Dickinson, Hamburg, Germany) in blocking solution (TBS containing 3% non-fat milk powder and 0.1% Tween 20) and a horseradish peroxidase (HRP)-conjugated anti-biotin antibody (1 : 2000; New England Biolabs, Frankfurt, Germany) in blocking solution. Activation of c-Jun and effector caspases was detected with a polyclonal phospho-c-Jun (Ser63) II antibody (New England Biolabs, Frankfurt, Germany), and a rabbit polyclonal anti-active p20 caspase 3 antiserum (Merck-Frosst, Kirkland, Canada; 1 : 1000 dilution). c-Jun protein levels were detected with the monoclonal c-jun (ab-3) antibody (Merck Biosciences). To ensure equal loading of the samples, membranes were incubated with a mouse monoclonal anti-JNK1 antibody (clone 333, 1 : 1000; BD Biosciences, Hamburg, Germany). All antibodies were diluted in blocking solution and blots were incubated overnight at 4°C. Primary antibodies were detected using HRP-conjugated goat anti-mouse or anti-rabbit IgG (Promega, Heidelberg, Germany) used at 1 : 5000 in blocking solution. Blots were developed using the Pierce SuperSignal substrate chemiluminescence reagent.
In vitro kinase assays
For determination of JNK activity, a SAPK/JNK Assay Kit was employed (New England Biolabs, Frankfurt, Germany). Briefly, PC12 clones were seeded in six-well plates and UV-irradiated as described. Lysates of UV-treated cultures and control cultures were prepared and incubated with c-Jun fusion beads as described by the manufacturer. After incubation of the obtained pellets in kinase buffer supplemented with 100 μM ATP for 30 min, kinase reactions were terminated by addition of SDS sample buffer. Reactions were subjected to SDS–PAGE and Western blotting with a Phospho-Ser63 c-Jun antibody and an HRP-conjugated anti-rabbit secondary antibody. Blots were developed using the Pierce SuperSignal substrate chemiluminescence reagent.
Microarray analysis
Total RNA of cell lines PC12 neo O1, PC12 APP M5 and PC12 APP N10 was extracted using the RNeasy Midi Kit (Qiagen, Hilden, Germany). cDNA was synthesized from 20–40 μg of the total RNA by using the Superscript Choice Kit (Invitrogen Life Technologies, Karlsruhe, Germany). The cRNA was prepared and biotin-labelled by in vitro transcription (Enzo Biochemical, Farmingdale, NY, USA). Labelled cRNA was fragmented by incubation at 94°C for 35 min in the presence of fragmentation buffer (40 mM Tris acetate, pH 8.1, 100 mM potassium acetate and 30 mM magnesium acetate). A 15 μg portion of fragmented cRNA was hybridized for 16 h at 45°C to a rat U34A array (Affymetrix, Santa Clara, CA, USA). After hybridization, the gene chips were automatically washed and stained with streptavidin–phycoerythrin by using a fluidics station. Finally, probe arrays were scanned at 3 μm resolution using the Genechip System confocal scanner (Aligent Technologies, Palo Alto, CA, USA). The Affymetrix Microarray Suite 5.0 was used to scan and analyze the relative abundance of each gene from the average difference of intensities. For each individual gene, the signal ratio between perfect match and mismatch probe cells was used to determine its ‘Absolute Call’, indicating whether the corresponding transcript was present (P), absent (A) or marginal (M). For analysis of differential target gene expression, the ‘Difference Call Decision Matrix’ was employed. Individual transcript levels were ranked as either increased (I), marginally increased (MI), decreased (D), marginally decreased (MD) or not changed (NC). To be considered as differentially regulated, the Absolute Call of individual transcripts had to be present or marginal, and the Difference Call increased/decreased or marginally increased/marginally decreased. The gene identity of the obtained EST sequences represented on the Chips is referring to the assignment given on the Affymetrix Homepage, NetAffx Analysis Center (www.affymetrix.com). EST sequences without given gene identities were analyzed by PubMed database searches (standard nucleotide–nucleotide BLAST (blastn), www.ncbi.nlm.nih.gov).52
Semiquantitative RT-PCR
A 1 μg portion of total RNA was used in an RT reaction together with 2.5 μg of random hexamers and dNTPs at a final concentration of 0.5 mM. After heating to 65°C for 5 min, the mixtures were chilled on ice and supplemented with reaction buffer, DTT (final concentration 10 mM) and 200 U Moloney murine leukemia virus reverse transcriptase (Invitrogen Life Technologies). First-strand cDNA synthesis was carried out at 42°C for 50 min. Reactions were stopped by incubation at 70°C for 15 min. cDNAs were subsequently analyzed for expression of putative APP target genes with GAPDH serving as internal control. PCR reactions were performed with Taq polymerase (Eppendorf, Hamburg, Germany) according to the instructions of the manufacturer. Cycle numbers were 25 for c-Jun, SPARC/osteonectin and GAPDH, 30 for Sel-10 and 35 for MnSOD (the sequences of the employed primers can be obtained upon request from the authors). PCR products were separated by agarose gel electrophoresis (1.5%) and visualized with 0.1% ethidium bromide under UV transillumination using a CCD camera-based gel documentation system (MWG, Ebersberg, Germany). Levels of expression in PC12 APPwt clones relative to PC12 neo O1 controls were quantified densitometrically using the public domain Java image processing program ImageJ.
Statistics
Data are given as means±S.E.M. For statistical comparison, t-test or one-way ANOVA followed by Tukey's test was employed. P-values smaller than 0.05 were considered to be statistically significant.
Abbreviations
- AD:
-
Alzheimer's disease
- AICD:
-
APP intracellular domain
- AP-1:
-
activator protein-1
- APP:
-
amyloid precursor protein
- ECM:
-
extracellular matrix
- JIP-1b:
-
c-Jun N-terminal kinase (JNK)-interacting protein-1b
- JNK:
-
c-Jun N-terminal kinase
- MLK:
-
mixed-lineage kinase
- MnSOD:
-
manganese superoxide dismutase
- NF-κB:
-
nuclear factor-kappa B
- SPARC:
-
secreted protein, acidic and rich in cysteine
References
Selkoe DJ (2001) Alzheimer's disease: genes, proteins, and therapy. Physiol. Rev. 81: 741–766
LaFerla FM (2002) Calcium dyshomeostasis and intracellular signalling in Alzheimer's disease. Nat. Rev. Neurosci. 3: 862–872
Selkoe DJ and Podlisny MB (2002) Deciphering the genetic basis of Alzheimer's disease. Annu. Rev. Genomics Hum. Genet. 3: 67–99
Barger SW and Mattson MP (1996) Induction of neuroprotective kappa B-dependent transcription by secreted forms of the Alzheimer's beta-amyloid precursor. Brain Res. Mol. Brain Res. 40: 116–126
Tanzi RE and Bertram L (2001) New frontiers in Alzheimer's disease genetics. Neuron 32: 181–184
De Strooper B (2003) Aph-1, Pen-2, and Nicastrin with Presenilin generate an active gamma-Secretase complex. Neuron 38: 9–12
Passer B, Pellegrini L, Russo C, Siegel RM, Lenardo MJ, Schettini G, Bachmann M, Tabaton M and D'Adamio L (2000) Generation of an apoptotic intracellular peptide by gamma-secretase cleavage of Alzheimer's amyloid beta protein precursor. J. Alzheimers Dis. 2: 289–301
Cao X and Sudhof TC (2001) A transcriptionally [correction of transcriptively] active complex of APP with Fe65 and histone acetyltransferase Tip60. Science 293: 115–120
Kimberly WT, Zheng JB, Guenette SY and Selkoe DJ (2001) The intracellular domain of the beta-amyloid precursor protein is stabilized by Fe65 and translocates to the nucleus in a notch-like manner. J. Biol. Chem. 276: 40288–40292
Matsuda S, Yasukawa T, Homma Y, Ito Y, Niikura T, Hiraki T, Hirai S, Ohno S, Kita Y, Kawasumi M, Kouyama K, Yamamoto T, Kyriakis JM and Nishimoto I (2001) c-Jun n-terminal kinase (jnk)-interacting protein-1b/islet-brain-1 scaffolds alzheimer's amyloid precursor protein with jnk. J. Neurosci. 21: 6597–6607
Scheinfeld MH, Roncarati R, Vito P, Lopez PA, Abdallah M and D'Adamio L (2002) Jun NH2-terminal kinase (JNK) interacting protein 1 (JIP1) binds the cytoplasmic domain of the Alzheimer's beta-amyloid precursor protein (APP). J. Biol. Chem. 277: 3767–3775
Taru H, Kirino Y and Suzuki T (2002) Differential roles of JIP scaffold proteins in the modulation of amyloid precursor protein metabolism. J. Biol. Chem. 277: 27567–27574
Whitmarsh AJ, Kuan CY, Kennedy NJ, Kelkar N, Haydar TF, Mordes JP, Appel M, Rossini AA, Jones SN, Flavell RA, Rakic P and Davis RJ (2001) Requirement of the JIP1 scaffold protein for stress-induced JNK activation. Genes Dev. 15: 2421–2432
Yasuda J, Whitmarsh AJ, Cavanagh J, Sharma M and Davis RJ (1999) The JIP group of mitogen-activated protein kinase scaffold proteins. Mol. Cell Biol. 19: 7245–7254
Herdegen T, Skene P and Bahr M (1997) The c-Jun transcription factor – bipotential mediator of neuronal death, survival and regeneration. Trends Neurosci. 20: 227–231
Kögel D, Schomburg R, Schurmann T, Reimertz C, König HG, Poppe M, Eckert A, Müller WE and Prehn JH (2003) The amyloid precursor protein protects PC12 cells against endoplasmic reticulum stress-induced apoptosis. J. Neurochem. 87: 248–256
Eckert A, Steiner B, Marques C, Leutz S, Romig H, Haass C and Muller WE (2001) Elevated vulnerability to oxidative stress-induced cell death and activation of caspase-3 by the Swedish amyloid precursor protein mutation. J. Neurosci. Res. 64: 183–192
Marques CA, Keil U, Bonert A, Steiner B, Haass C, Muller WE and Eckert A (2003) Neurotoxic mechanisms caused by the Alzheimer's disease-linked Swedish amyloid precursor protein mutation: oxidative stress, caspases, and the JNK pathway. J. Biol. Chem. 278: 28294–28302
Briggs J, Chamboredon S, Castellazzi M, Kerry JA and Bos TJ (2002) Transcriptional upregulation of SPARC, in response to c-Jun overexpression, contributes to increased motility and invasion of MCF7 breast cancer cells. Oncogene 21: 7077–7091
Minet E, Michel G, Mottet D, Piret JP, Barbieux A, Raes M and Michiels C (2001) c-JUN gene induction and AP-1 activity is regulated by a JNK-dependent pathway in hypoxic HepG2 cells. Exp. Cell Res. 265: 114–124
Rishton GM, Retz DM, Tempest PA, Novotny J, Kahn S, Treanor JJ, Lile JD and Citron M (2000) Fenchylamine sulfonamide inhibitors of amyloid beta peptide production by the gamma-secretase proteolytic pathway: potential small-molecule therapeutic agents for the treatment of Alzheimer's disease. J. Med. Chem. 43: 2297–2299
Mielke K, Damm A, Yang DD and Herdegen T (2000) Selective expression of JNK isoforms and stress-specific JNK activity in different neural cell lines. Brain Res. Mol. Brain Res. 75: 128–137
Noguchi K, Kokubu A, Kitanaka C, Ichijo H and Kuchino Y (2001) ASK1-signaling promotes c-Myc protein stability during apoptosis. Biochem. Biophys. Res. Commun. 281: 1313–1320
Tournier C, Hess P, Yang DD, Xu J, Turner TK, Nimnual A, Bar-Sagi D, Jones SN, Flavell RA and Davis RJ (2000) Requirement of JNK for stress-induced activation of the cytochrome c-mediated death pathway. Science 288: 870–874
Selznick LA, Zheng TS, Flavell RA, Rakic P and Roth KA (2000) Amyloid beta-induced neuronal death is bax-dependent but caspase-independent. J. Neuropathol. Exp. Neurol. 59: 271–279
Murakata C, Kaneko M, Gessner G, Angeles TS, Ator MA, O'Kane TM, McKenna BA, Thomas BA, Mathiasen JR, Saporito MS, Bozyczko-Coyne D and Hudkins RL (2002) Mixed lineage kinase activity of indolocarbazole analogues. Bioorg. Med. Chem. Lett. 12: 147–150
Mielke K and Herdegen T (2002) Fatal shift of signal transduction is an integral part of neuronal differentiation: JNKs realize TNFalpha-mediated apoptosis in neuronlike, but not naive, PC12 cells. Mol. Cell Neurosci. 20: 211–224
Xu Y, Kiningham KK, Devalaraja MN, Yeh CC, Majima H, Kasarskis EJ and St Clair DK (1999) An intronic NF-kappaB element is essential for induction of the human manganese superoxide dismutase gene by tumor necrosis factor-alpha and interleukin-1beta DNA. Cell Biol. 18: 709–722
Krueger WH, Gonye GE, Madison DL, Murray KE, Kumar M, Spoerel N and Pfeiffer SE (1997) TPO1, a member of a novel protein family, is developmentally regulated in cultured oligodendrocytes. J. Neurochem. 69: 1343–1355
Roher AE, Weiss N, Kokjohn TA, Kuo YM, Kalback W, Anthony J, Watson D, Luehrs DC, Sue L, Walker D, Emmerling M, Goux W and Beach T (2002) Increased A beta peptides and reduced cholesterol and myelin proteins characterize white matter degeneration in Alzheimer's disease. Biochemistry 41: 11080–11090
Hirai K, Aliev G, Nunomura A, Fujioka H, Russell RL, Atwood CS, Johnson AB, Kress Y, Vinters HV, Tabaton M, Shimohama S, Cash AD, Siedlak SL, Harris PL, Jones PK, Petersen RB, Perry G and Smith MA (2001) Mitochondrial abnormalities in Alzheimer's disease. J. Neurosci. 21: 3017–3023
Michalak M, Robert Parker JM and Opas M (2002) Ca(2+) signaling and calcium binding chaperones of the endoplasmic reticulum. Cell Calcium 32: 269–278
Pu Y and Chang DC (2001) Cytosolic Ca(2+) signal is involved in regulating UV-induced apoptosis in HeLa cells. Biochem. Biophys. Res. Commun. 282: 84–89
Mattson MP (2000) Apoptosis in neurodegenerative disorders. Nat. Rev. Mol. Cell Biol. 1: 120–129
Holzberg D, Knight CG, Dittrich-Breiholz O, Schneider H, Dorrie A, Hoffmann E, Resch K and Kracht M (2003) Disruption of the c-JUN–JNK complex by a cell-permeable peptide containing the c-JUN delta domain induces apoptosis and affects a distinct set of IL-1-induced inflammatory genes. J. Biol. Chem. 278: 40213–40223
Bornstein P and Sage EH (2002) Matricellular proteins: extracellular modulators of cell function. Curr. Opin. Cell Biol. 14: 608–616
Nishimura I, Takazaki R, Kuwako K, Enokido Y and Yoshikawa K (2003) Upregulation and antiapoptotic role of endogenous Alzheimer amyloid precursor protein in dorsal root ganglion neurons. Exp. Cell Res. 286: 241–251
Eilers A, Whitfield J, Shah B, Spadoni C, Desmond H and Ham J (2001) Direct inhibition of c-Jun N-terminal kinase in sympathetic neurones prevents c-jun promoter activation and NGF withdrawal-induced death. J. Neurochem. 76: 1439–1454
Xu Z, Maroney AC, Dobrzanski P, Kukekov NV and Greene LA (2001) The MLK family mediates c-Jun N-terminal kinase activation in neuronal apoptosis. Mol. Cell Biol. 21: 4713–4724
Tsuruta F, Sunayama J, Mori Y, Hattori S, Shimizu S, Tsujimoto Y, Yoshioka K, Masuyama N and Gotoh Y (2004) JNK promotes Bax translocation to mitochondria through phosphorylation of 14-3-3 proteins. EMBO J. 23: 1889–1899
Lei K and Davis RJ (2003) JNK phosphorylation of Bim-related members of the Bcl2 family induces Bax-dependent apoptosis. Proc. Natl. Acad. Sci. USA 100: 2432–2437
Manning AM and Davis RJ (2003) Targeting JNK for therapeutic benefit: from junk to gold? Nat. Rev. Drug Discov. 2: 554–565
Bozyczko-Coyne D, Saporito MS and Hudkins RL (2002) Targeting the JNK pathway for therapeutic benefit in CNS disease. Curr. Drug Target CNS Neurol. Disord. 1: 31–49
Morishima Y, Gotoh Y, Zieg J, Barrett T, Takano H, Flavell R, Davis RJ, Shirasaki Y and Greenberg ME (2001) Beta-amyloid induces neuronal apoptosis via a mechanism that involves the c-Jun N-terminal kinase pathway and the induction of Fas ligand. J. Neurosci. 21: 7551–7560
Shoji M, Iwakami N, Takeuchi S, Waragai M, Suzuki M, Kanazawa I, Lippa CF, Ono S and Okazawa H (2000) JNK activation is associated with intracellular beta-amyloid accumulation. Brain Res. Mol. Brain Res. 85: 221–233
Zhu X, Raina AK, Rottkamp CA, Aliev G, Perry G, Boux H and Smith MA (2001) Activation and redistribution of c-jun N-terminal kinase/stress activated protein kinase in degenerating neurons in Alzheimer's disease. J. Neurochem. 76: 435–441
Luo JJ, Wallace MS, Hawver DB, Kusiak JW and Wallace WC (2001) Characterization of the neurotrophic interaction between nerve growth factor and secreted alpha-amyloid precursor protein. J. Neurosci. Res. 63: 410–420
Burton TR, Dibrov A, Kashour T and Amara FM (2002) Anti-apoptotic wild-type Alzheimer amyloid precursor protein signaling involves the p38 mitogen-activated protein kinase/MEF2 pathway. Brain Res. Mol. Brain Res. 108: 102–120
Barthwal MK, Sathyanarayana P, Kundu CN, Rana B, Pradeep A, Sharma C, Woodgett JR and Rana A (2003) Negative regulation of mixed lineage kinase 3 by protein kinase B/AKT leads to cell survival. J. Biol. Chem. 278: 3897–3902
Mudher A, Chapman S, Richardson J, Asuni A, Gibb G, Pollard C, Killick R, Iqbal T, Raymond L, Varndell I, Sheppard P, Makoff A, Gower E, Soden PE, Lewis P, Murphy M, Golde TE, Rupniak HT, Anderton BH and Lovestone S (2001) Dishevelled regulates the metabolism of amyloid precursor protein via protein kinase C/mitogen-activated protein kinase and c-Jun terminal kinase. J. Neurosci. 21: 4987–4995
Garcia-Calvo M, Peterson EP, Rasper DM, Vaillancourt JP, Zamboni R, Nicholson DW and Thornberry NA (1999) Purification and catalytic properties of human caspase family members. Cell Death Differ. 6: 362–369
Altschul SF, Gish W, Miller W, Myers EW and Lipman DJ (1990) Basic local alignment search tool. J. Mol. Biol. 215: 403–410
Acknowledgements
We thank Hanni Bähler, Petra Mech, Christiane Schettler and Hildegard Schweers for technical assistance, Drs. Christian Haass and Anne Eckert for cell lines, Drs. Stephen P Trusko and Marcel Leist for CEP-11004 and Dr. Kirsten Mielke for helpful discussions. This work was supported by grants from Alzheimer Forschung Initiative e.V. to DK (AFI Grant 03809) and JHMP (AFI Grant 00808), as well as Science Foundation Ireland (03/RP1/B344) and Interdisciplinary Center for Clinical Research (IZKF), University of Münster (Bundesministerium für Bildung, Wissenschaft, Forschung und Technologie Grant 01 KS 9604/0) to JHMP.
Author information
Authors and Affiliations
Corresponding author
Additional information
Edited by G Melino
Rights and permissions
About this article
Cite this article
Kögel, D., Schomburg, R., Copanaki, E. et al. Regulation of gene expression by the amyloid precursor protein: inhibition of the JNK/c-Jun pathway. Cell Death Differ 12, 1–9 (2005). https://doi.org/10.1038/sj.cdd.4401495
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/sj.cdd.4401495
Keywords
This article is cited by
-
Is brain iron trafficking part of the physiology of the amyloid precursor protein?
JBIC Journal of Biological Inorganic Chemistry (2019)
-
Amyloid Precursor Protein Mediates Neuronal Protection from Rotenone Toxicity
Molecular Neurobiology (2019)
-
URMC-099 facilitates amyloid-β clearance in a murine model of Alzheimer’s disease
Journal of Neuroinflammation (2018)
-
Modulation of BAG3 Expression and Proteasomal Activity by sAPPα Does Not Require Membrane-Tethered Holo-APP
Molecular Neurobiology (2016)
-
Holo-APP and G-protein-mediated signaling are required for sAPPα-induced activation of the Akt survival pathway
Cell Death & Disease (2014)